Sample QC metrics
An overview of quality control (QC) metrics for all samples, both before and after filtering. The QC metrics include:
no. of cells: Total number of barcodes called as cells for each sample.no. of UMIs: Sum of counts across all features for each cell with log-transformed values displayed.no. of genes: Number of unique features with non-zero counts for each cell with log-transformed values displayed.mito pct.: The proportion of reads that mapped to genes in the mitochondrial genome. High proportions are indicative of poor-quality cells (compromised membranes allow individual RNA molecules to escape while retaining mitochondria, leading to an increased relative abundance of mitochondrial transcript) or nuclei (incomplete removal of cytoplasm). Logit-transformed values are displayed as applying the logit (inverse logistic) transformation to mitochondrial proportions yields approximately Gaussian distributions.spliced pct.: The proportion of spliced reads. In single-nuclei RNA sequencing, high spliced proportions may indicate inadequate removal of cytoplasmic material from the nuclei. Logit-transformed values are displayed as applying the logit (inverse logistic) transformation to spliced proportions results in approximately Gaussian distributions.
suppressWarnings({
cat("### Pre-QC\n")
print(plot_qc_ranges_marginals(qc_dt, b_lvls, qc_names, qc_lu, cuts_dt, batch_var))
cat("\n\n")
cat("### After light QC\n")
print(plot_qc_ranges_marginals(kept_dt, b_lvls, qc_names, qc_lu, cuts_dt, batch_var))
cat("\n\n")
})Pre-QC
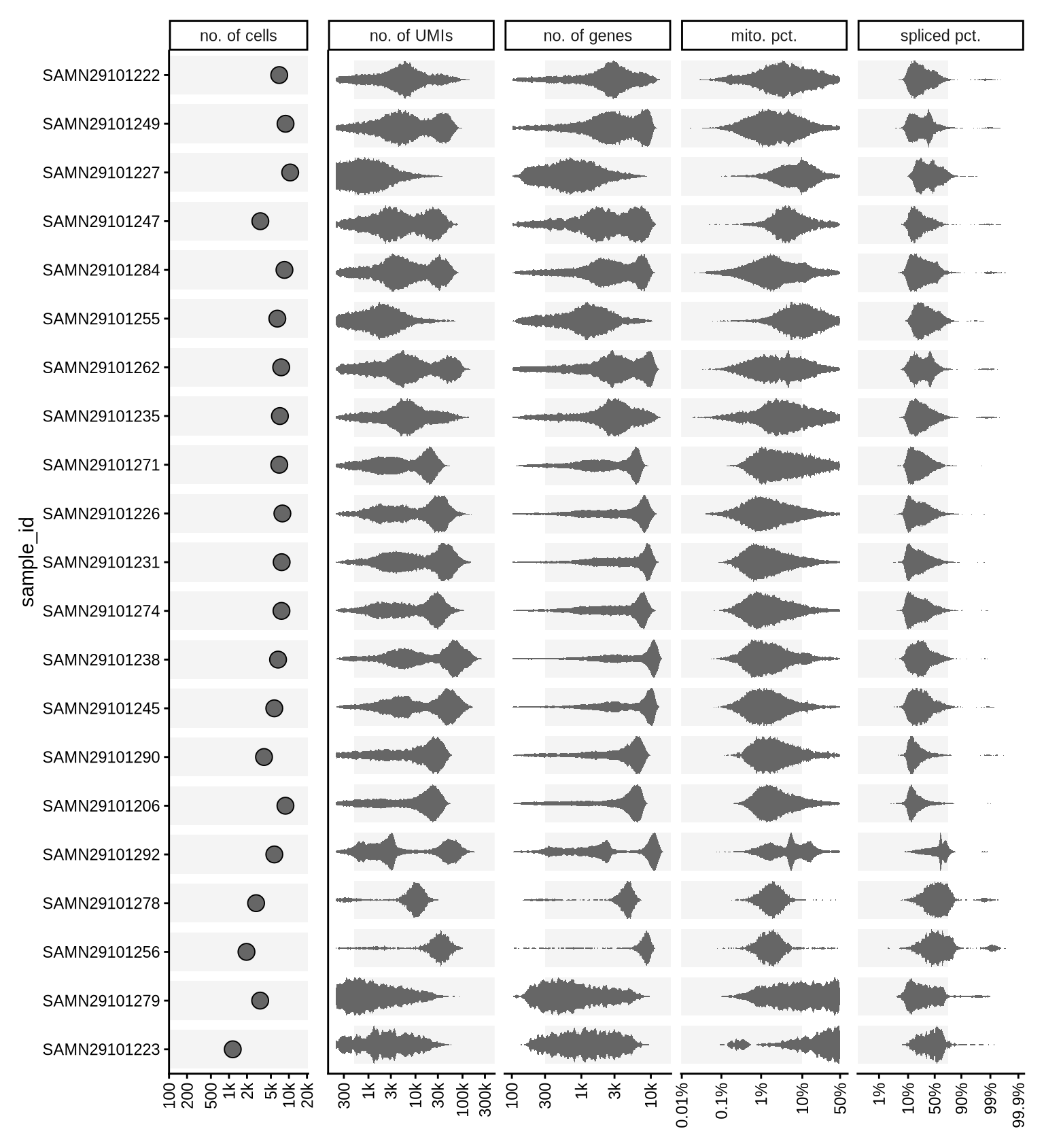
After light QC
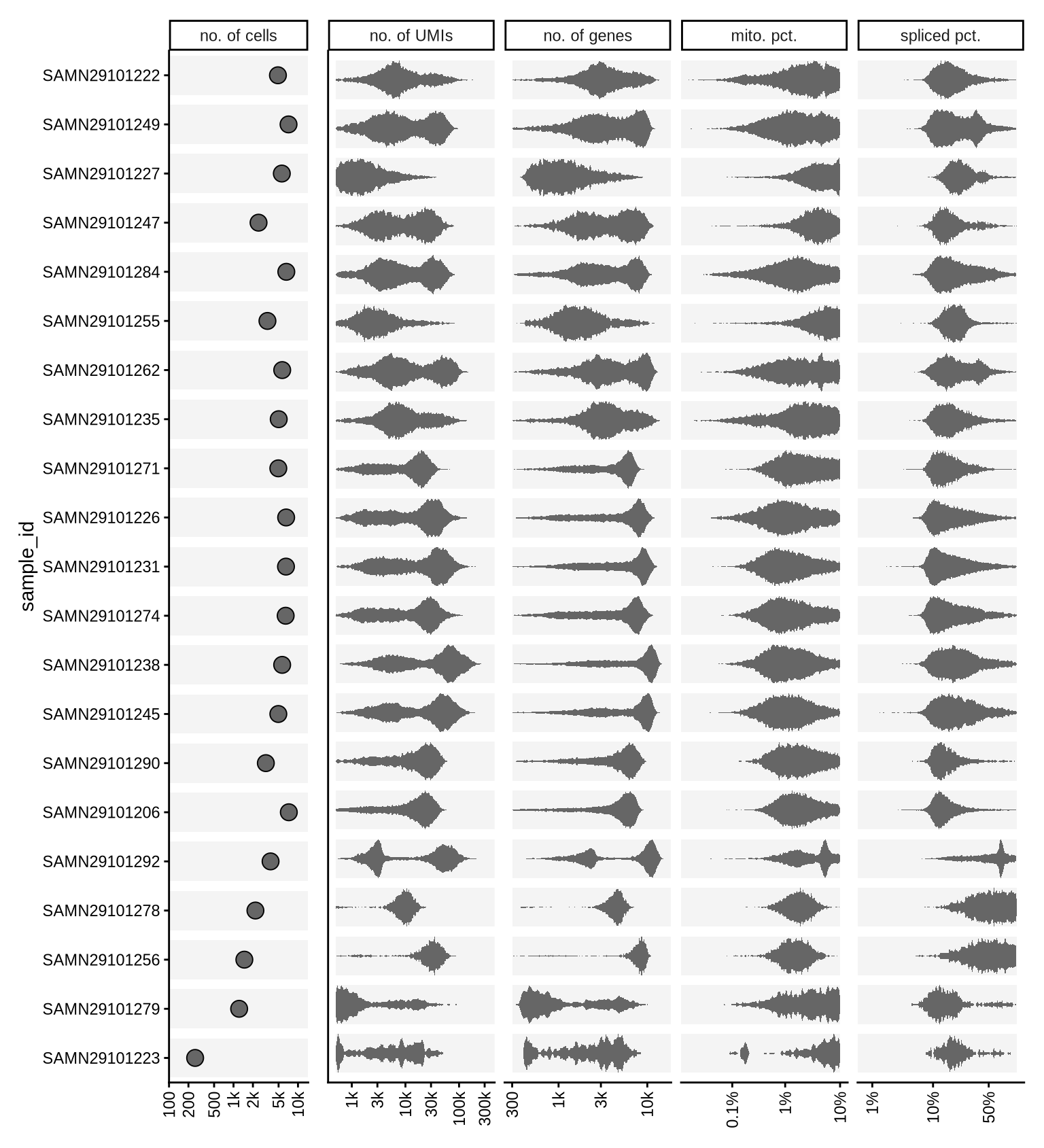
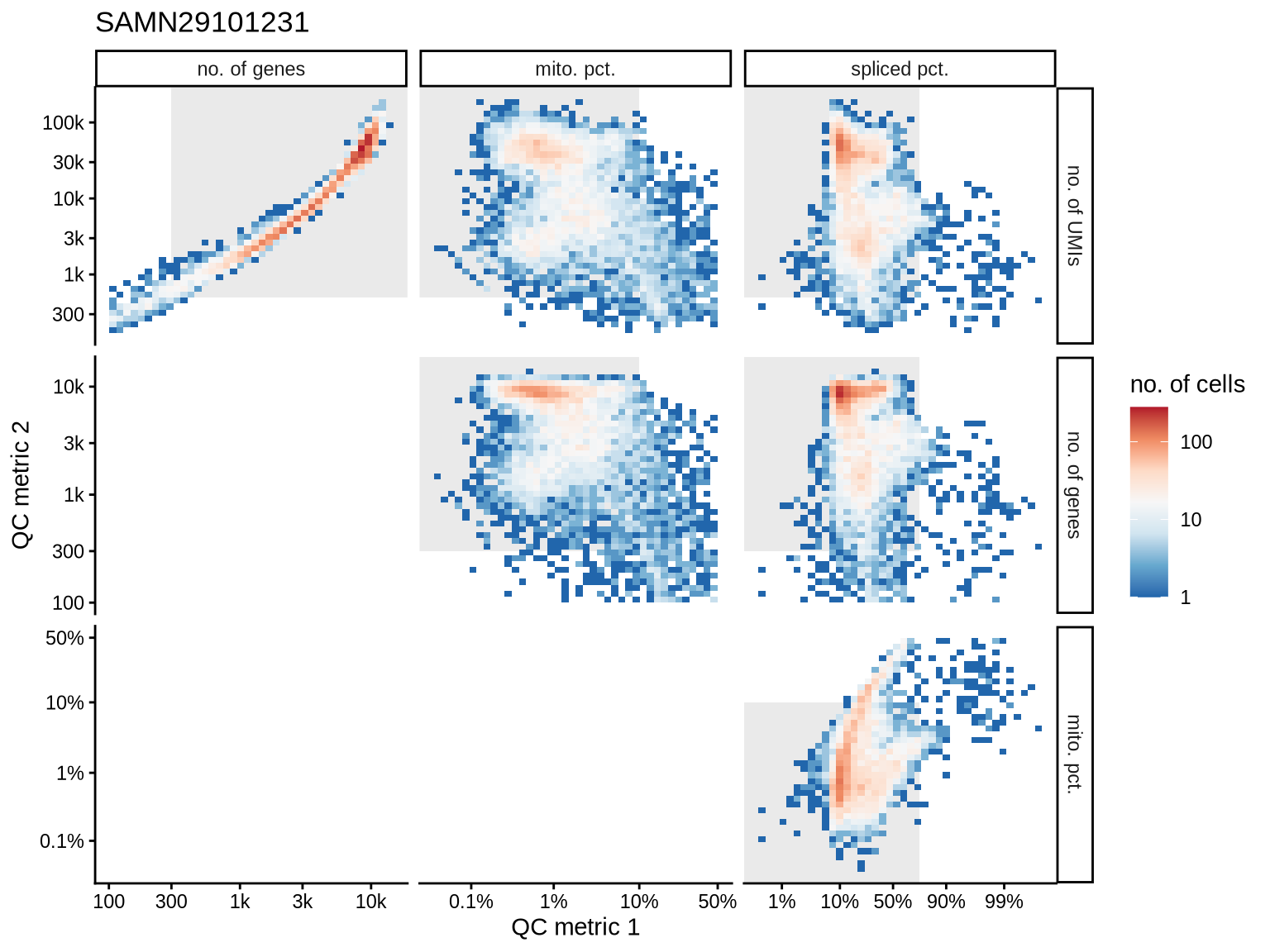
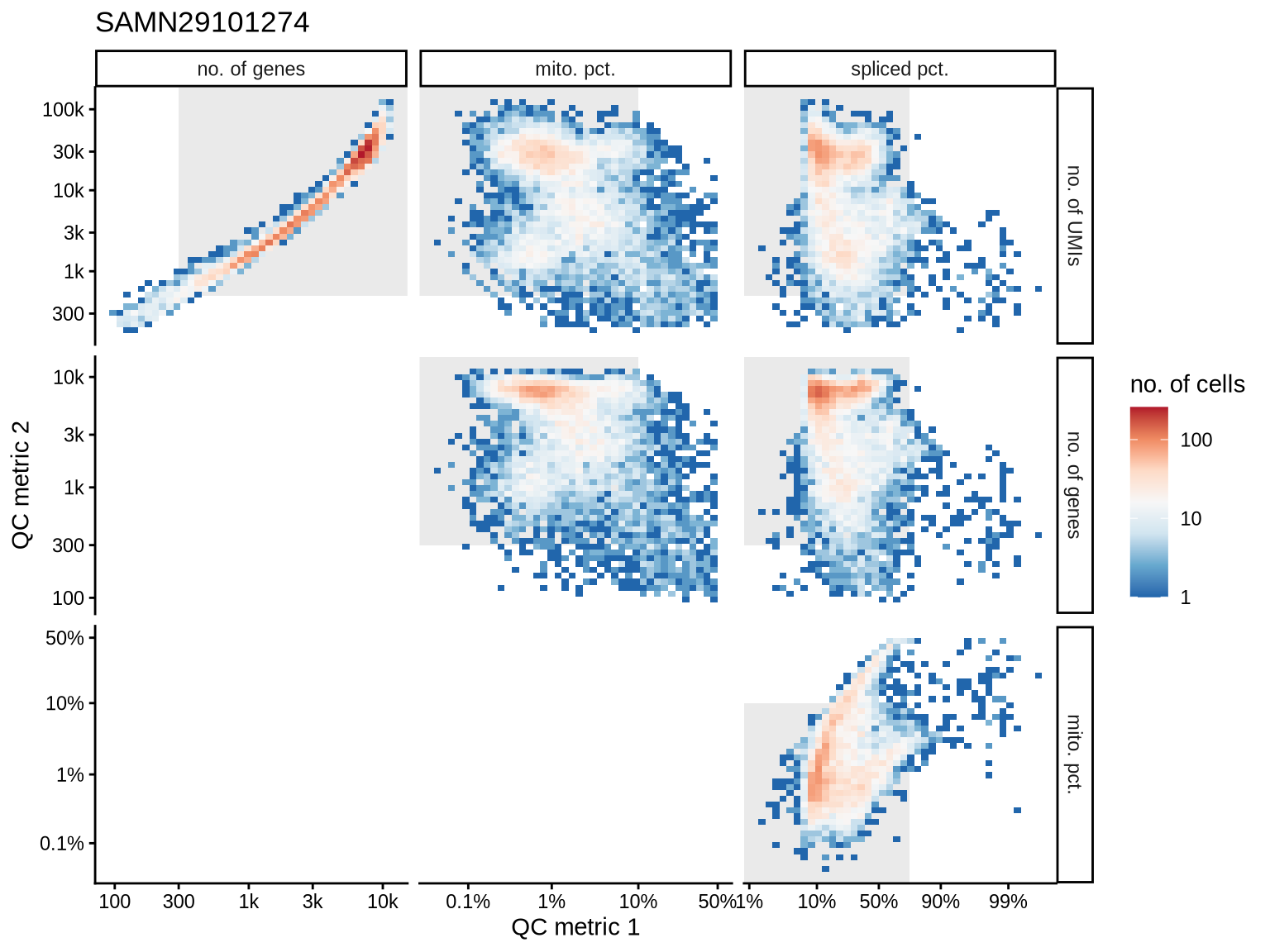
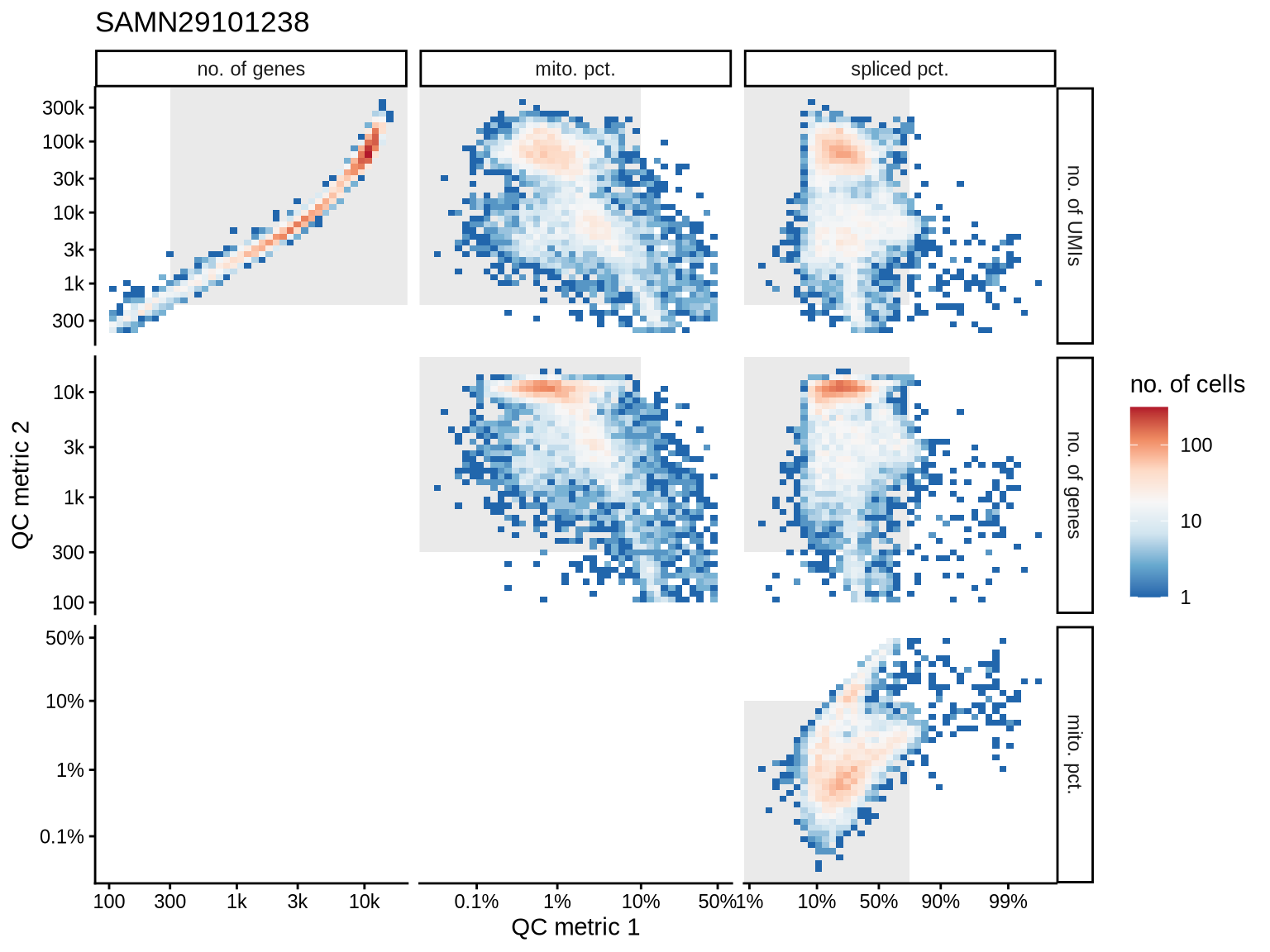
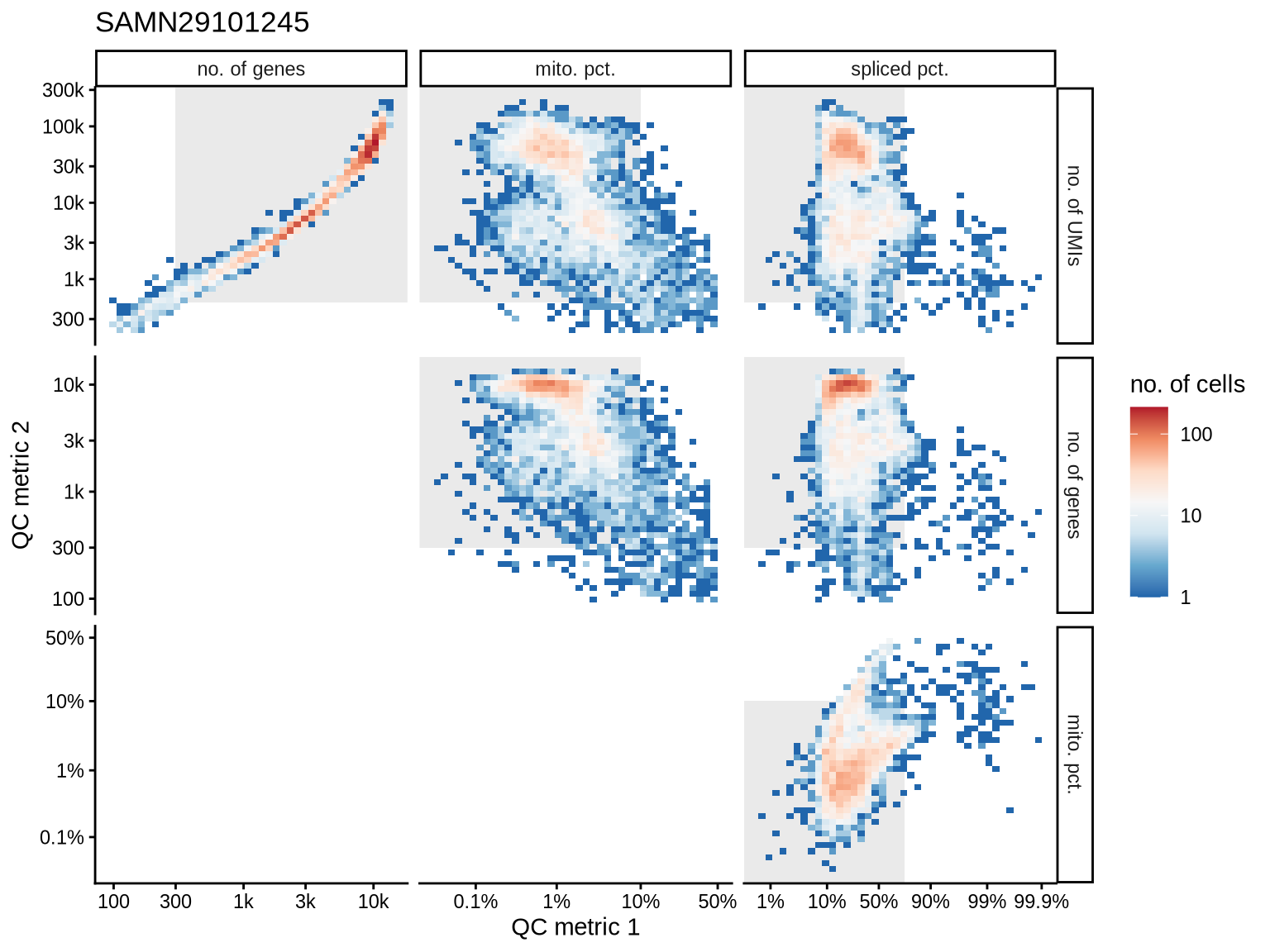
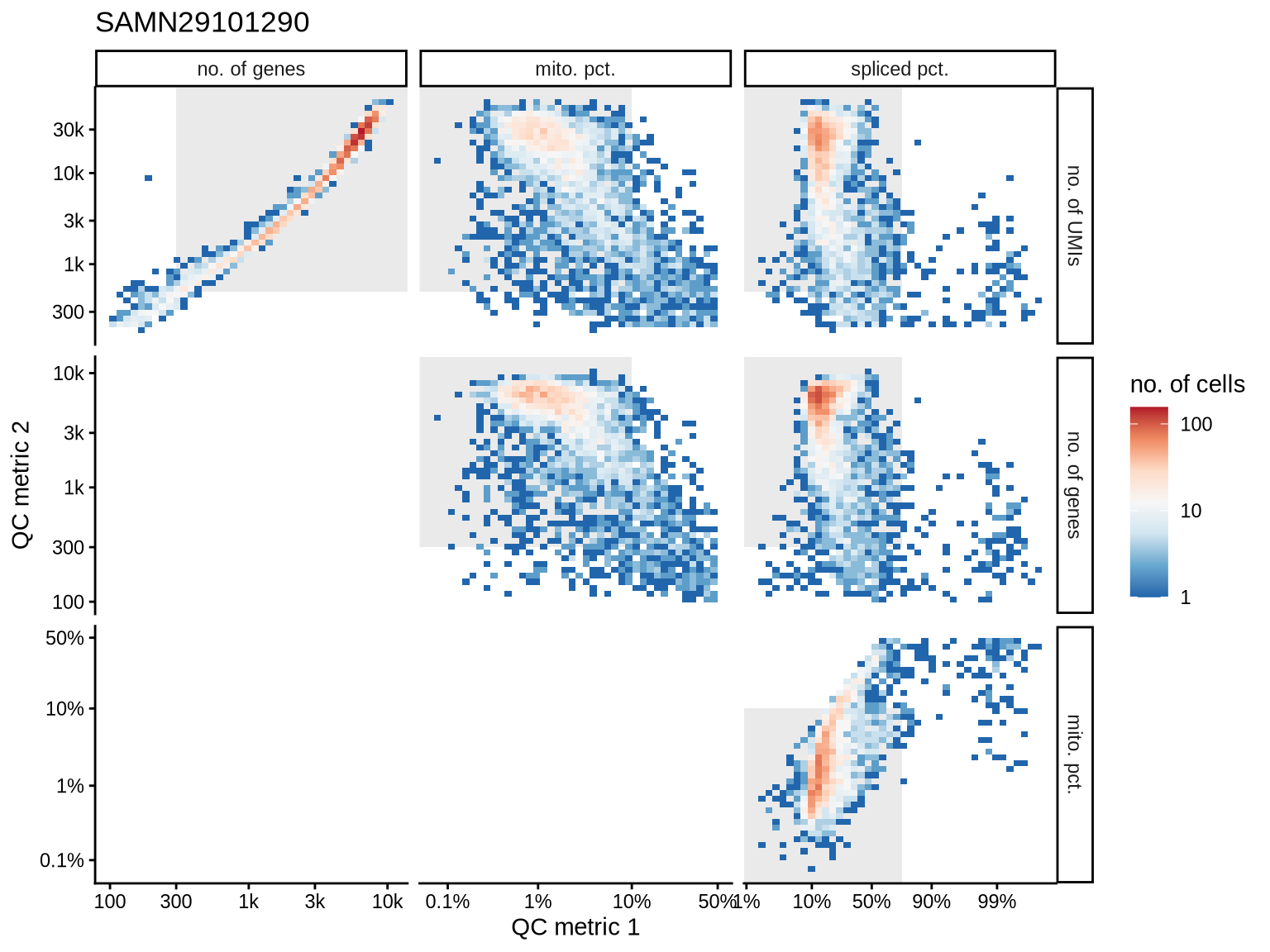
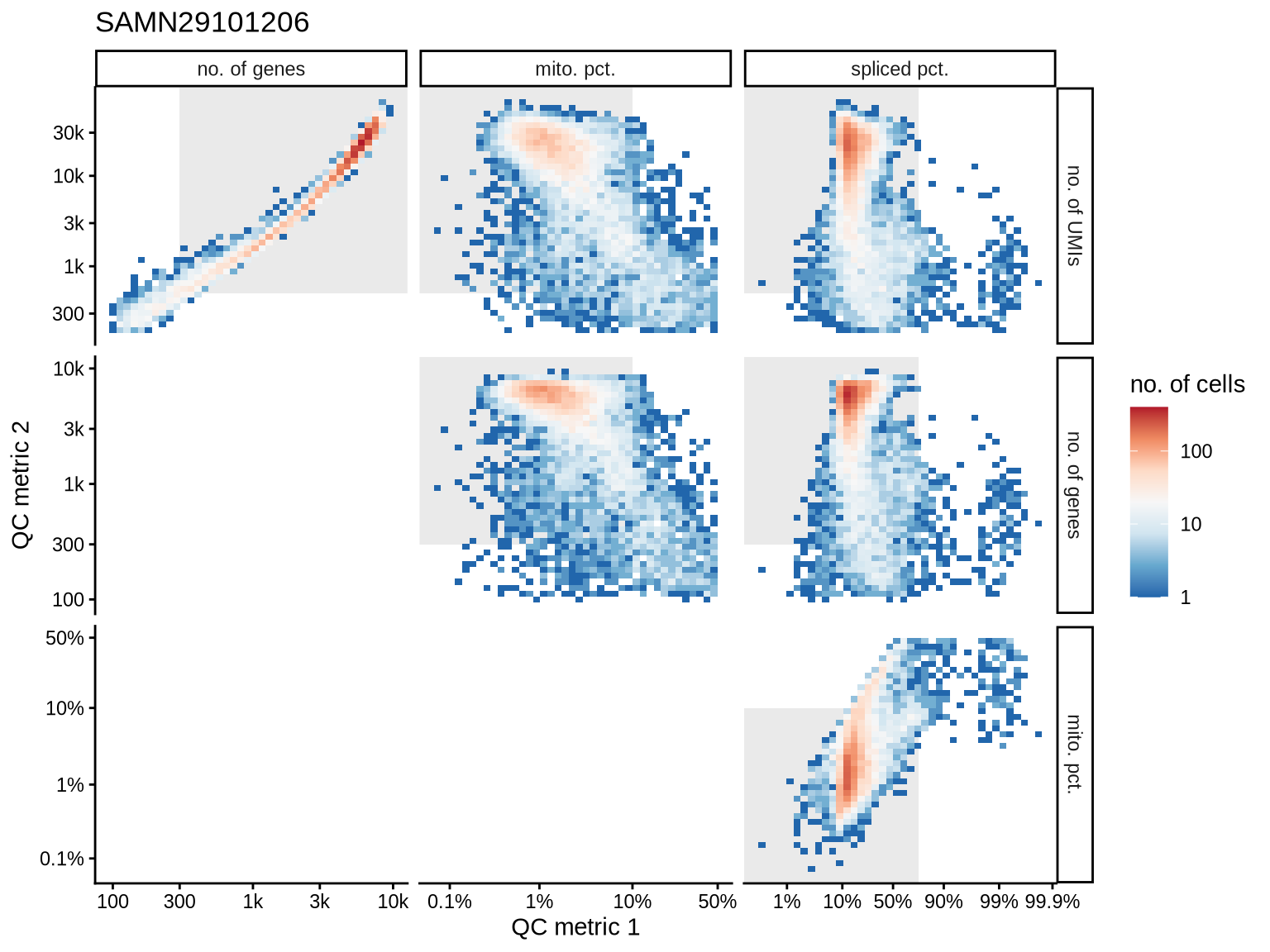
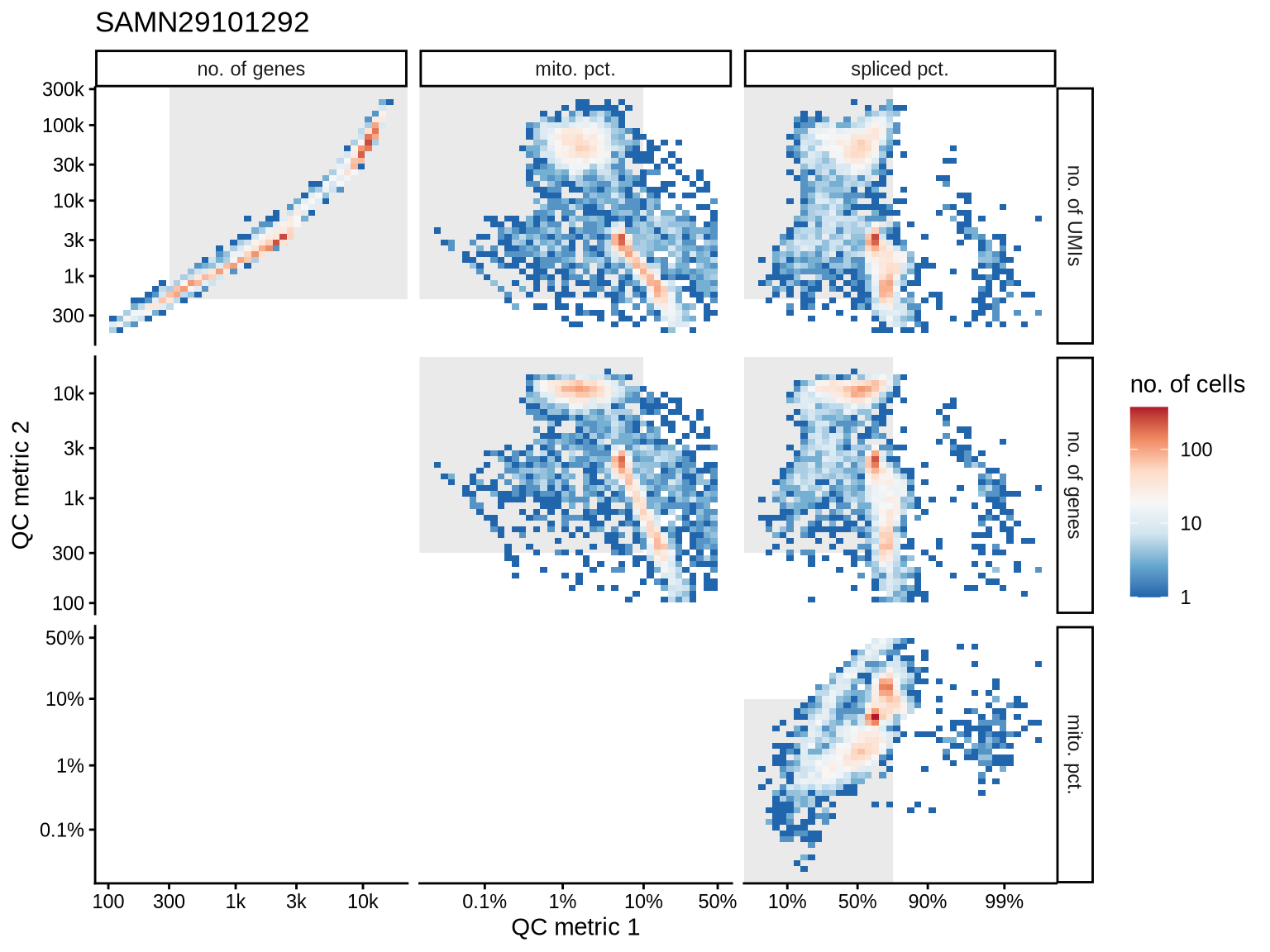
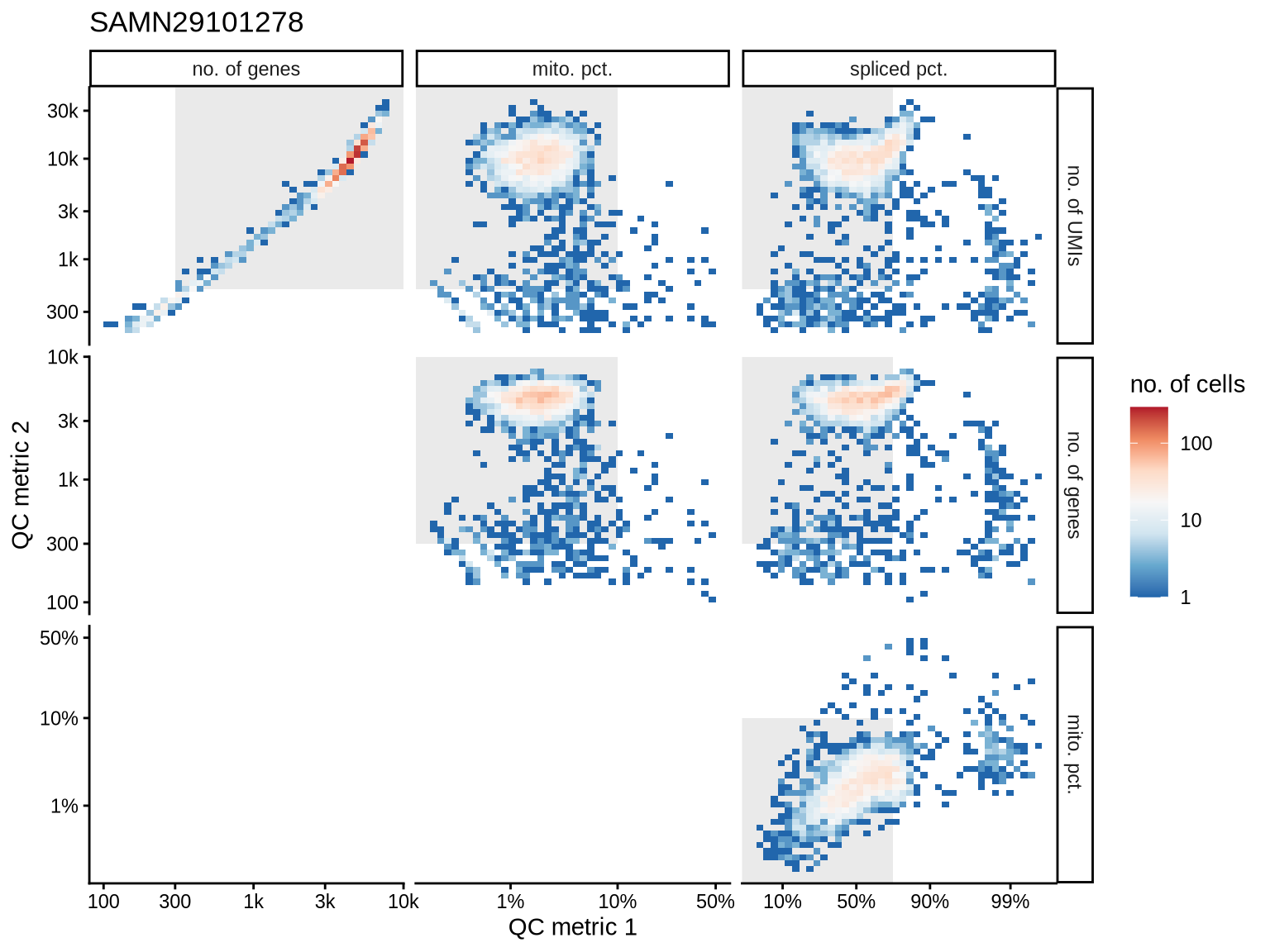
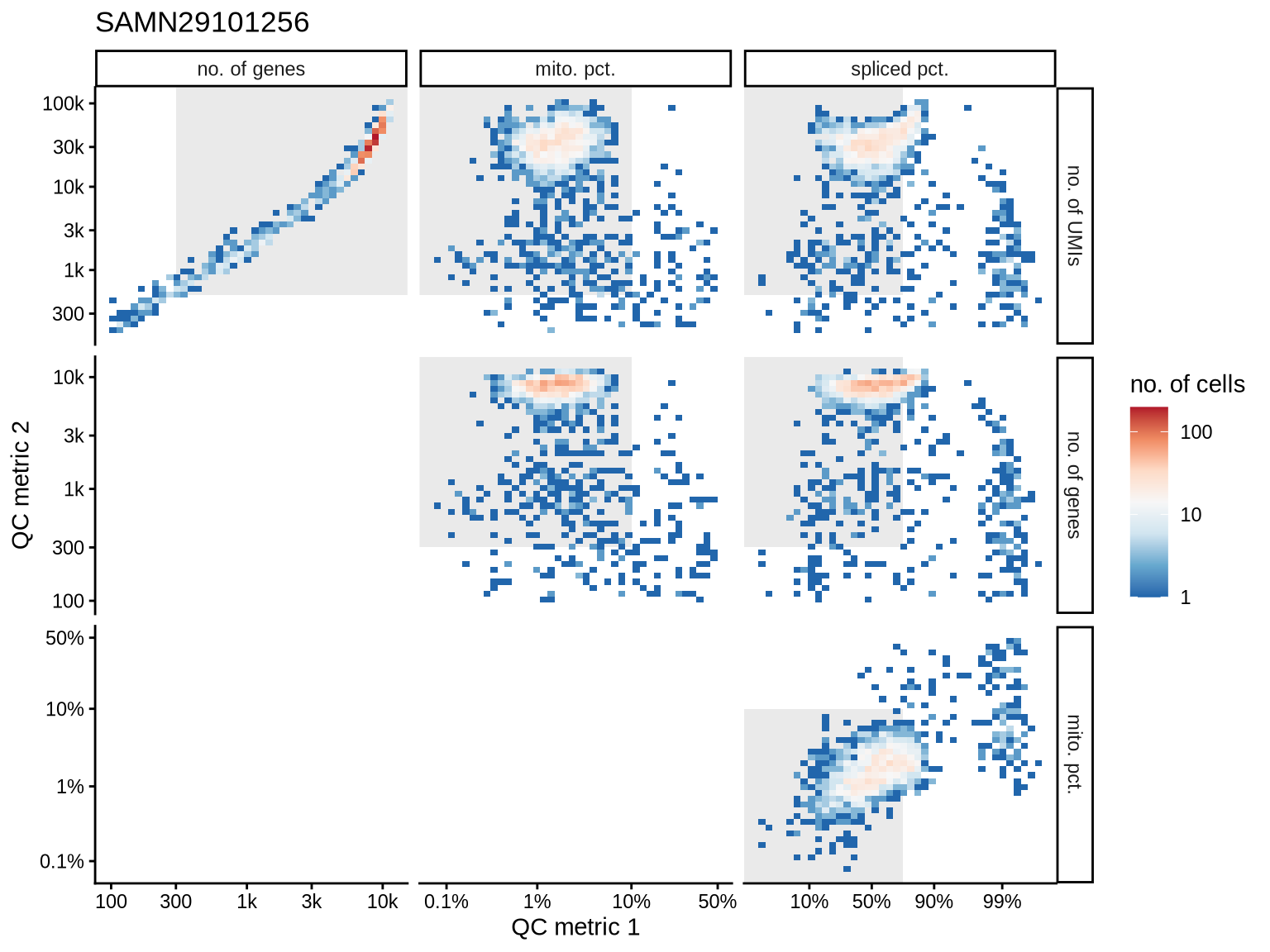
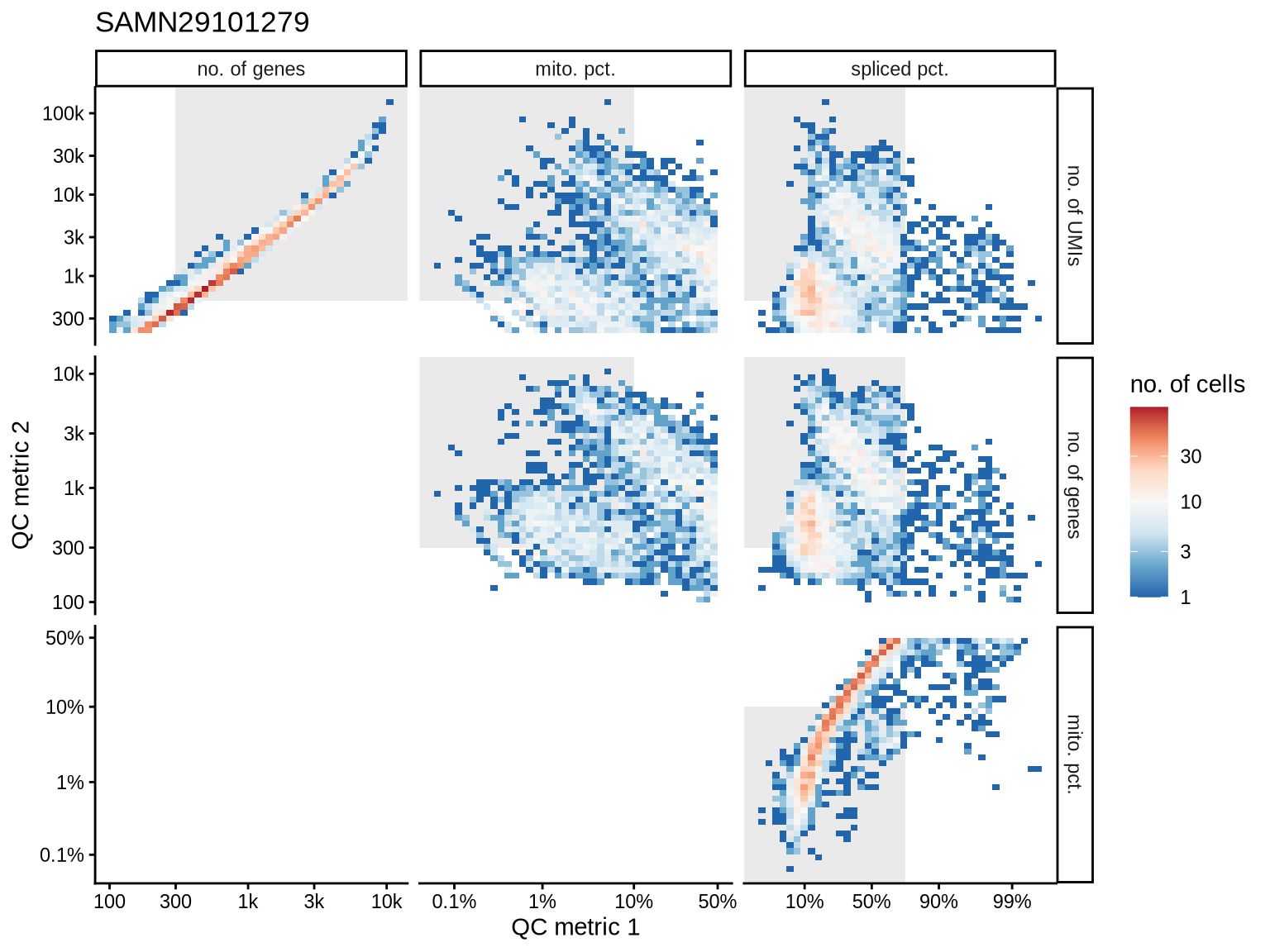
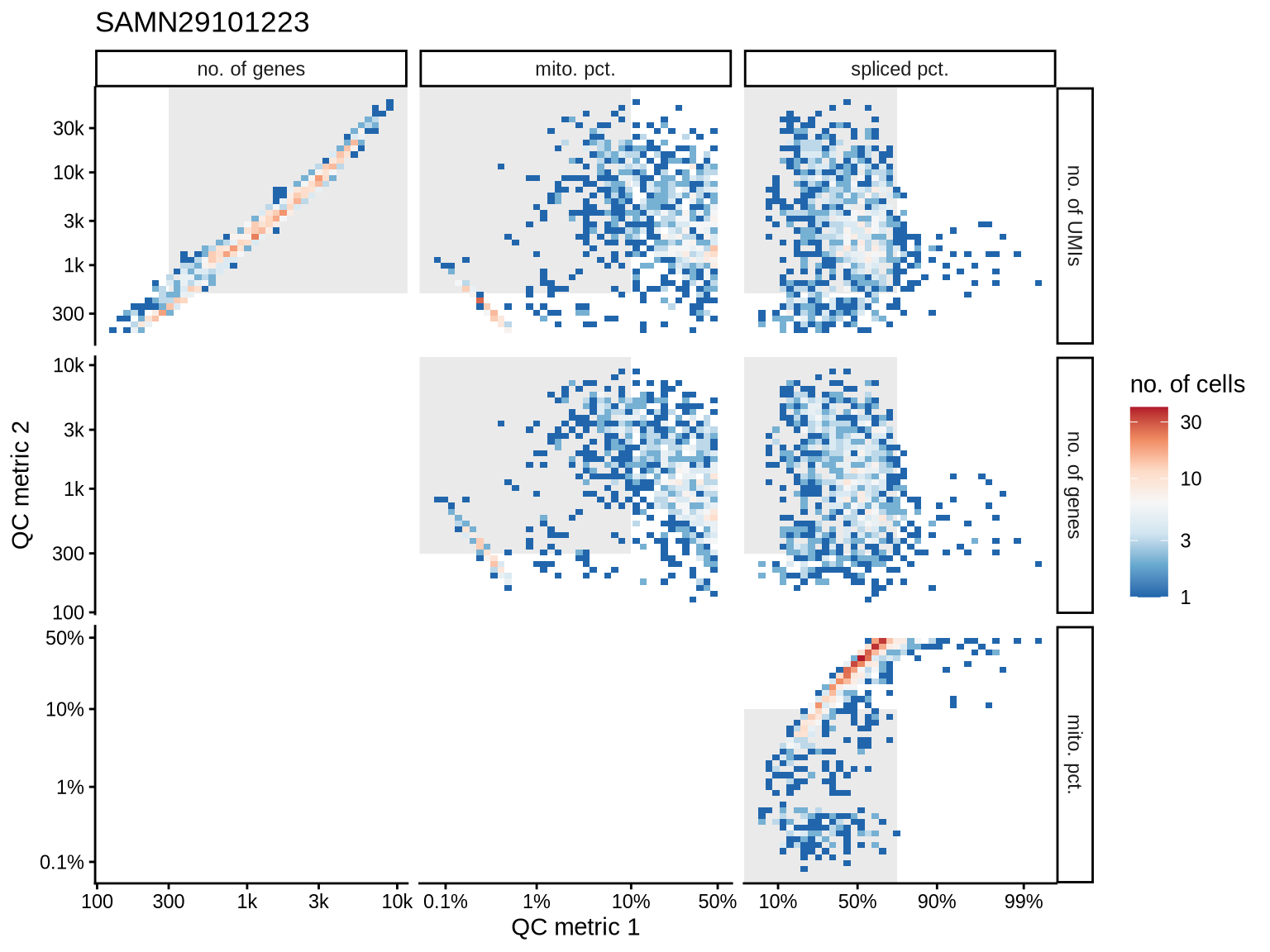
QC metrics scatterplots
Pairwise relationships of QC metrics at individual cell level.
for (b in b_lvls) {
qc_tmp = qc_dt[ batch_var == b ]
if (nrow(qc_tmp) == 1)
next
cat('### ', b, '\n')
print(plot_qc_metric_scatter(qc_tmp, qc_names, qc_lu, cuts_dt[ batch_var == b ], b))
cat('\n\n')
}SAMN29101222
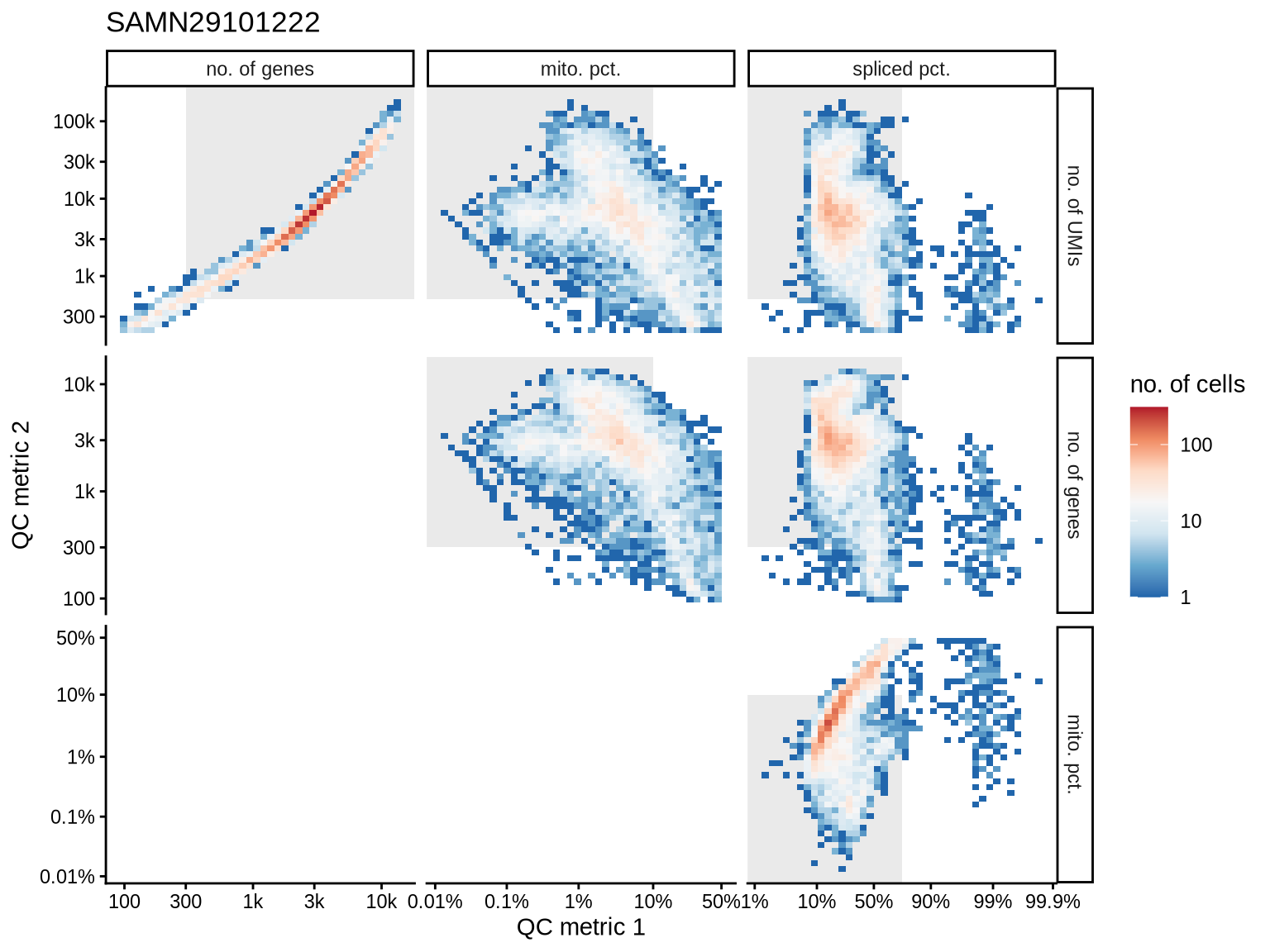
SAMN29101249
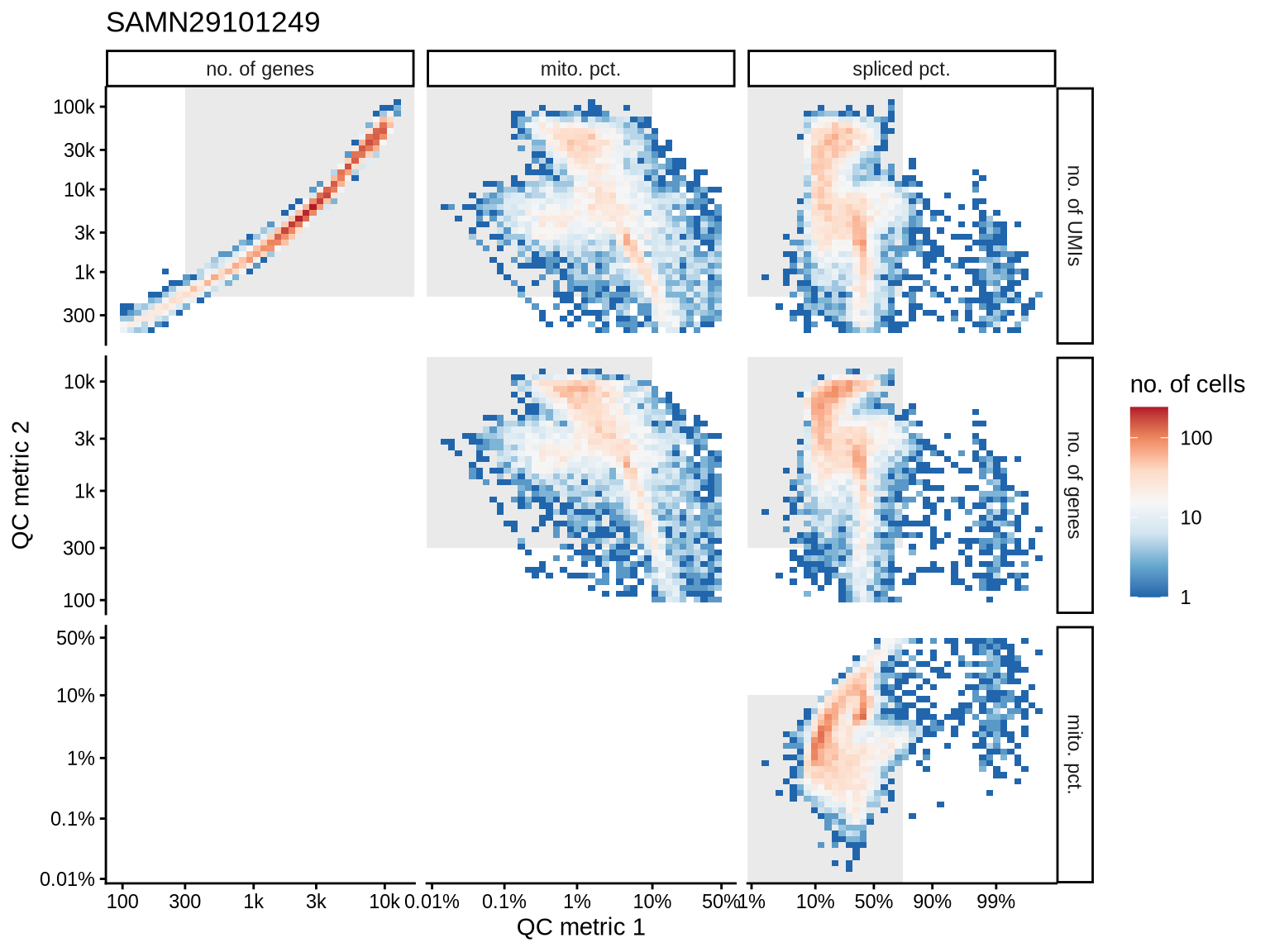
SAMN29101227
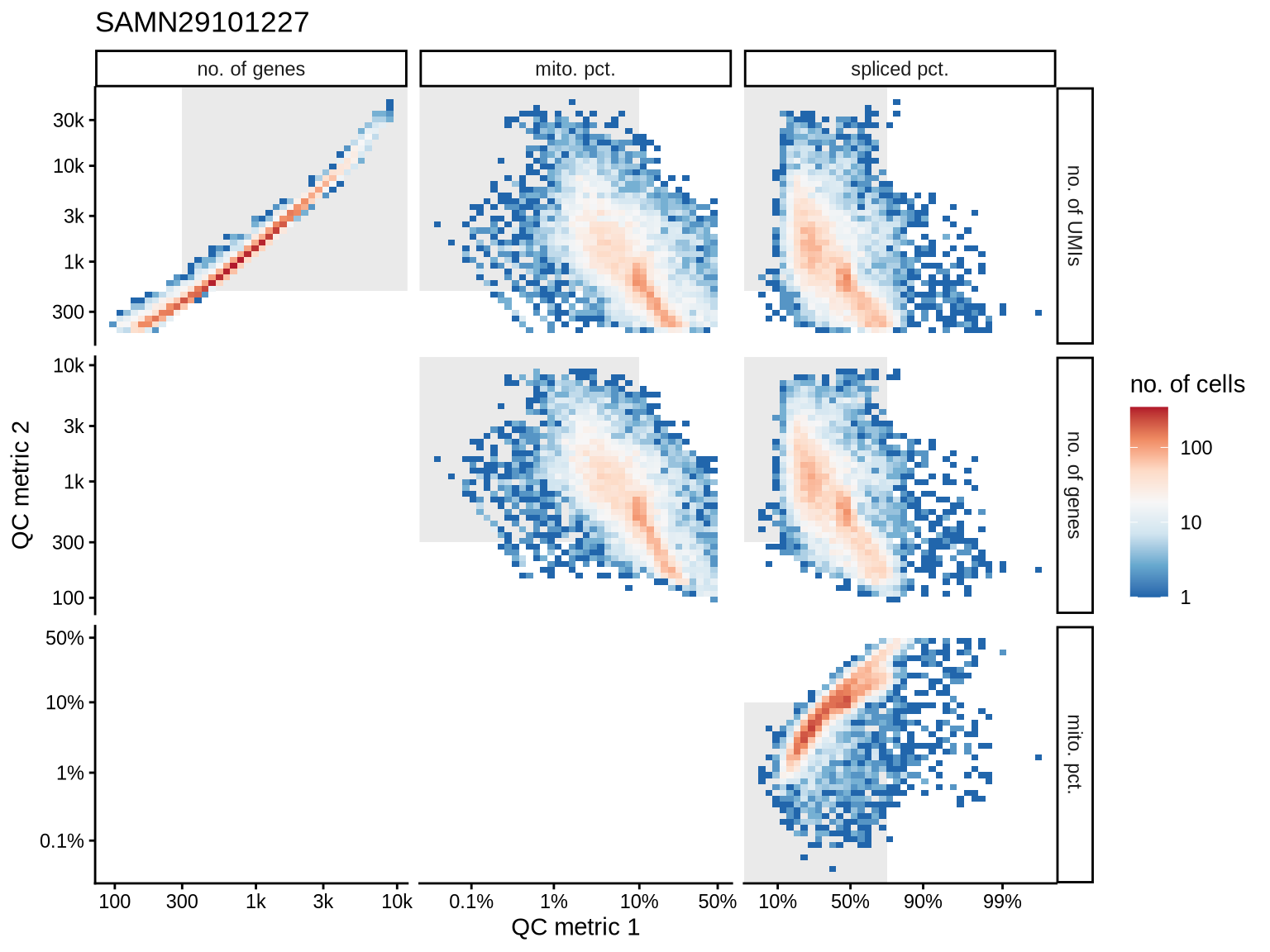
SAMN29101247
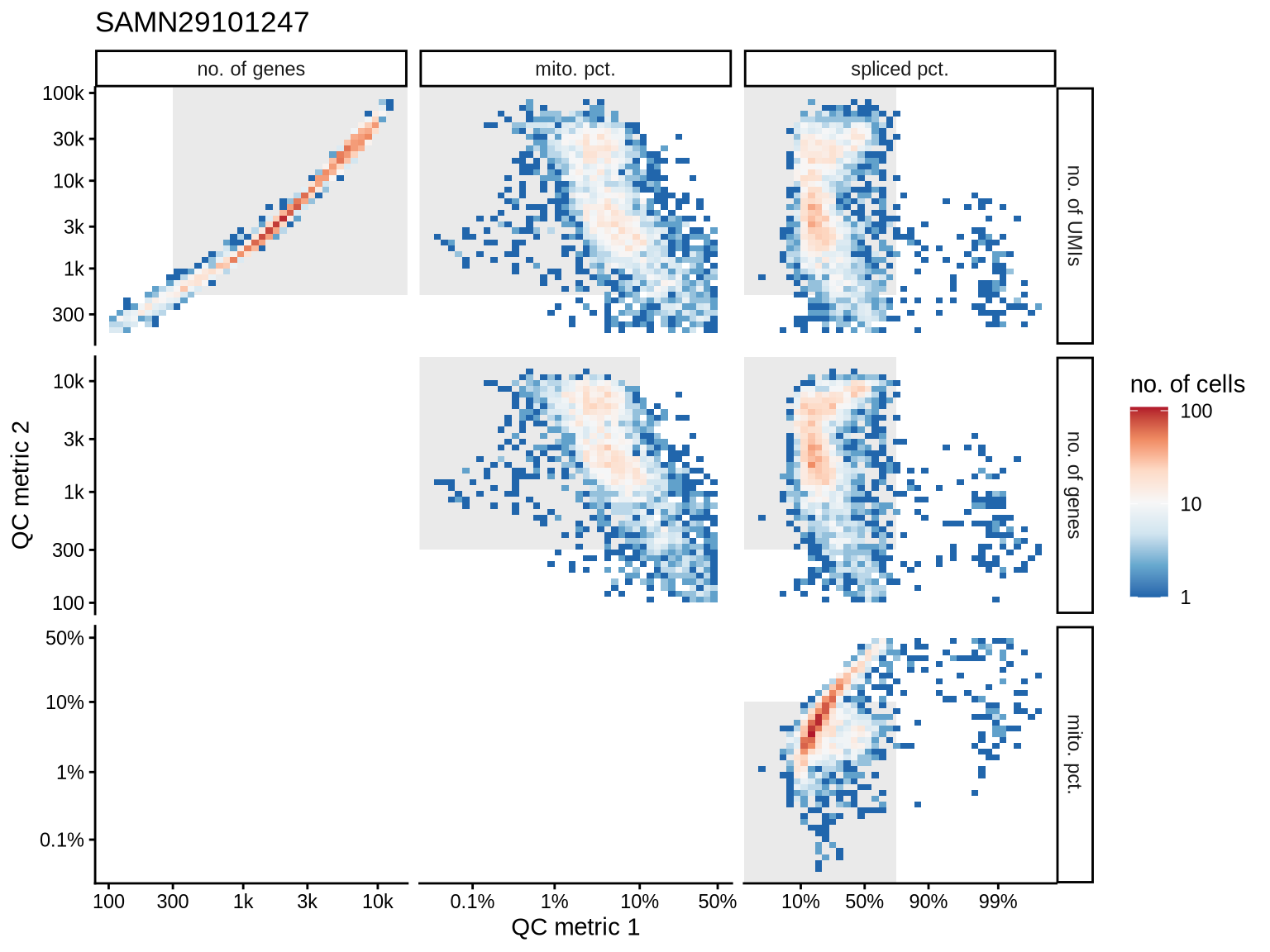
SAMN29101284
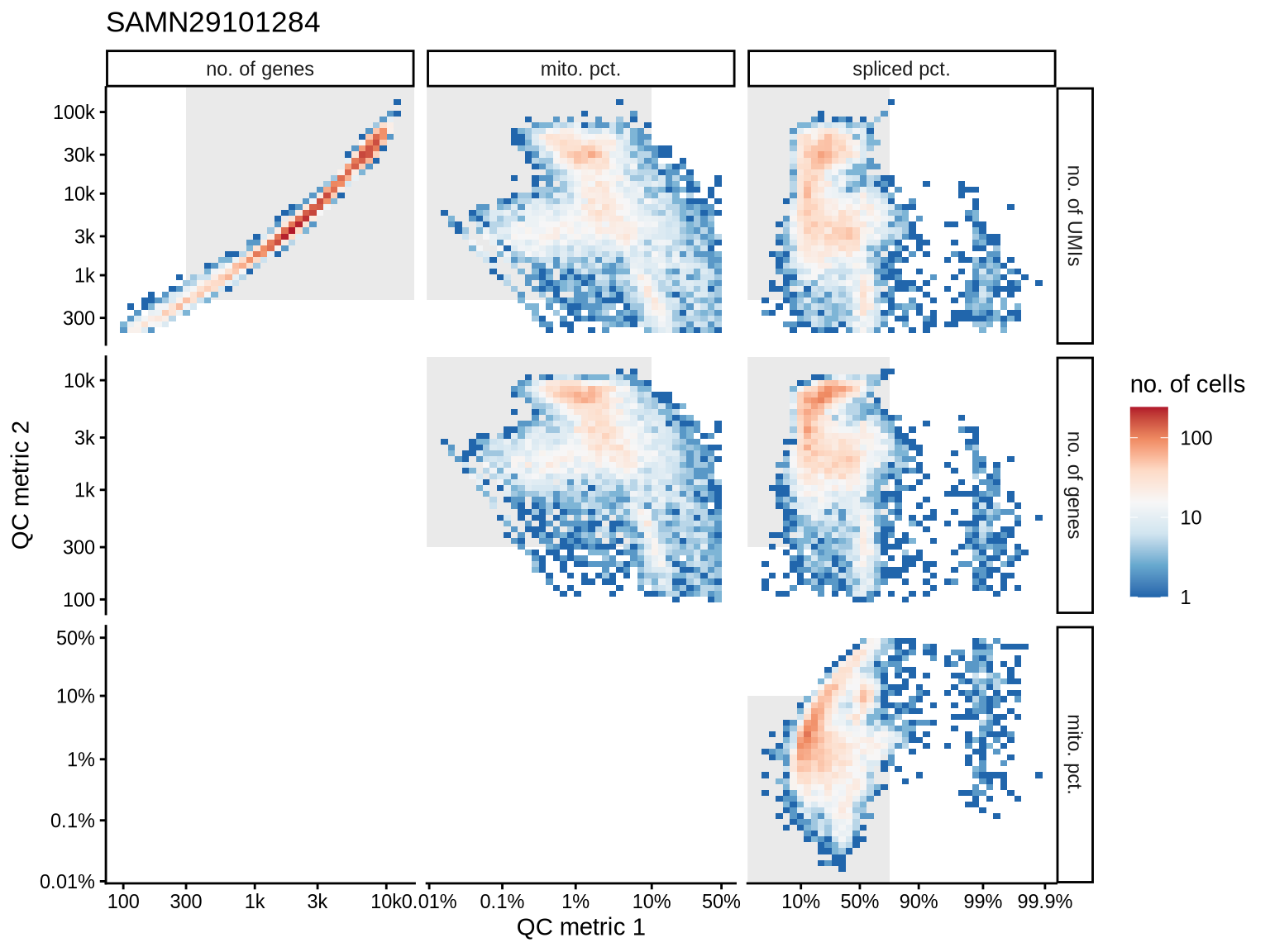
SAMN29101255
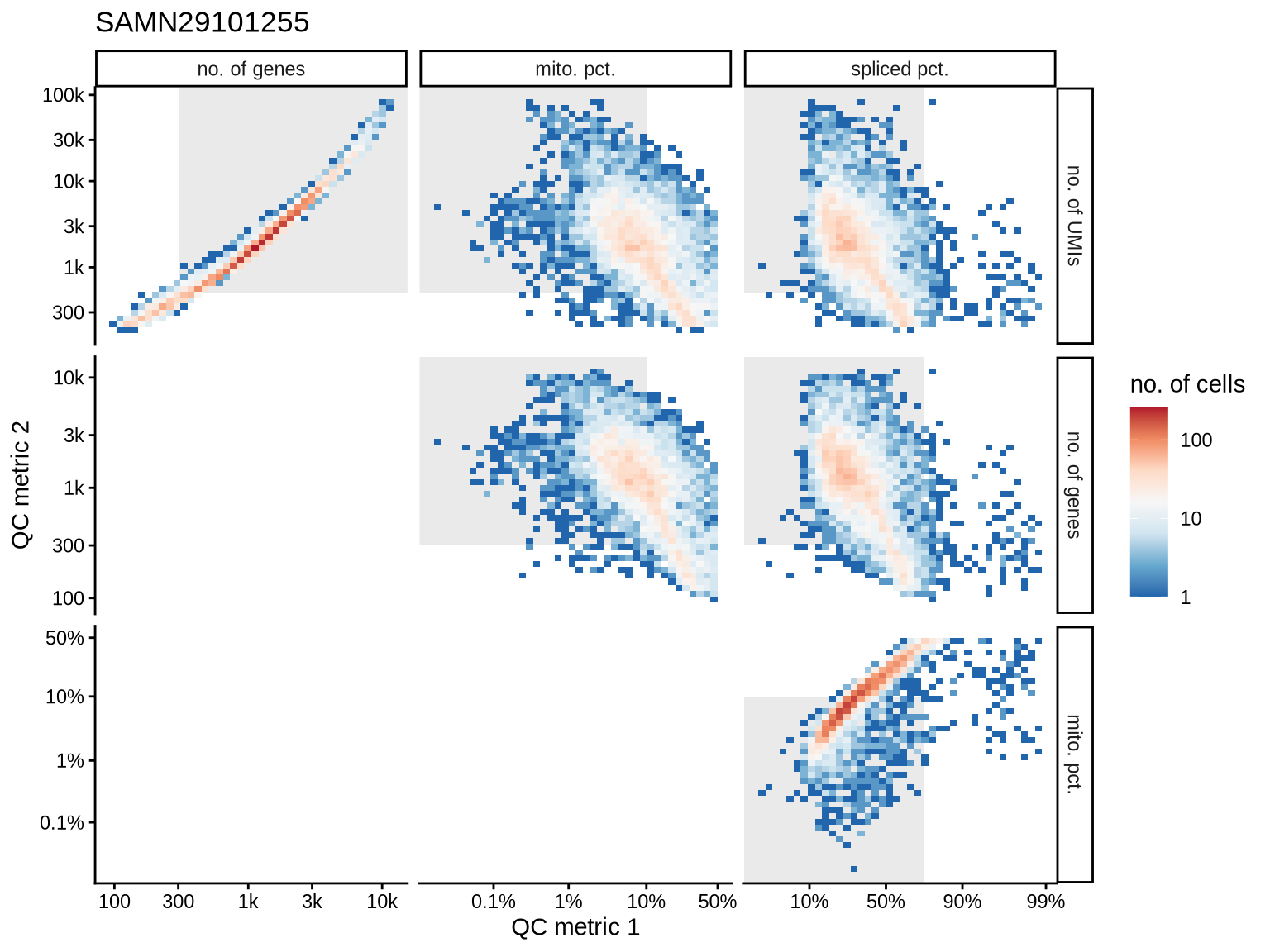
SAMN29101262
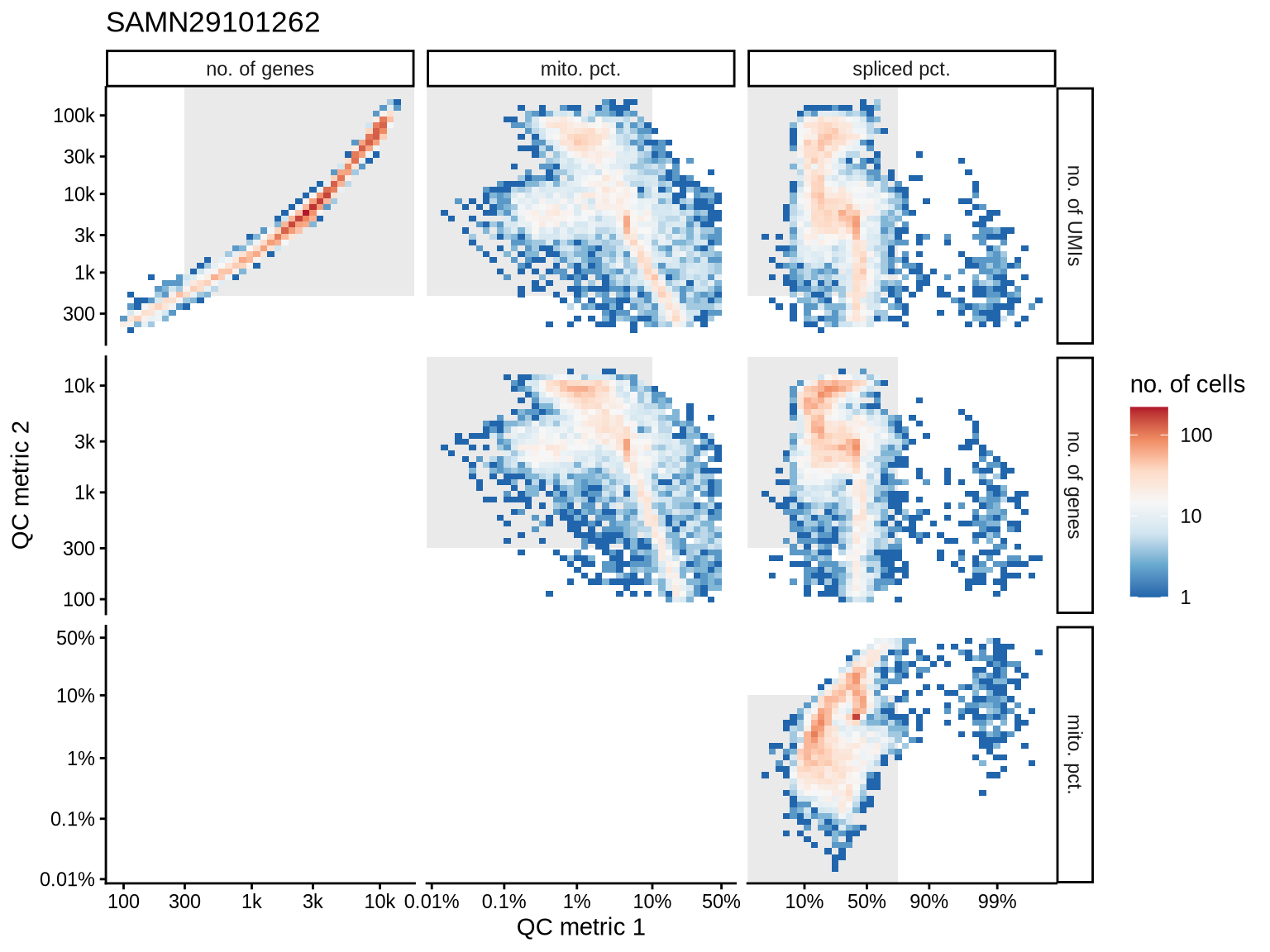
SAMN29101235
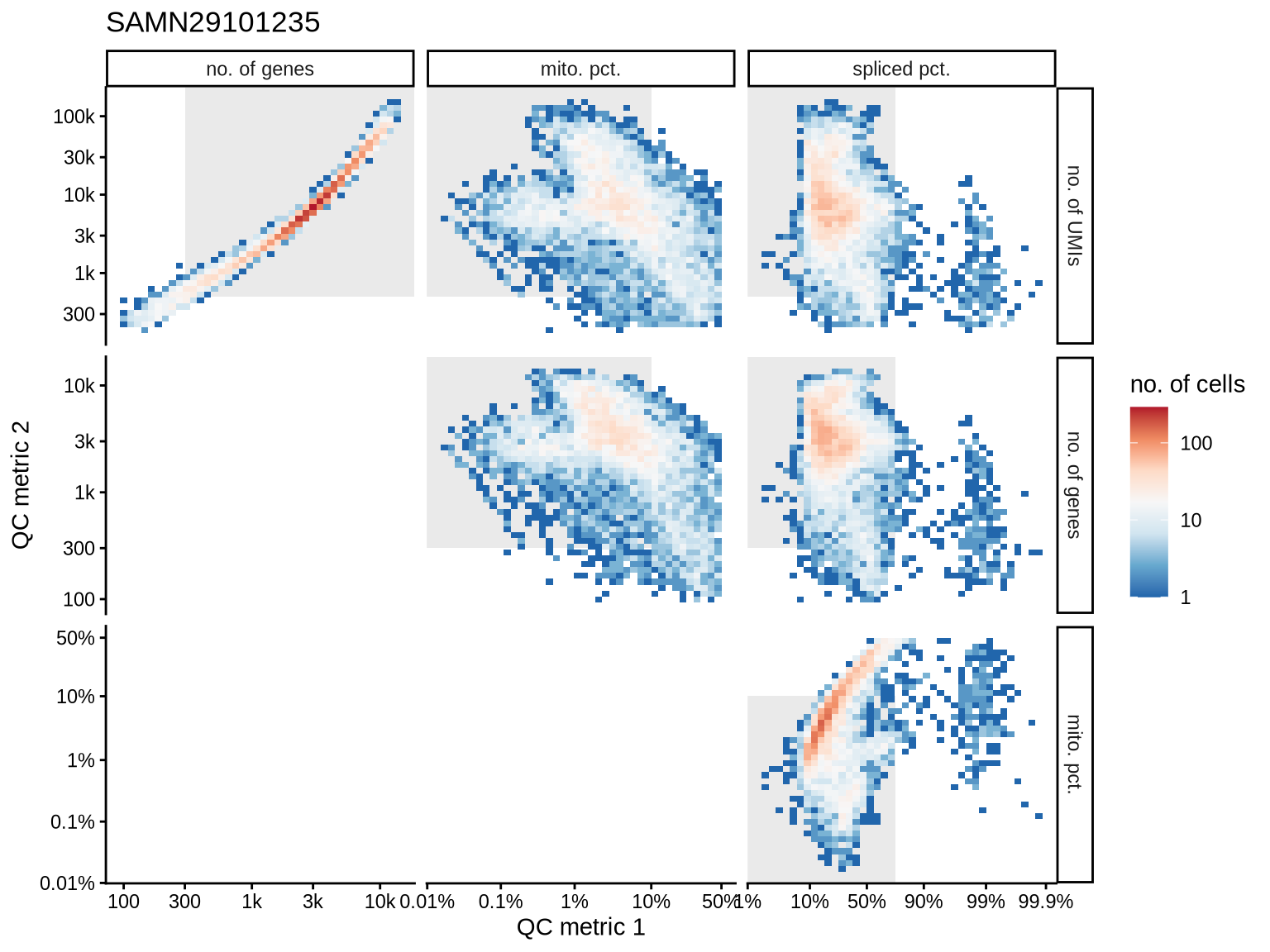
SAMN29101271
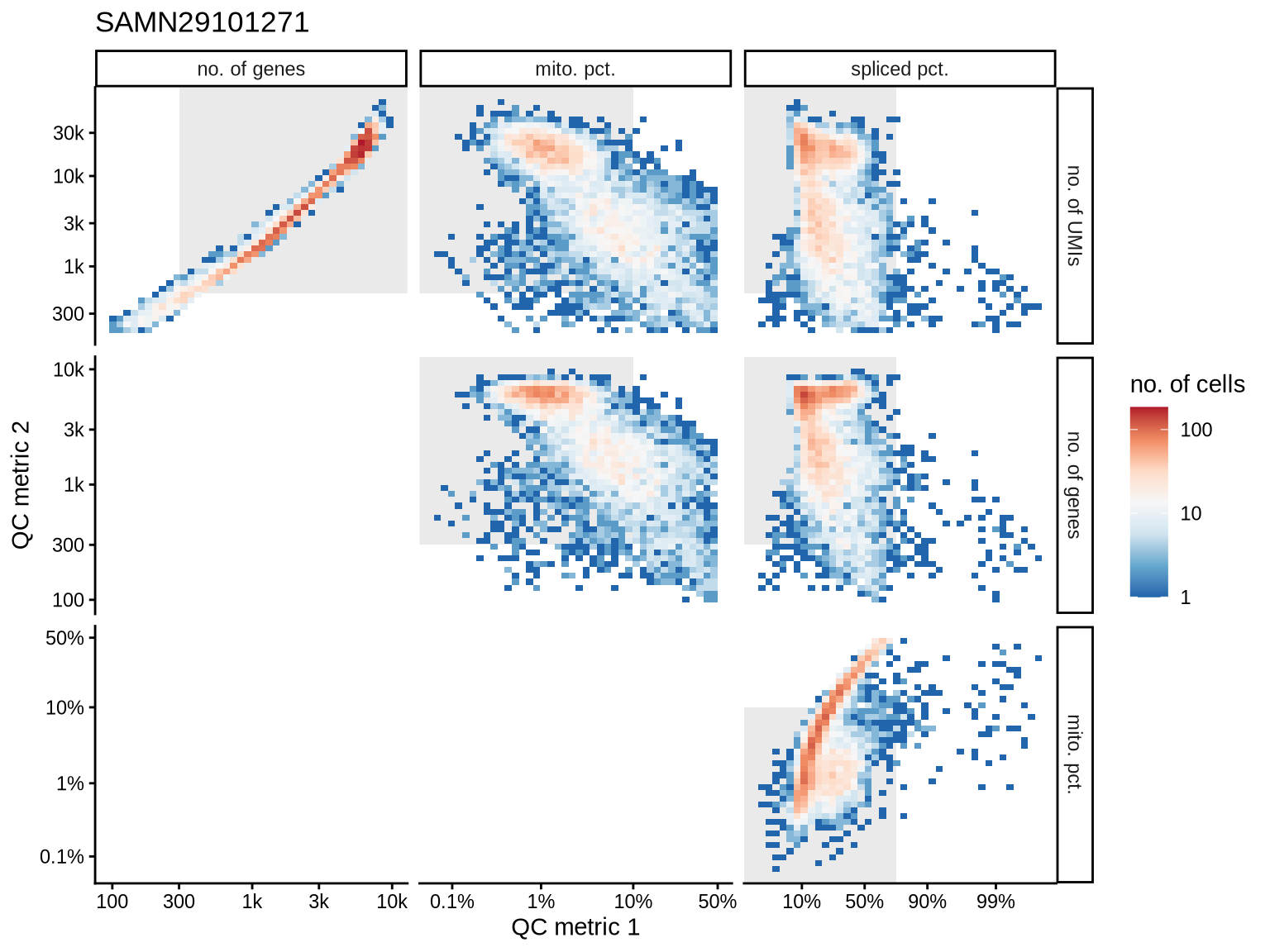
SAMN29101226
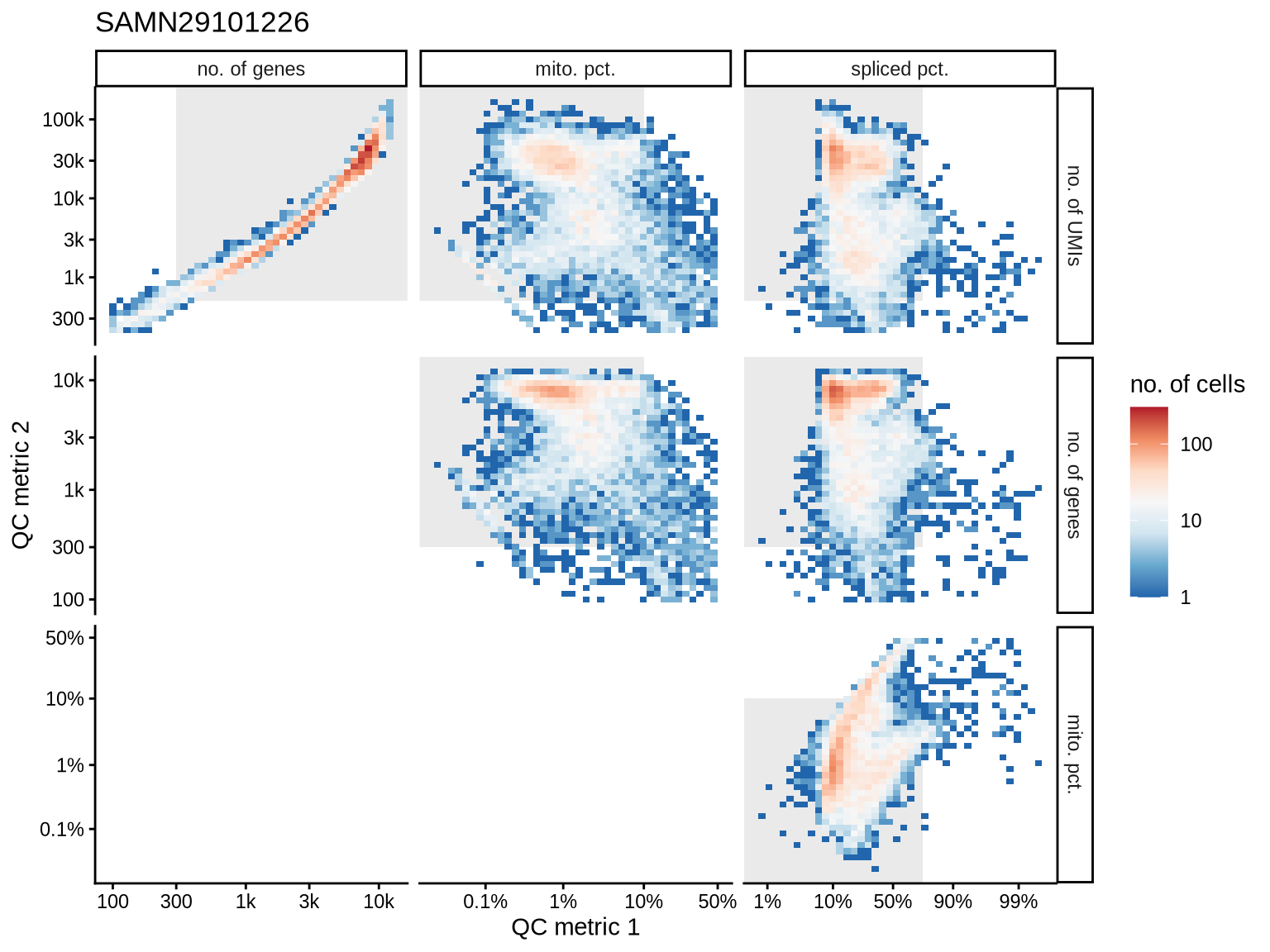
SAMN29101231
SAMN29101274
SAMN29101238
SAMN29101245
SAMN29101290
SAMN29101206
SAMN29101292
SAMN29101278
SAMN29101256
SAMN29101279
SAMN29101223
How many cells and samples retained?
Table summarizing the number of cells before and after qc filtering. Some samples may have been excluded from further analysis due to insufficient number of cells.
calc_qc_summary(qc_dt, kept_dt, cuts_dt, qc_lu, batch_var) %>% knitr::kable()| sample_id | excluded | N pre-QC | N post-QC | N excluded | pct. excluded | pct. excluded by no. of UMIs | pct. excluded by no. of genes | pct. excluded by mito. pct. | pct. excluded by spliced pct. |
|---|---|---|---|---|---|---|---|---|---|
| SAMN29101223 | FALSE | 1157 | 254 | 903 | 78.0 | 13.7 | 10.2 | 66.8 | 5.5 |
| SAMN29101279 | FALSE | 3314 | 1223 | 2091 | 63.1 | 24.5 | 19.3 | 44.3 | 7.2 |
| SAMN29101255 | FALSE | 6405 | 3359 | 3046 | 47.6 | 14.9 | 12.6 | 45.6 | 3.5 |
| SAMN29101227 | FALSE | 10497 | 5613 | 4884 | 46.5 | 26.7 | 19.3 | 39.7 | 5.4 |
| SAMN29101292 | FALSE | 5710 | 3756 | 1954 | 34.2 | 5.9 | 6.4 | 27.9 | 10.2 |
| SAMN29101222 | FALSE | 6889 | 4885 | 2004 | 29.1 | 7.4 | 7.7 | 26.2 | 4.0 |
| SAMN29101235 | FALSE | 7075 | 5026 | 2049 | 29.0 | 5.7 | 6.7 | 25.4 | 4.6 |
| SAMN29101271 | FALSE | 6899 | 4953 | 1946 | 28.2 | 7.3 | 6.1 | 25.1 | 1.7 |
| SAMN29101247 | FALSE | 3337 | 2445 | 892 | 26.7 | 7.0 | 7.3 | 24.4 | 3.4 |
| SAMN29101256 | FALSE | 1959 | 1473 | 486 | 24.8 | 3.1 | 4.0 | 3.9 | 22.7 |
| SAMN29101262 | FALSE | 7431 | 5692 | 1739 | 23.4 | 7.1 | 8.0 | 20.5 | 3.6 |
| SAMN29101278 | FALSE | 2834 | 2187 | 647 | 22.8 | 8.3 | 6.2 | 1.5 | 16.4 |
| SAMN29101284 | FALSE | 8422 | 6590 | 1832 | 21.8 | 6.9 | 6.9 | 17.0 | 4.8 |
| SAMN29101249 | FALSE | 8779 | 7135 | 1644 | 18.7 | 5.8 | 6.2 | 15.0 | 4.4 |
| SAMN29101206 | FALSE | 8748 | 7208 | 1540 | 17.6 | 7.1 | 8.1 | 13.6 | 3.1 |
| SAMN29101290 | FALSE | 3842 | 3172 | 670 | 17.4 | 7.2 | 7.4 | 13.7 | 3.3 |
| SAMN29101226 | FALSE | 7781 | 6550 | 1231 | 15.8 | 4.1 | 5.2 | 12.3 | 2.6 |
| SAMN29101274 | FALSE | 7510 | 6439 | 1071 | 14.3 | 4.0 | 4.6 | 10.9 | 2.6 |
| SAMN29101231 | FALSE | 7562 | 6513 | 1049 | 13.9 | 3.2 | 4.5 | 10.9 | 2.5 |
| SAMN29101238 | FALSE | 6588 | 5682 | 906 | 13.8 | 3.5 | 5.3 | 10.8 | 2.8 |
| SAMN29101245 | FALSE | 5702 | 4956 | 746 | 13.1 | 3.2 | 4.2 | 9.0 | 3.8 |
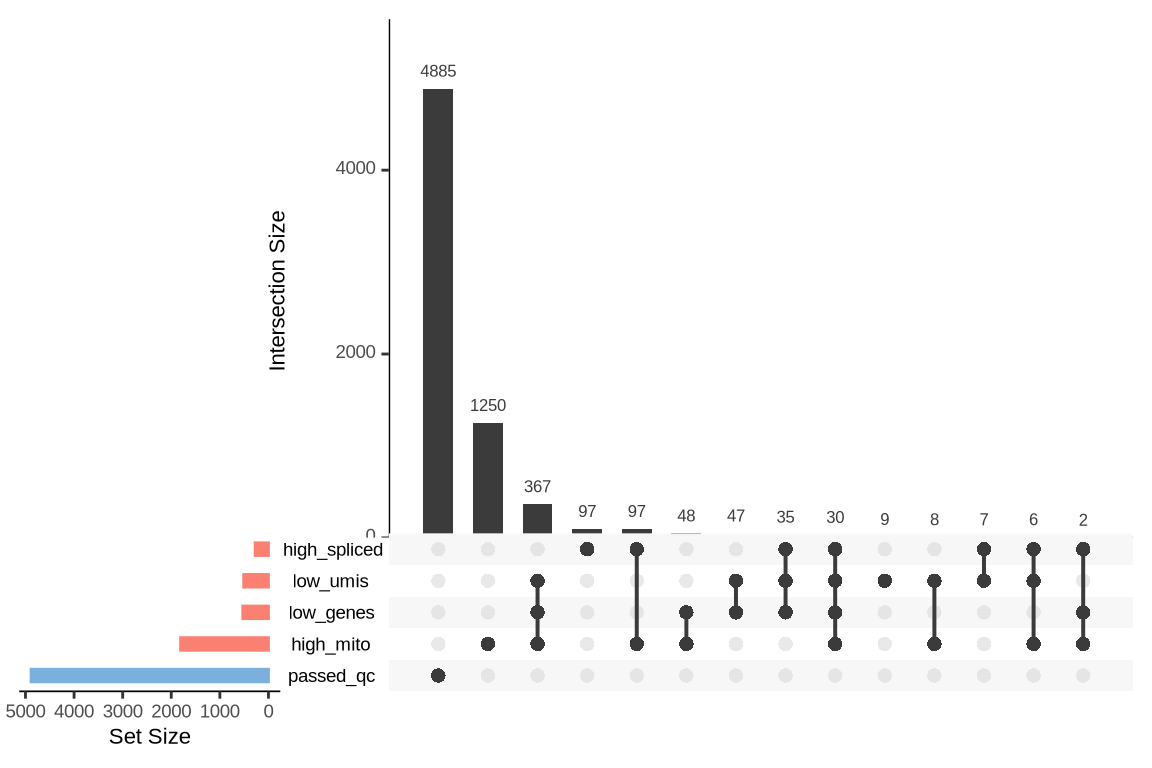
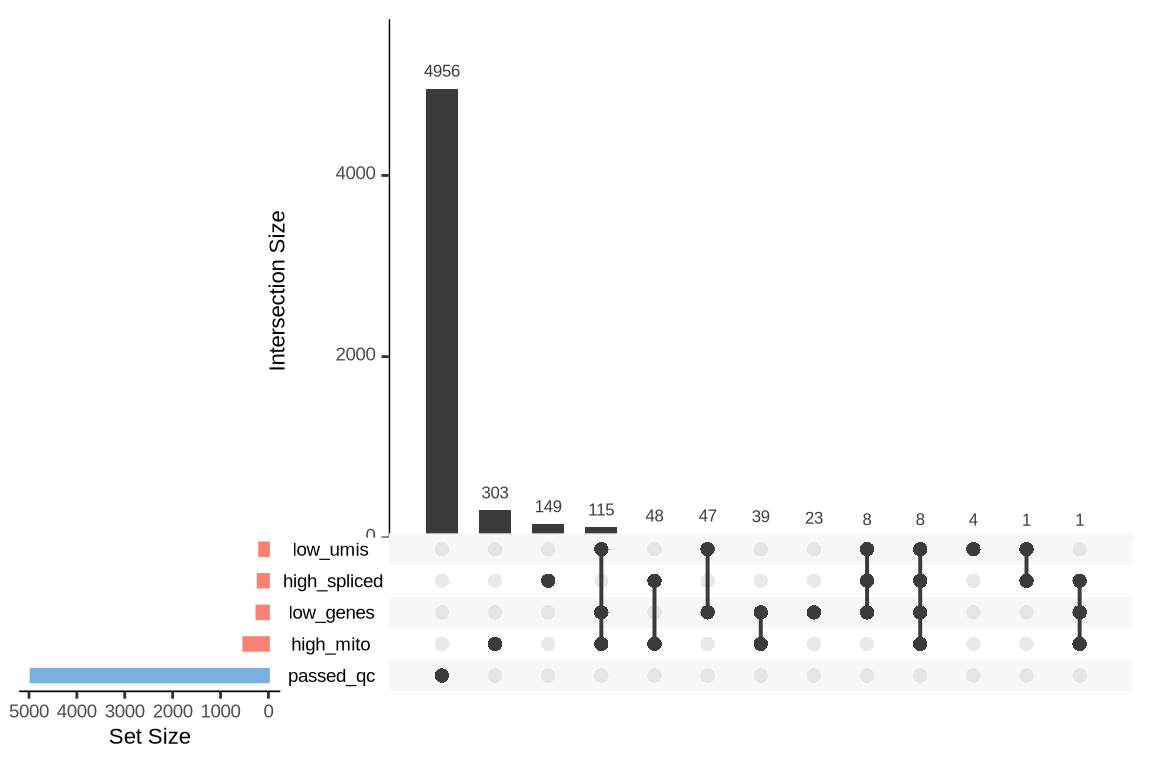
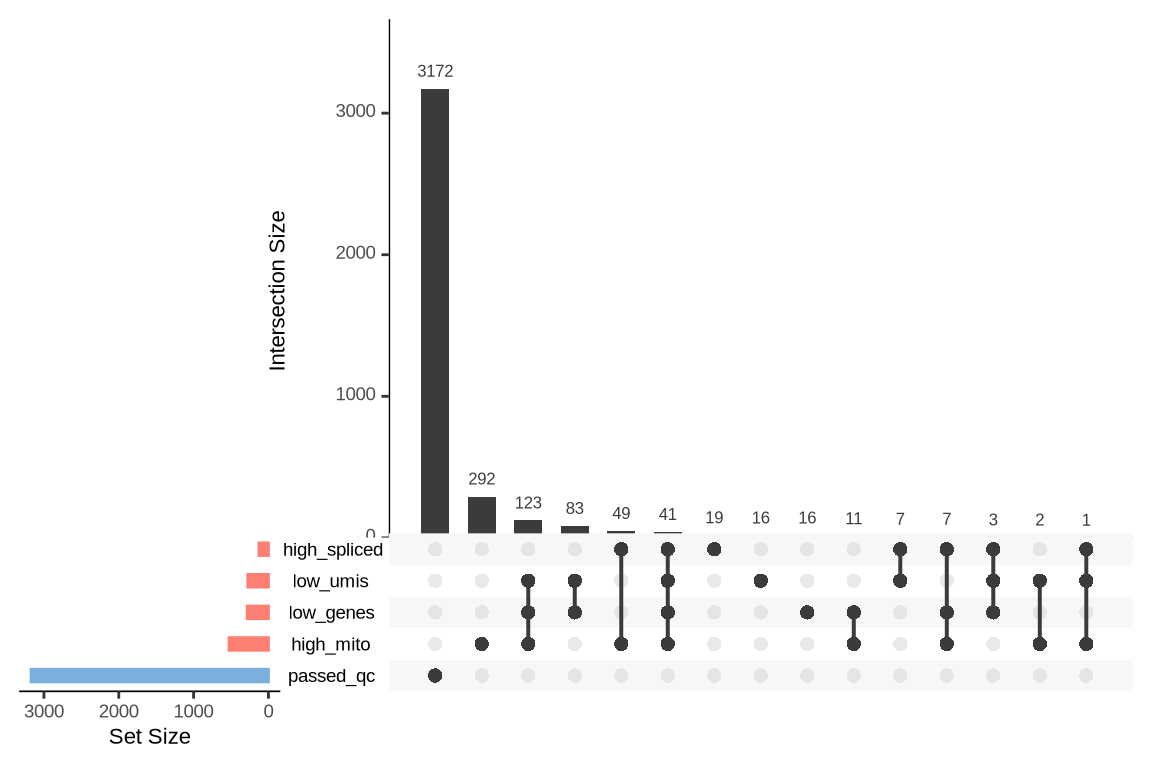
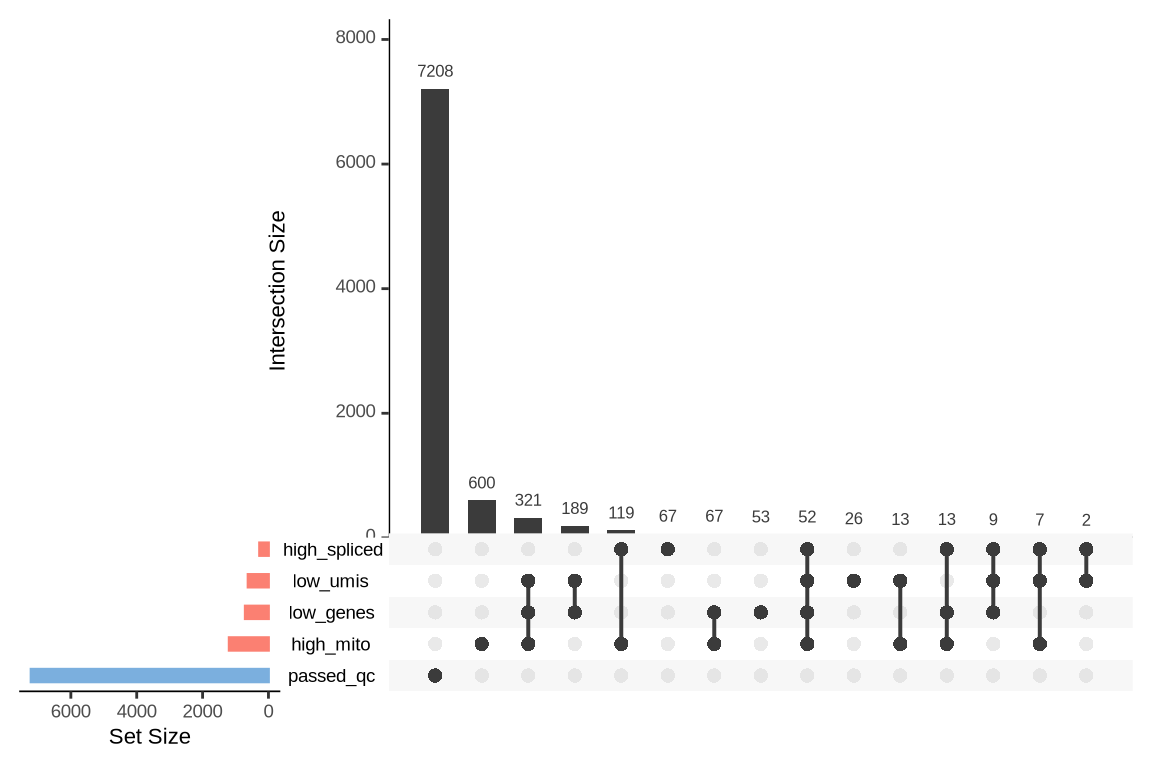
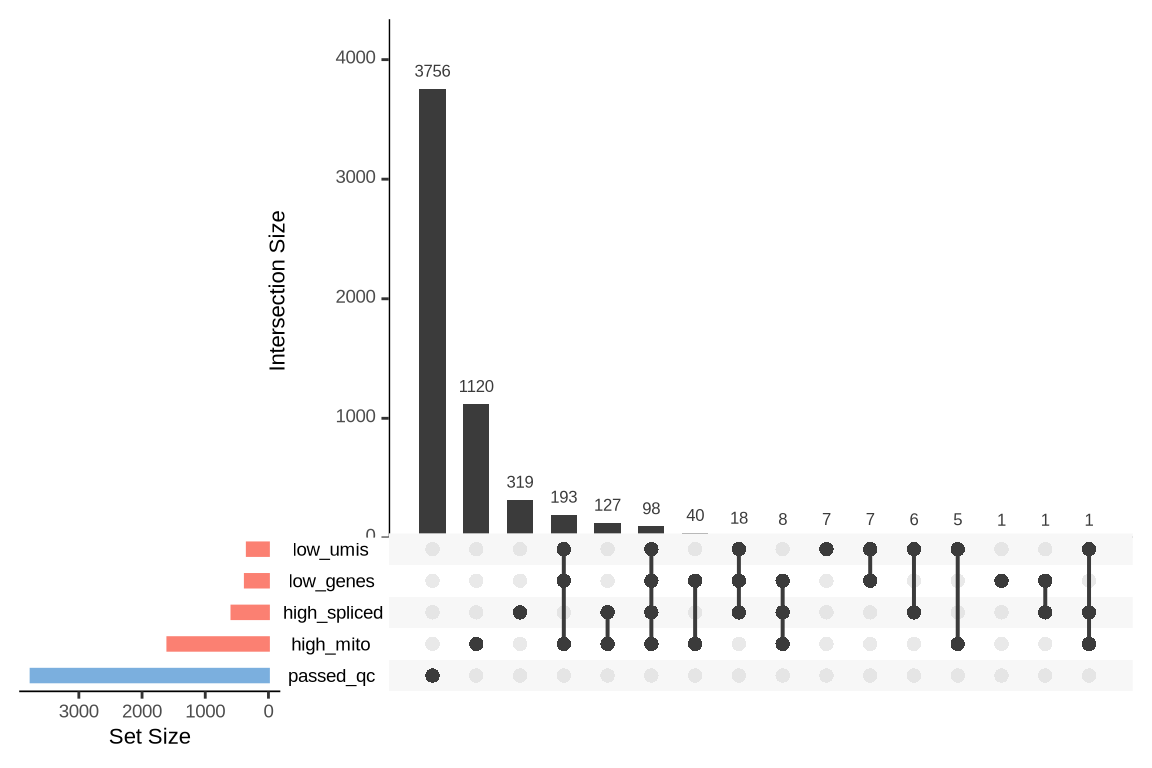
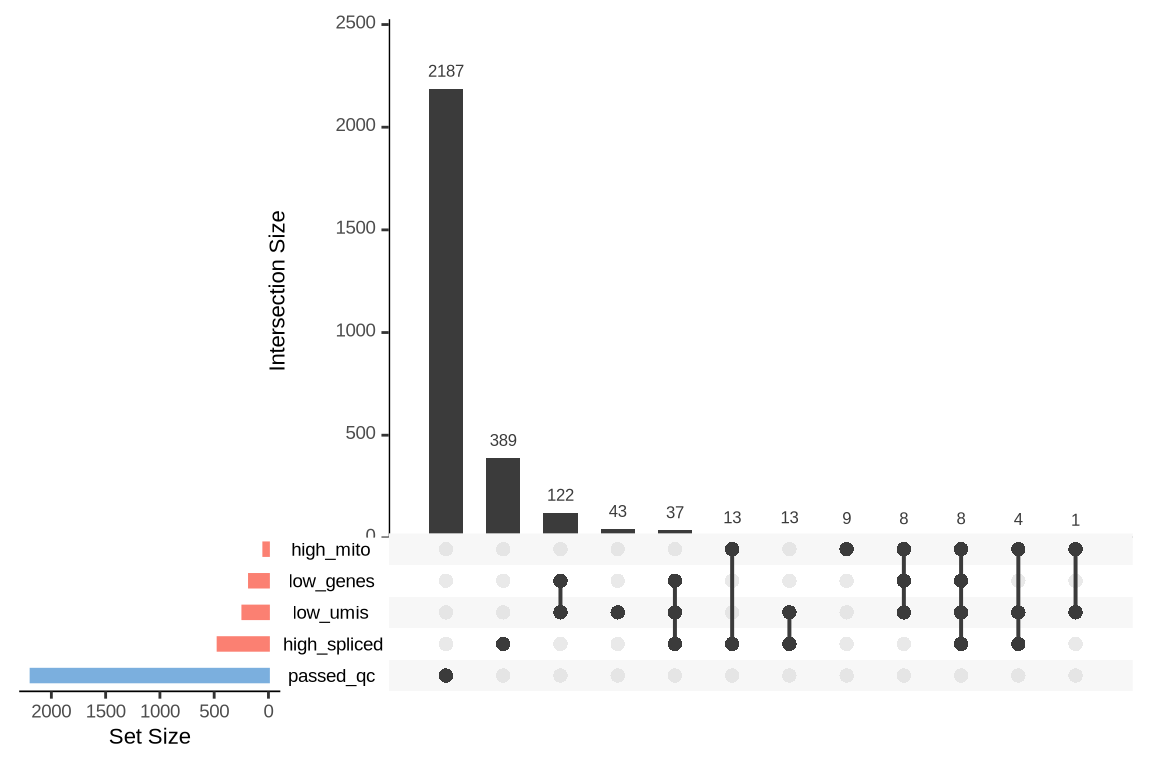
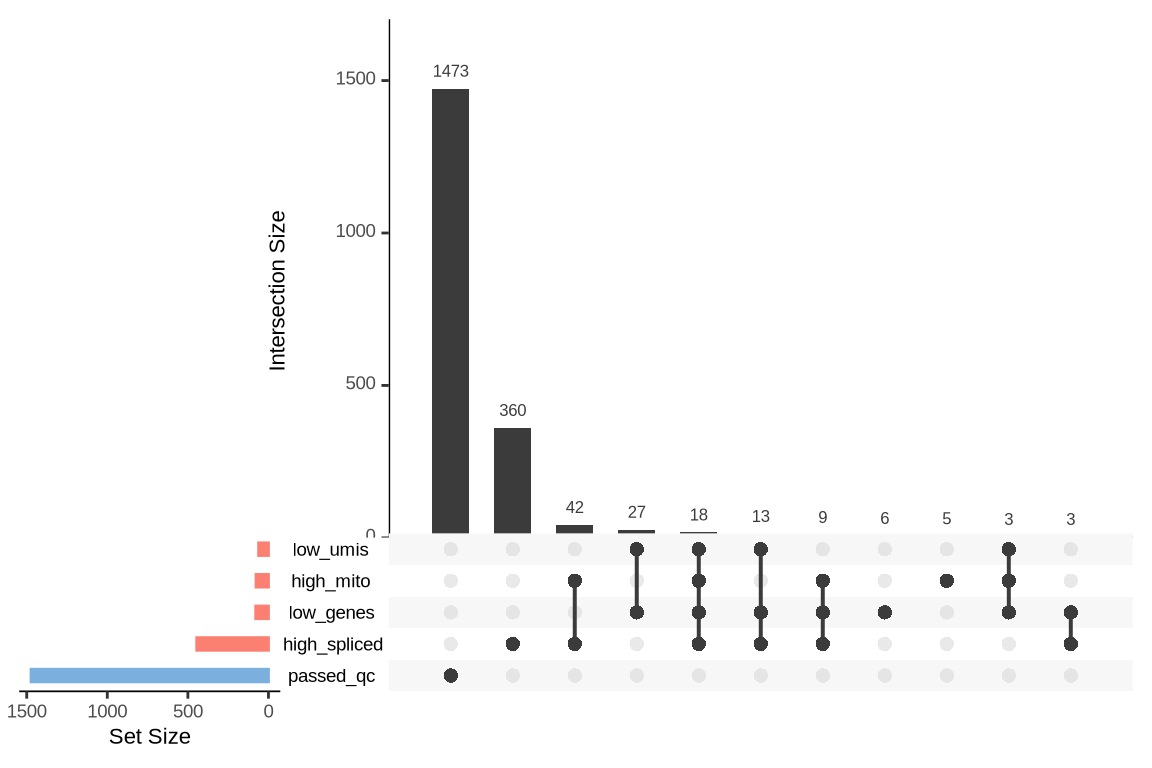
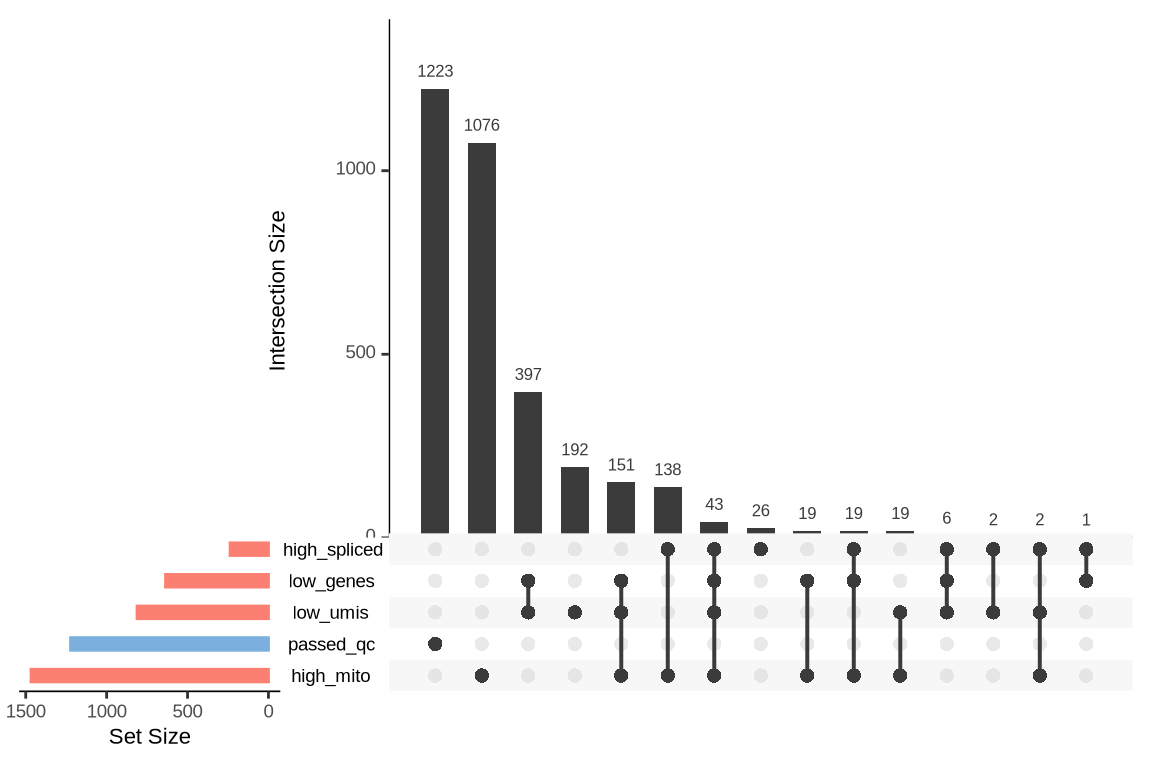
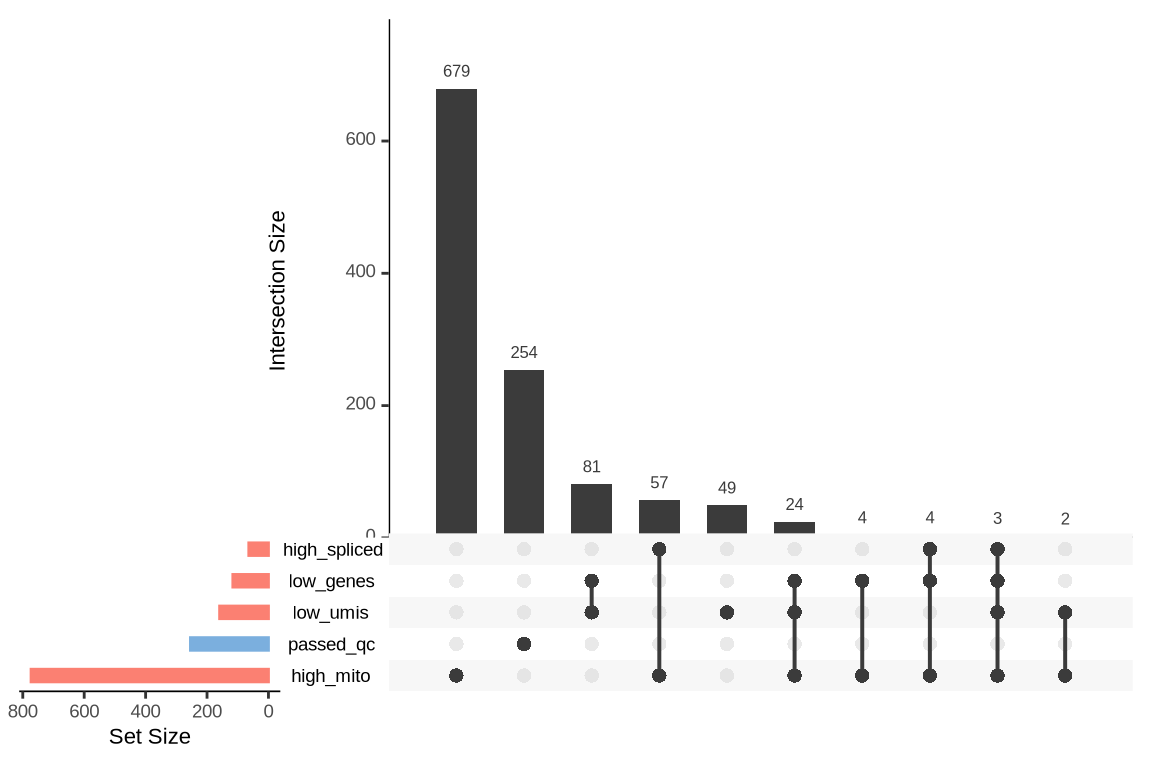
What were the reasons for excluding barcodes?
An upset plot is generated for each sample to visualize the reasons for barcode exclusion. The plot shows the overlap between different exclusion criteria, with vertical bars representing the size of each intersection. Plots are omitted for samples where no barcodes were excluded.
for (bb in b_lvls) {
qc_tmp = qc_dt[ batch_var == bb ]
if (nrow(qc_tmp) == 1 | sum(qc_tmp$keep) == nrow(qc_tmp))
next
cat('### ', bb, '\n')
suppressMessages( print(plot_upset_of_exclusions(qc_tmp, qc_names, qc_lu, cuts_dt)) )
cat('\n\n')
}SAMN29101222
SAMN29101249
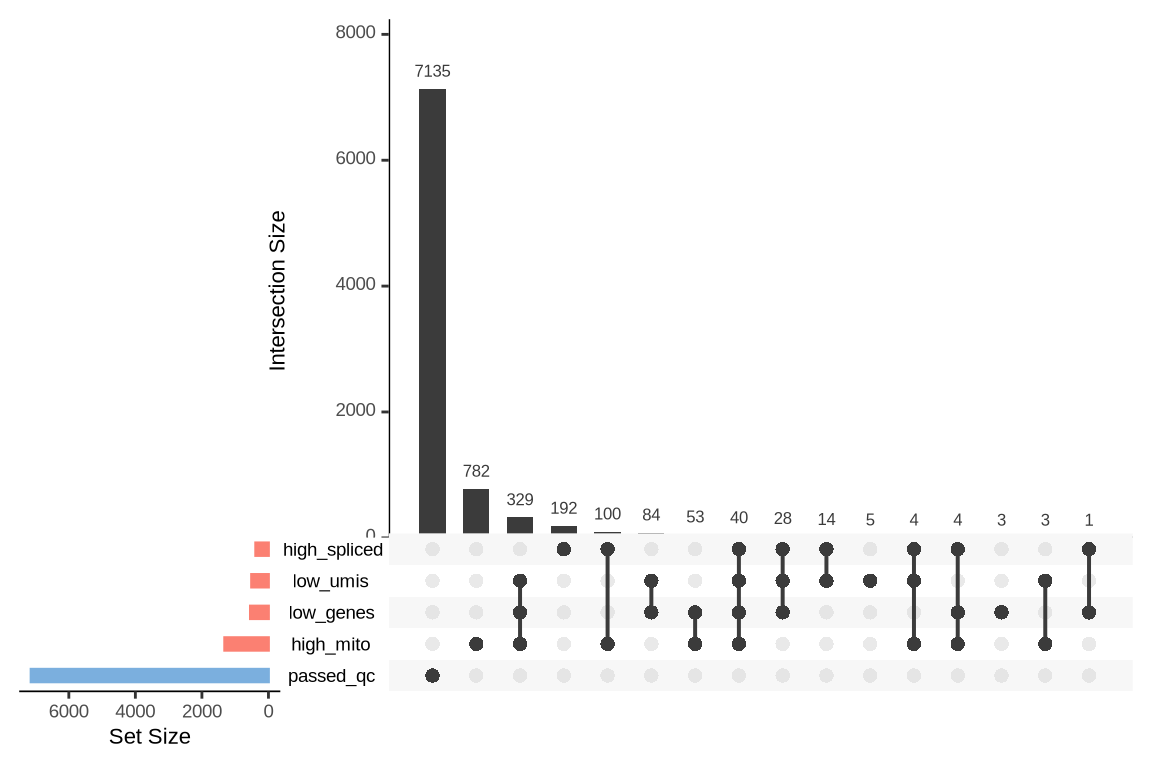
SAMN29101227
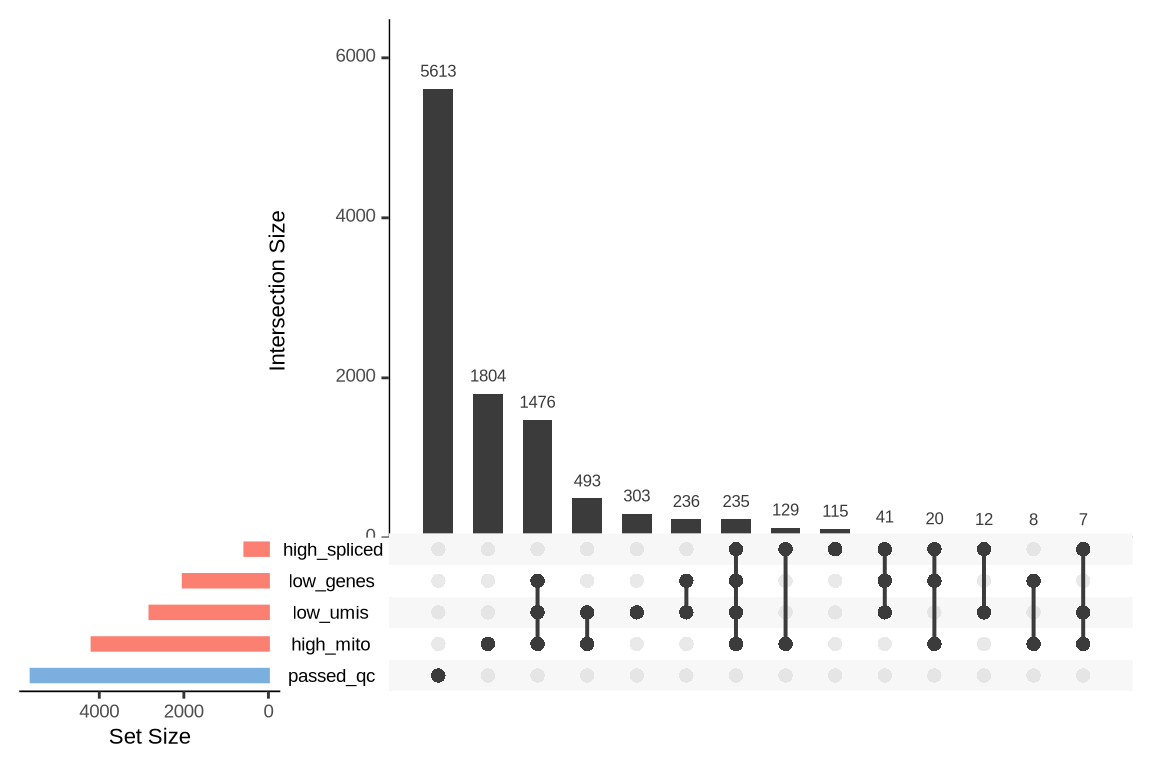
SAMN29101247
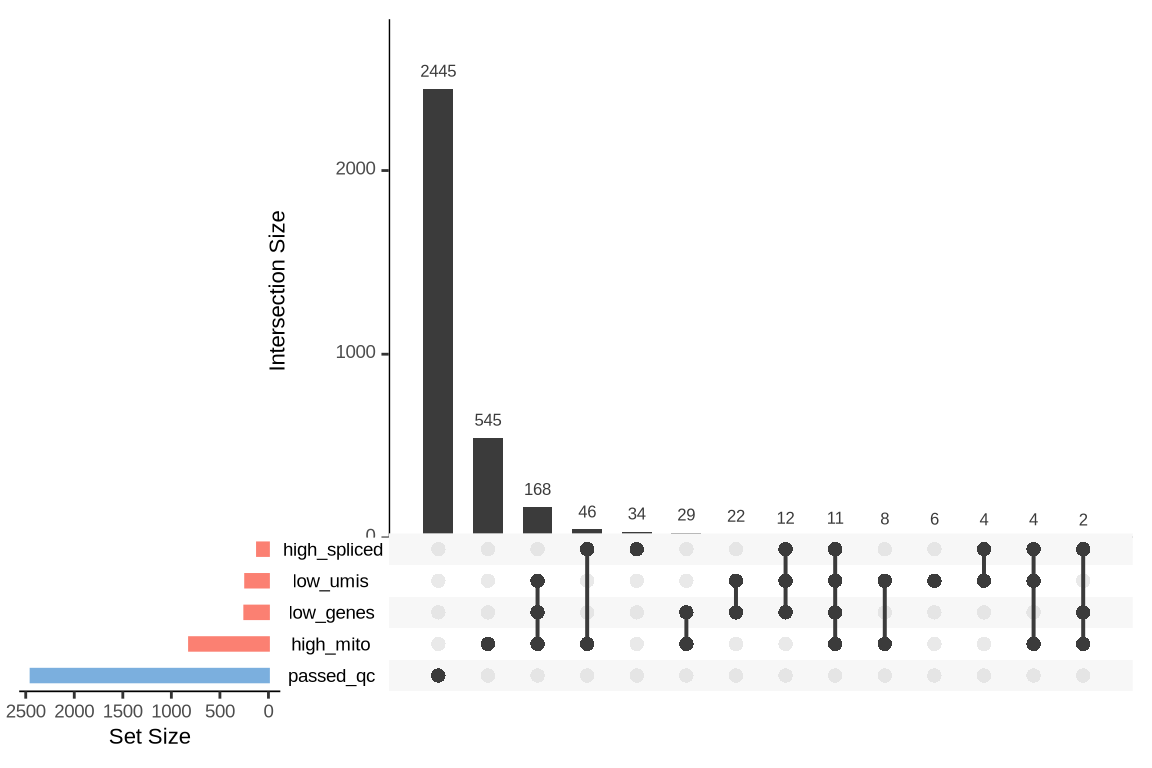
SAMN29101284
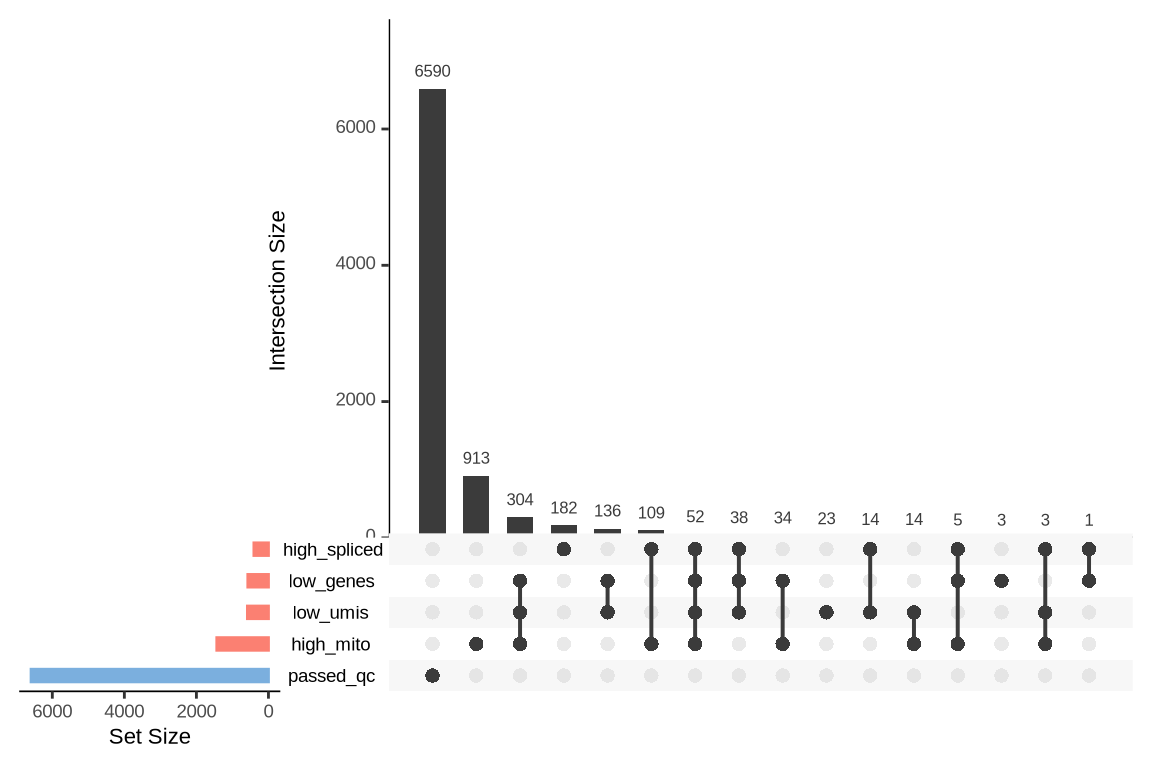
SAMN29101255
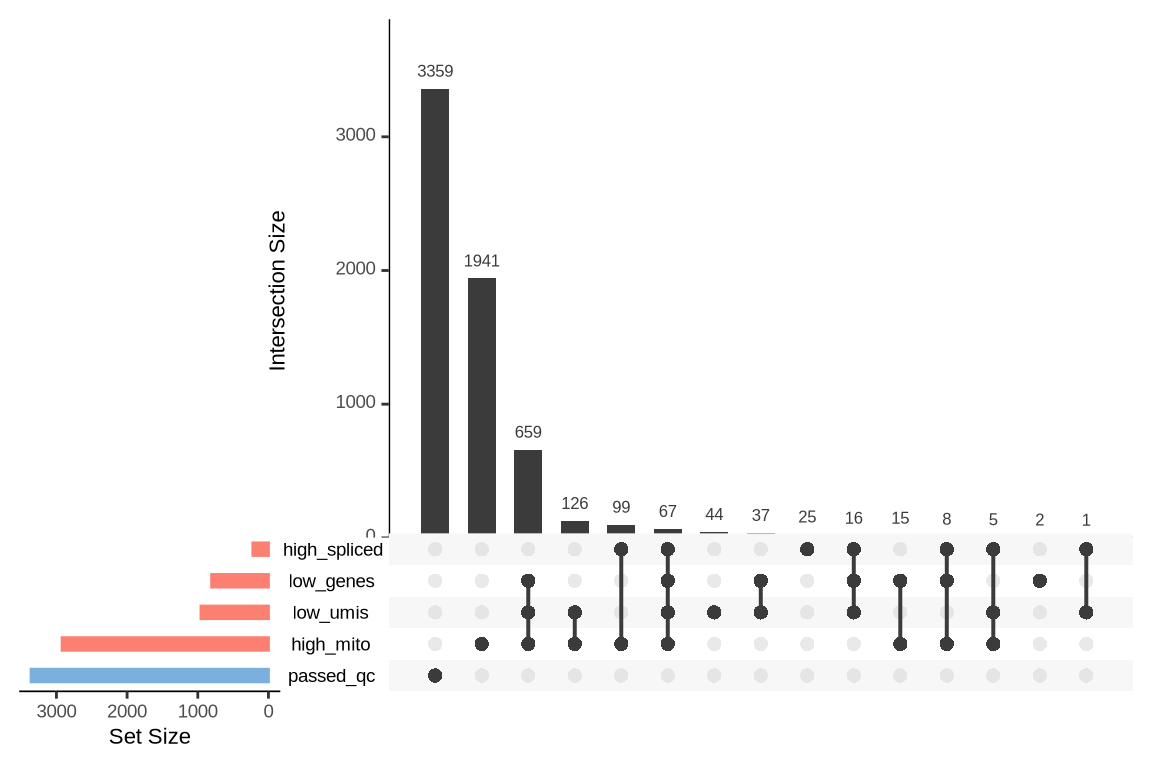
SAMN29101262
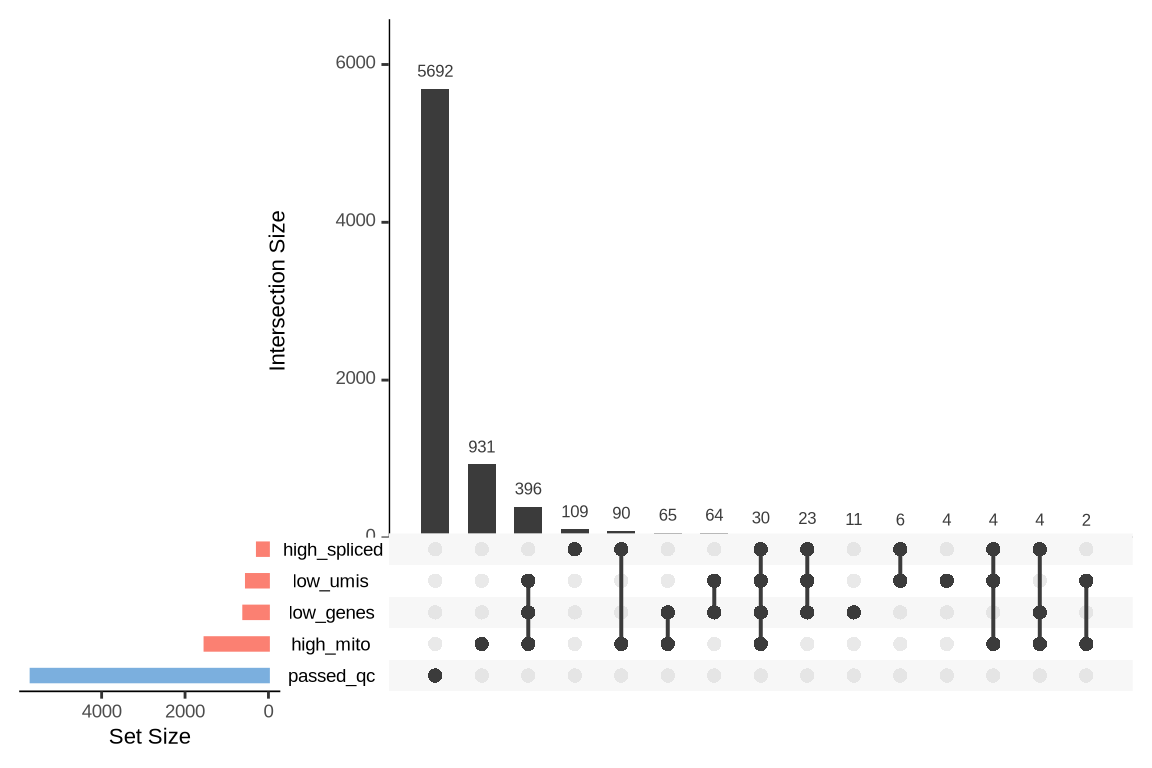
SAMN29101235
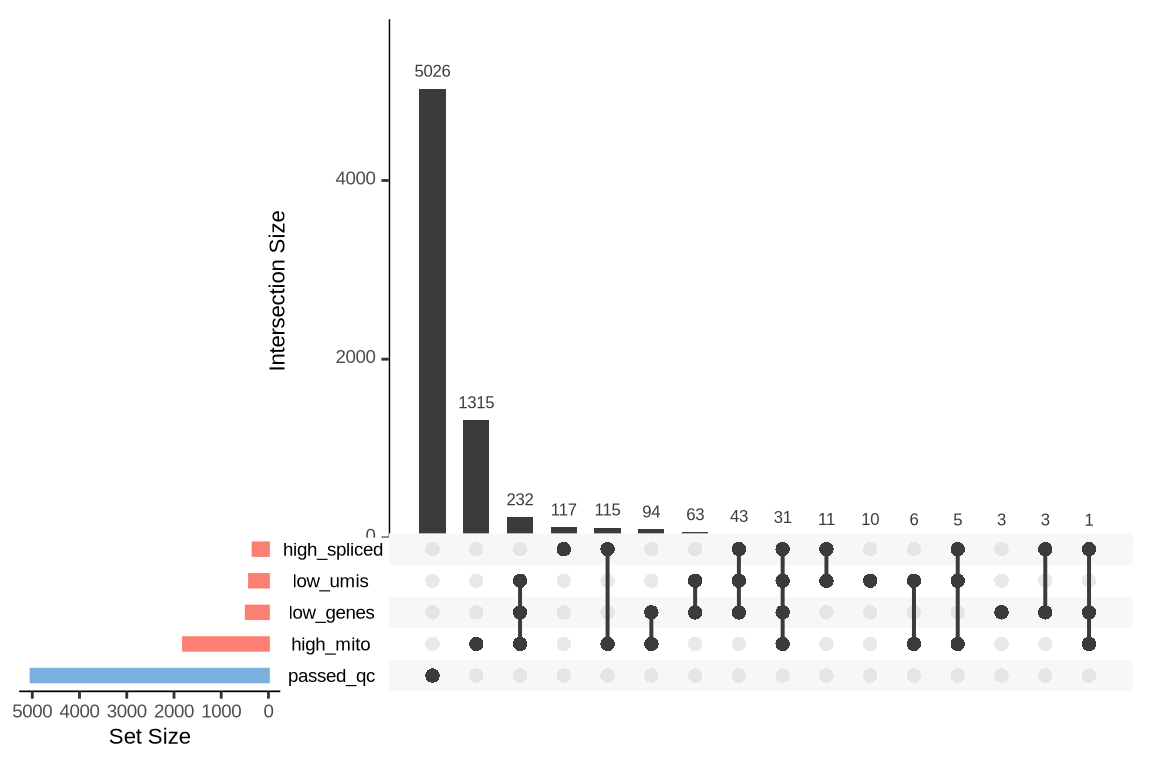
SAMN29101271
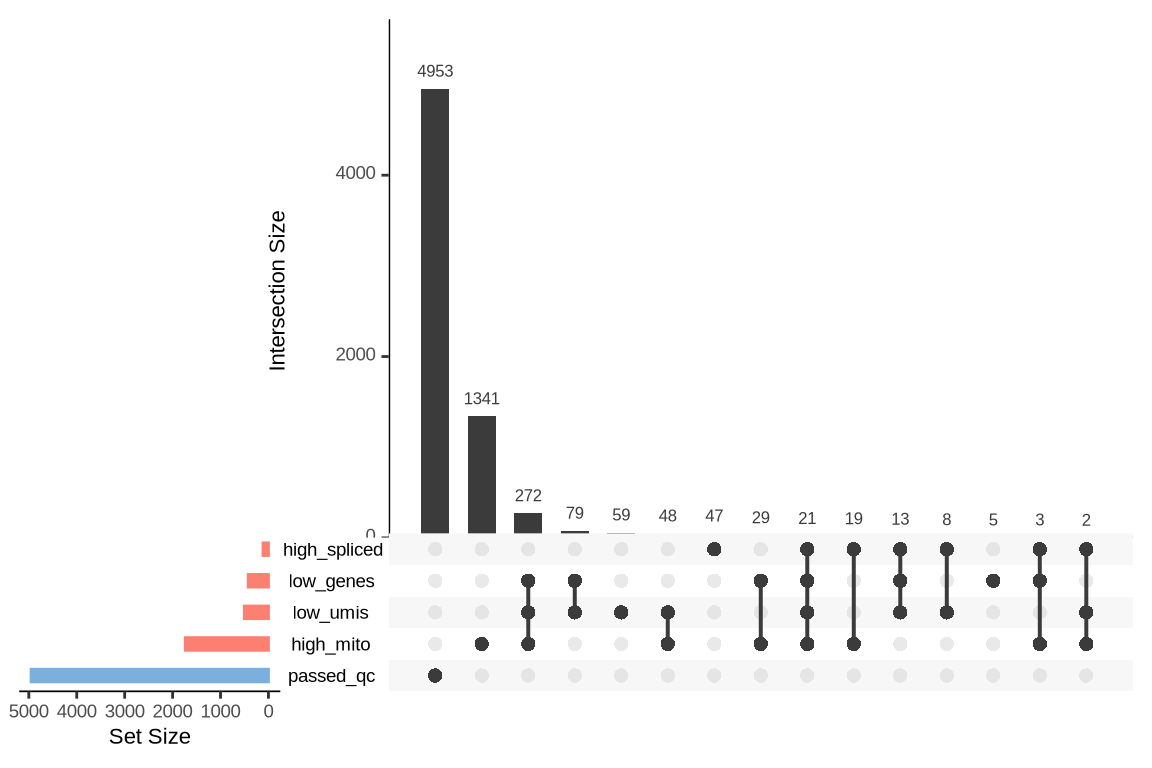
SAMN29101226
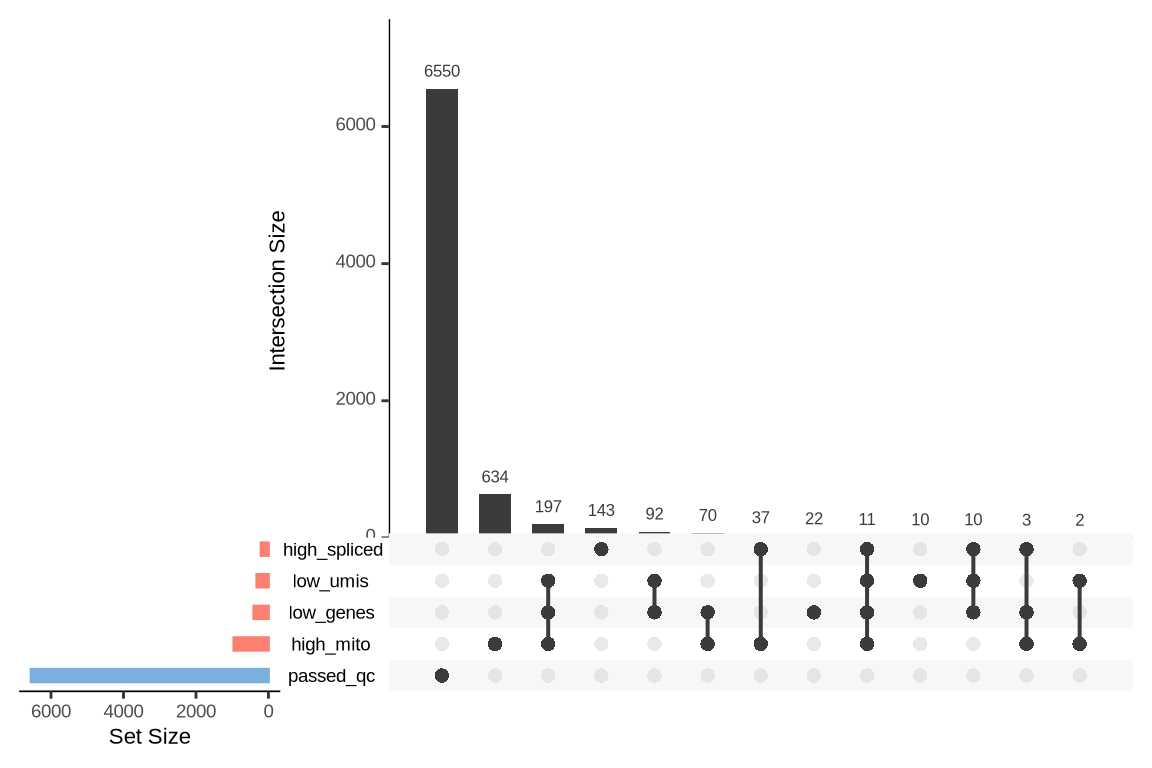
SAMN29101231
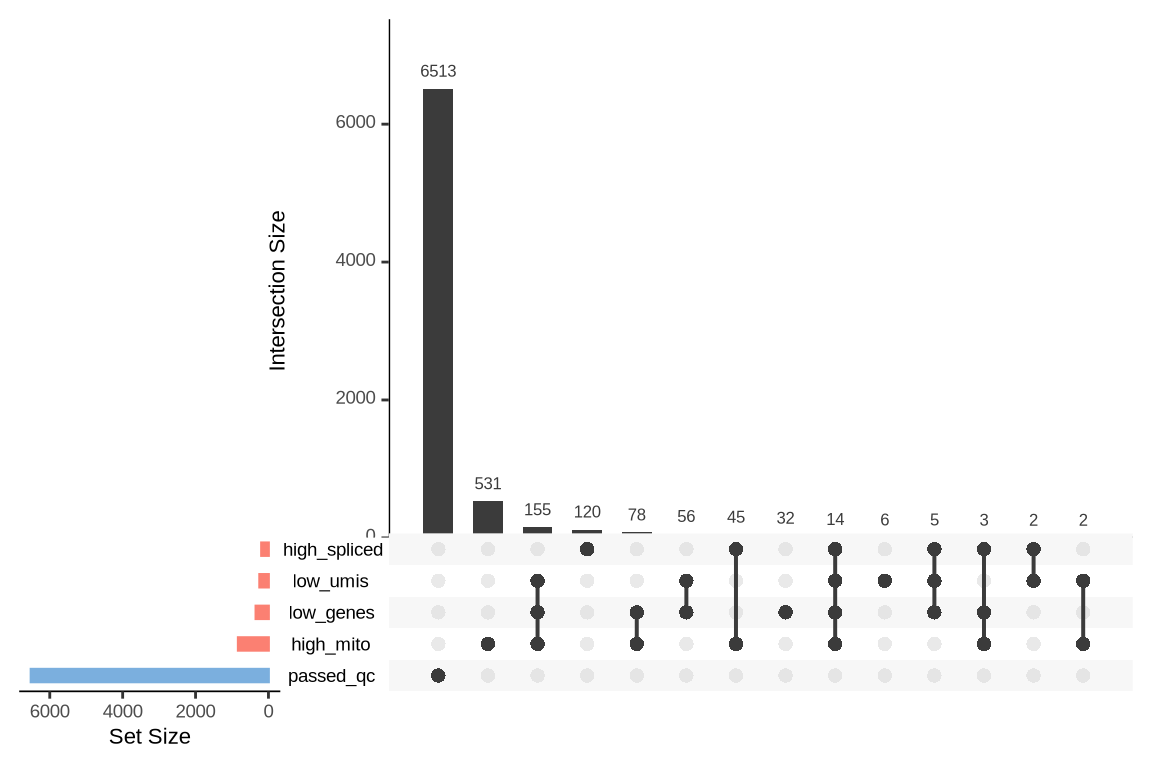
SAMN29101274
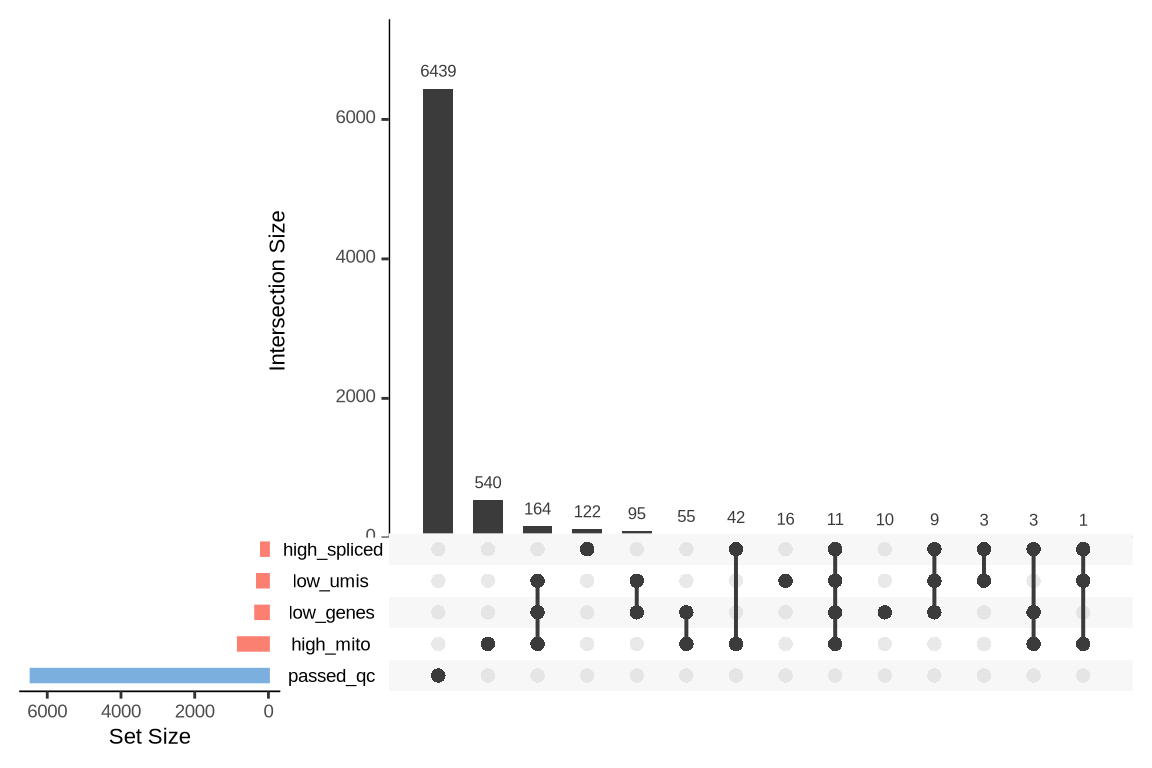
SAMN29101238
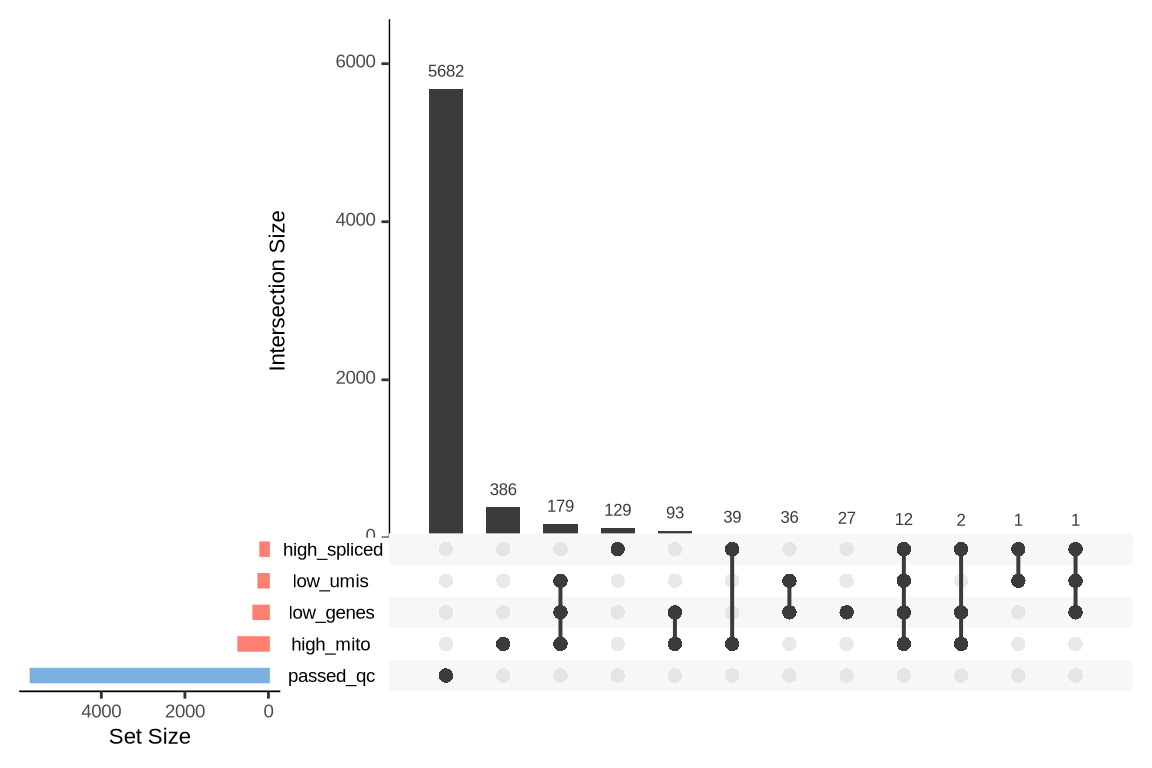
SAMN29101245
SAMN29101290
SAMN29101206
SAMN29101292
SAMN29101278
SAMN29101256
SAMN29101279
SAMN29101223
R session info
Details of the R package versions used are given below.
devtools::session_info()## ─ Session info ───────────────────────────────────────────────────────────────
## setting value
## version R version 4.4.3 (2025-02-28)
## os Red Hat Enterprise Linux 8.10 (Ootpa)
## system x86_64, linux-gnu
## ui X11
## language (EN)
## collate en_US.UTF-8
## ctype en_US.UTF-8
## tz Europe/Zurich
## date 2026-03-25
## pandoc 3.8.2.1 @ /home/macnairw/packages/scprocess/.snakemake/conda/4fef11cadd34f9d2d13a0d6139d09340_/bin/ (via rmarkdown)
## quarto NA
##
## ─ Packages ───────────────────────────────────────────────────────────────────
## package * version date (UTC) lib source
## abind * 1.4-8 2024-09-12 [1] CRAN (R 4.4.3)
## assertthat * 0.2.1 2019-03-21 [1] CRAN (R 4.4.3)
## beachmat 2.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## beeswarm 0.4.0 2021-06-01 [1] CRAN (R 4.4.3)
## Biobase * 2.66.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocGenerics * 0.52.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocIO 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocManager 1.30.27 2025-11-14 [1] CRAN (R 4.4.3)
## BiocNeighbors 2.0.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## BiocParallel * 1.40.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocSingular 1.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## BiocStyle * 2.34.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## Biostrings 2.74.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## bitops 1.0-9 2024-10-03 [1] CRAN (R 4.4.3)
## bluster 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## bookdown 0.45 2025-10-03 [1] CRAN (R 4.4.3)
## bslib 0.9.0 2025-01-30 [1] CRAN (R 4.4.3)
## ca 0.71.1 2020-01-24 [1] CRAN (R 4.4.3)
## cachem 1.1.0 2024-05-16 [1] CRAN (R 4.4.3)
## callr 3.7.6 2024-03-25 [1] CRAN (R 4.4.3)
## cellranger 1.1.0 2016-07-27 [1] CRAN (R 4.4.3)
## circlize * 0.4.16 2024-02-20 [1] CRAN (R 4.4.3)
## cli 3.6.5 2025-04-23 [1] CRAN (R 4.4.3)
## clue 0.3-66 2024-11-13 [1] CRAN (R 4.4.3)
## cluster 2.1.8.1 2025-03-12 [1] CRAN (R 4.4.3)
## codetools 0.2-20 2024-03-31 [1] CRAN (R 4.4.3)
## colorspace 2.1-2 2025-09-22 [1] CRAN (R 4.4.3)
## ComplexHeatmap * 2.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## crayon 1.5.3 2024-06-20 [1] CRAN (R 4.4.3)
## curl 7.0.0 2025-08-19 [1] CRAN (R 4.4.3)
## data.table * 1.17.8 2025-07-10 [1] CRAN (R 4.4.3)
## DelayedArray * 0.32.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## devtools 2.4.6 2025-10-03 [1] CRAN (R 4.4.3)
## digest 0.6.39 2025-11-19 [1] CRAN (R 4.4.3)
## doParallel 1.0.17 2022-02-07 [1] CRAN (R 4.4.3)
## dplyr * 1.1.4 2023-11-17 [1] CRAN (R 4.4.3)
## dqrng 0.3.2 2023-11-29 [1] CRAN (R 4.4.3)
## edgeR 4.4.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## ellipsis 0.3.2 2021-04-29 [1] CRAN (R 4.4.3)
## evaluate 1.0.5 2025-08-27 [1] CRAN (R 4.4.3)
## farver 2.1.2 2024-05-13 [1] CRAN (R 4.4.3)
## fastmap 1.2.0 2024-05-15 [1] CRAN (R 4.4.3)
## forcats * 1.0.1 2025-09-25 [1] CRAN (R 4.4.3)
## foreach 1.5.2 2022-02-02 [1] CRAN (R 4.4.3)
## fs 1.6.6 2025-04-12 [1] CRAN (R 4.4.3)
## generics 0.1.4 2025-05-09 [1] CRAN (R 4.4.3)
## GenomeInfoDb * 1.42.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## GenomeInfoDbData 1.2.13 2026-03-05 [1] Bioconductor
## GenomicAlignments 1.42.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## GenomicRanges * 1.58.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## GetoptLong 1.0.5 2020-12-15 [1] CRAN (R 4.4.3)
## getPass 0.2-4 2023-12-10 [1] CRAN (R 4.4.3)
## ggbeeswarm * 0.7.2 2023-04-29 [1] CRAN (R 4.4.3)
## ggh4x * 0.3.1 2025-05-30 [1] CRAN (R 4.4.3)
## ggplot.multistats * 1.0.1 2024-09-25 [1] CRAN (R 4.4.3)
## ggplot2 * 4.0.1 2025-11-14 [1] CRAN (R 4.4.3)
## ggrepel * 0.9.6 2024-09-07 [1] CRAN (R 4.4.3)
## git2r 0.35.0 2024-10-20 [1] CRAN (R 4.4.3)
## GlobalOptions 0.1.2 2020-06-10 [1] CRAN (R 4.4.3)
## glue 1.8.0 2024-09-30 [1] CRAN (R 4.4.3)
## gridExtra 2.3 2017-09-09 [1] CRAN (R 4.4.3)
## gtable 0.3.6 2024-10-25 [1] CRAN (R 4.4.3)
## HDF5Array * 1.34.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## hexbin 1.28.5 2024-11-13 [1] CRAN (R 4.4.3)
## hms 1.1.4 2025-10-17 [1] CRAN (R 4.4.3)
## htmltools 0.5.8.1 2024-04-04 [1] CRAN (R 4.4.3)
## httpuv 1.6.16 2025-04-16 [1] CRAN (R 4.4.3)
## httr 1.4.7 2023-08-15 [1] CRAN (R 4.4.3)
## igraph 2.1.4 2025-01-23 [1] CRAN (R 4.4.3)
## IRanges * 2.40.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## irlba 2.3.5.1 2022-10-03 [1] CRAN (R 4.4.3)
## iterators 1.0.14 2022-02-05 [1] CRAN (R 4.4.3)
## jquerylib 0.1.4 2021-04-26 [1] CRAN (R 4.4.3)
## jsonlite 2.0.0 2025-03-27 [1] CRAN (R 4.4.3)
## knitr 1.50 2025-03-16 [1] CRAN (R 4.4.3)
## labeling 0.4.3 2023-08-29 [1] CRAN (R 4.4.3)
## later 1.4.4 2025-08-27 [1] CRAN (R 4.4.3)
## lattice 0.22-7 2025-04-02 [1] CRAN (R 4.4.3)
## lifecycle 1.0.4 2023-11-07 [1] CRAN (R 4.4.3)
## limma 3.62.1 2024-11-03 [1] Bioconductor 3.20 (R 4.4.2)
## locfit 1.5-9.12 2025-03-05 [1] CRAN (R 4.4.3)
## lubridate * 1.9.4 2024-12-08 [1] CRAN (R 4.4.3)
## magrittr * 2.0.4 2025-09-12 [1] CRAN (R 4.4.3)
## MASS 7.3-65 2025-02-28 [1] CRAN (R 4.4.3)
## Matrix * 1.7-4 2025-08-28 [1] CRAN (R 4.4.3)
## MatrixGenerics * 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## matrixStats * 1.5.0 2025-01-07 [1] CRAN (R 4.4.3)
## memoise 2.0.1 2021-11-26 [1] CRAN (R 4.4.3)
## metapod 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## otel 0.2.0 2025-08-29 [1] CRAN (R 4.4.3)
## patchwork * 1.3.2 2025-08-25 [1] CRAN (R 4.4.3)
## pillar 1.11.1 2025-09-17 [1] CRAN (R 4.4.3)
## pkgbuild 1.4.8 2025-05-26 [1] CRAN (R 4.4.3)
## pkgconfig 2.0.3 2019-09-22 [1] CRAN (R 4.4.3)
## pkgload 1.4.1 2025-09-23 [1] CRAN (R 4.4.3)
## plyr 1.8.9 2023-10-02 [1] CRAN (R 4.4.3)
## png 0.1-8 2022-11-29 [1] CRAN (R 4.4.3)
## processx 3.8.6 2025-02-21 [1] CRAN (R 4.4.3)
## promises 1.5.0 2025-11-01 [1] CRAN (R 4.4.3)
## ps 1.9.1 2025-04-12 [1] CRAN (R 4.4.3)
## purrr * 1.2.0 2025-11-04 [1] CRAN (R 4.4.3)
## R.methodsS3 1.8.2 2022-06-13 [1] CRAN (R 4.4.3)
## R.oo 1.27.1 2025-05-02 [1] CRAN (R 4.4.3)
## R.utils 2.13.0 2025-02-24 [1] CRAN (R 4.4.3)
## R6 2.6.1 2025-02-15 [1] CRAN (R 4.4.3)
## RColorBrewer * 1.1-3 2022-04-03 [1] CRAN (R 4.4.3)
## Rcpp 1.1.0 2025-07-02 [1] CRAN (R 4.4.3)
## RCurl 1.98-1.17 2025-03-22 [1] CRAN (R 4.4.3)
## readr * 2.1.6 2025-11-14 [1] CRAN (R 4.4.3)
## readxl * 1.4.5 2025-03-07 [1] CRAN (R 4.4.3)
## registry 0.5-1 2019-03-05 [1] CRAN (R 4.4.3)
## remotes 2.5.0 2024-03-17 [1] CRAN (R 4.4.3)
## restfulr 0.0.16 2025-06-27 [1] CRAN (R 4.4.3)
## rhdf5 * 2.50.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## rhdf5filters 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## Rhdf5lib 1.28.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## rjson 0.2.23 2024-09-16 [1] CRAN (R 4.4.3)
## rlang 1.1.6 2025-04-11 [1] CRAN (R 4.4.3)
## rmarkdown 2.30 2025-09-28 [1] CRAN (R 4.4.3)
## rmdformats 1.0.4 2022-05-17 [1] CRAN (R 4.4.3)
## rprojroot 2.1.1 2025-08-26 [1] CRAN (R 4.4.3)
## Rsamtools 2.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## rstudioapi 0.17.1 2024-10-22 [1] CRAN (R 4.4.3)
## rsvd 1.0.5 2021-04-16 [1] CRAN (R 4.4.1)
## rtracklayer 1.66.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## S4Arrays * 1.6.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## S4Vectors * 0.44.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## S7 0.2.1 2025-11-14 [1] CRAN (R 4.4.3)
## sass 0.4.10 2025-04-11 [1] CRAN (R 4.4.3)
## ScaledMatrix 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## scales * 1.4.0 2025-04-24 [1] CRAN (R 4.4.3)
## scater * 1.34.1 2025-03-03 [1] Bioconductor 3.20 (R 4.4.2)
## scDblFinder * 1.23.4 2025-08-22 [1] Bioconductor 3.22 (R 4.4.3)
## scran 1.34.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## scuttle * 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## seriation * 1.5.8 2025-08-20 [1] CRAN (R 4.4.3)
## sessioninfo 1.2.3 2025-02-05 [1] CRAN (R 4.4.3)
## shape 1.4.6.1 2024-02-23 [1] CRAN (R 4.4.3)
## SingleCellExperiment * 1.28.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## SparseArray * 1.6.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## statmod 1.5.1 2025-10-09 [1] CRAN (R 4.4.3)
## strex * 2.0.1 2024-10-03 [1] CRAN (R 4.4.3)
## stringi 1.8.7 2025-03-27 [1] CRAN (R 4.4.3)
## stringr * 1.6.0 2025-11-04 [1] CRAN (R 4.4.3)
## SummarizedExperiment * 1.36.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## tibble * 3.3.0 2025-06-08 [1] CRAN (R 4.4.3)
## tidyr * 1.3.1 2024-01-24 [1] CRAN (R 4.4.3)
## tidyselect 1.2.1 2024-03-11 [1] CRAN (R 4.4.3)
## tidyverse * 2.0.0 2023-02-22 [1] CRAN (R 4.4.3)
## timechange 0.3.0 2024-01-18 [1] CRAN (R 4.4.3)
## TSP 1.2.6 2025-11-27 [1] CRAN (R 4.4.3)
## tzdb 0.5.0 2025-03-15 [1] CRAN (R 4.4.3)
## UCSC.utils 1.2.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## UpSetR * 1.4.0 2019-05-22 [1] CRAN (R 4.4.3)
## usethis 3.2.1 2025-09-06 [1] CRAN (R 4.4.3)
## vctrs 0.6.5 2023-12-01 [1] CRAN (R 4.4.3)
## vipor 0.4.7 2023-12-18 [1] CRAN (R 4.4.3)
## viridis * 0.6.5 2024-01-29 [1] CRAN (R 4.4.3)
## viridisLite * 0.4.2 2023-05-02 [1] CRAN (R 4.4.3)
## whisker 0.4.1 2022-12-05 [1] CRAN (R 4.4.3)
## withr 3.0.2 2024-10-28 [1] CRAN (R 4.4.3)
## workflowr * 1.7.2 2025-08-18 [1] CRAN (R 4.4.3)
## xfun 0.54 2025-10-30 [1] CRAN (R 4.4.3)
## xgboost 3.1.2.1 2026-01-06 [1] local
## XML 3.99-0.20 2025-11-08 [1] CRAN (R 4.4.3)
## XVector 0.46.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## yaml * 2.3.11 2025-11-28 [1] CRAN (R 4.4.3)
## zlibbioc 1.52.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
##
## [1] /home/macnairw/packages/scprocess/.snakemake/conda/4fef11cadd34f9d2d13a0d6139d09340_/lib/R/library
## * ── Packages attached to the search path.
##
## ──────────────────────────────────────────────────────────────────────────────