Clusters over UMAP
Clustering of data is performed at resolution 0.2. Cluster membership of cells is displayed on a UMAP.
g_cl = plot_umap_cluster(
umap_dt = int_dt[, .(cell_id, UMAP1, UMAP2) ],
clust_dt = int_dt[, .(cell_id, cluster) ],
name = sprintf('res = %s', sel_res))
g_dens = plot_umap_density(int_dt[, .(cell_id, UMAP1, UMAP2) ])
g = g_cl + g_dens
print(g)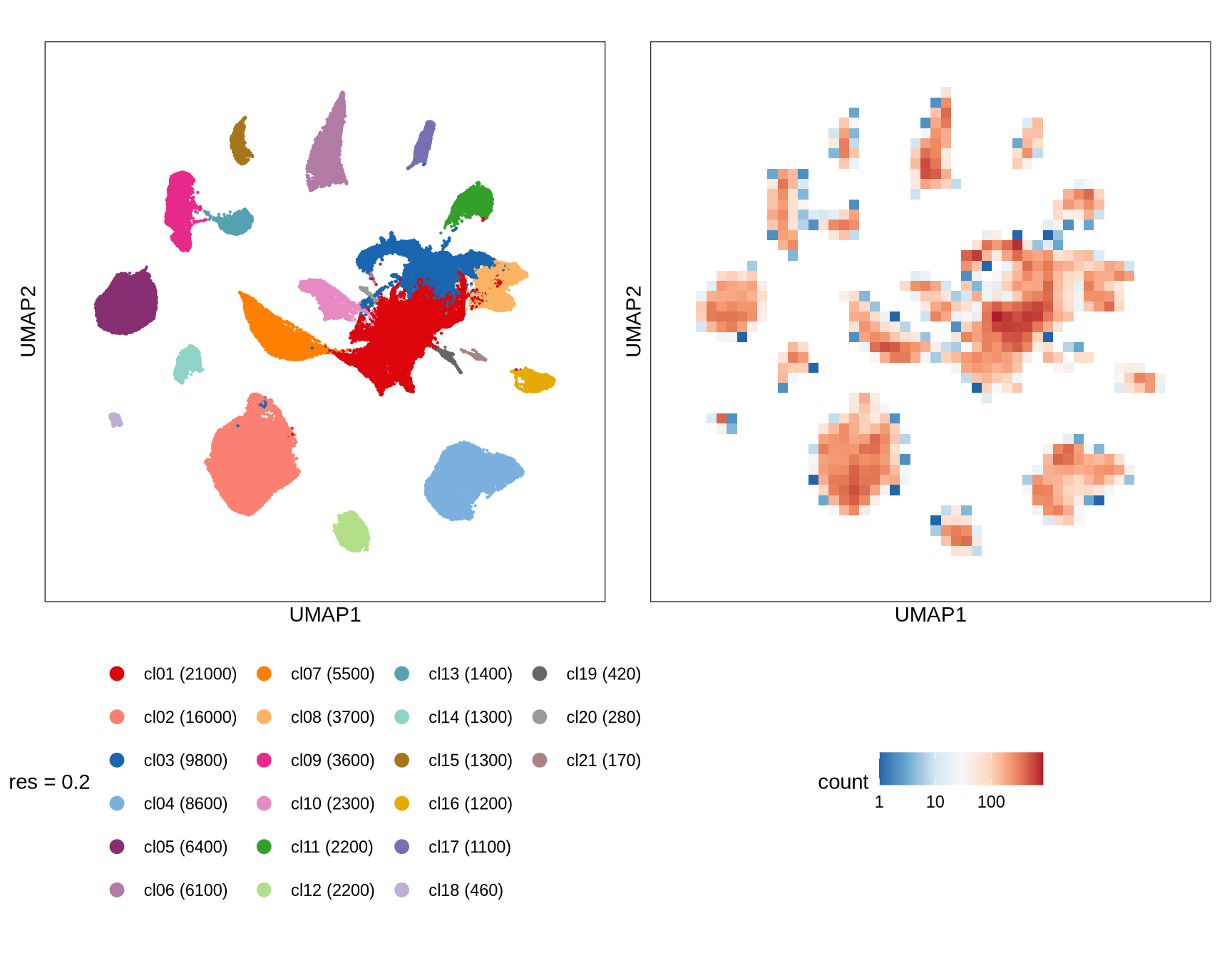
Marker genes
To identify marker genes for each cluster, pseudobulk counts are
generated by aggregating the expression values of cells within a
specific cluster for each sample. Similarly, pseudobulk counts are
generated for the remaining clusters by aggregating expression values
separately for each sample. The resulting pseudobulk values for the
target cluster are then compared to those of the remaining clusters
using edgeR.
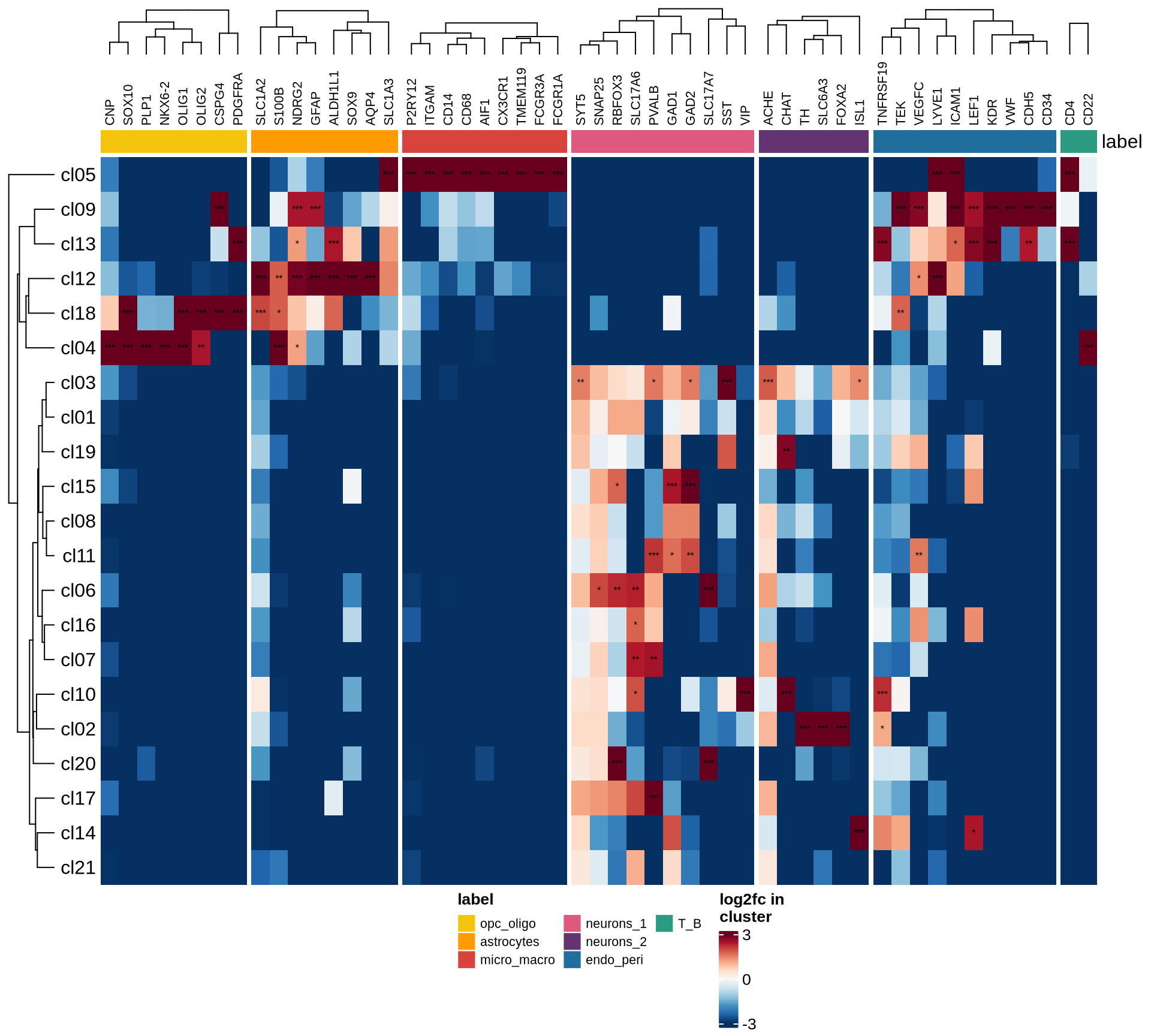
Heatmaps of marker genes
The resulting log2 fold change values (calculated using
edgeR) per cluster are shown for several genesets:
- selected canonical marker genes, if specified in the config file;
- the 50 most highly variable genes; and
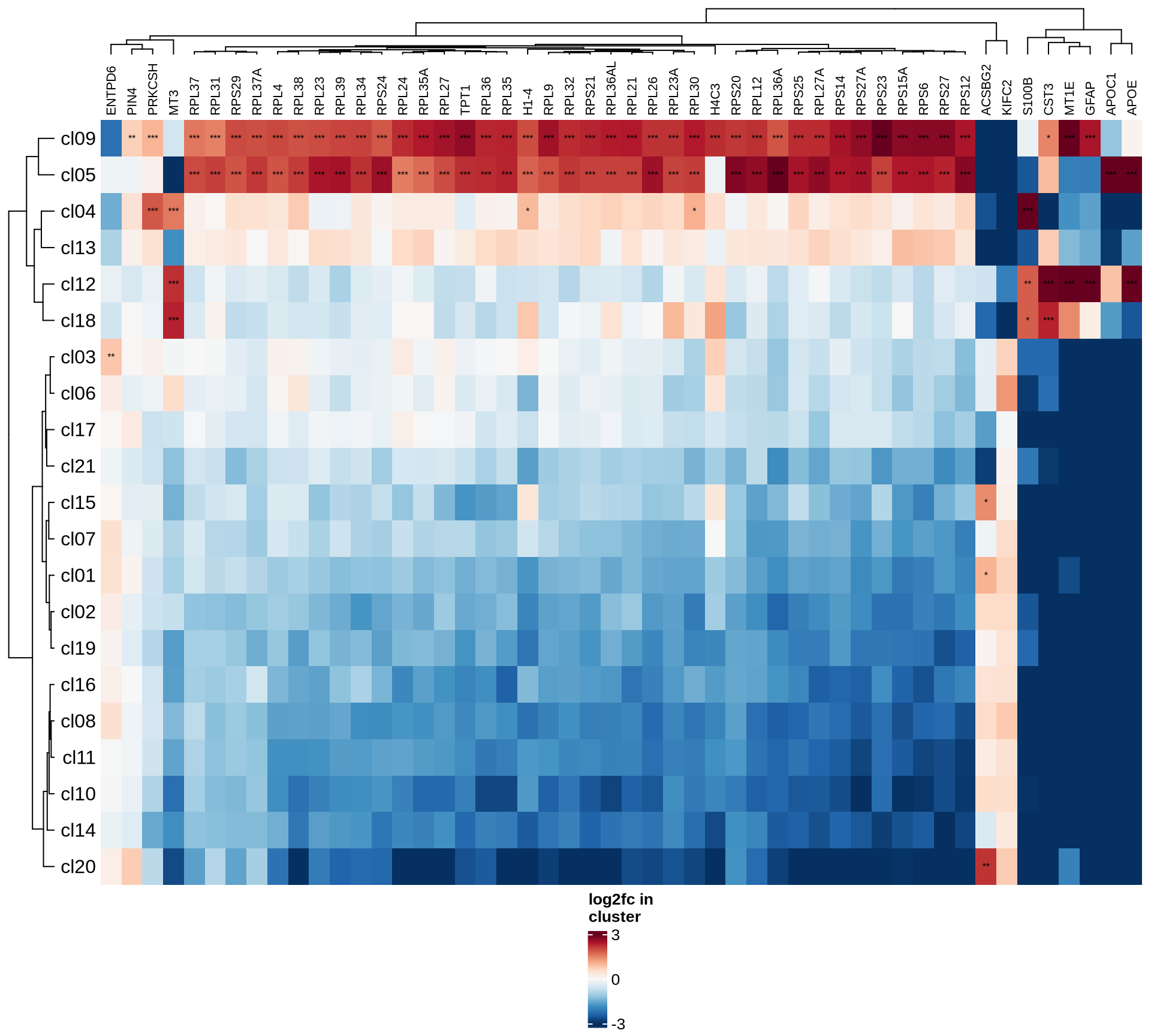
- the 50 genes most likely to represent ambient RNA contamination.
for (nn in names(mkrs_ls)) {
cat("#### ", nn, "\n")
suppressMessages(draw( plot_heatmap_of_selected_genes(mkrs_dt, mkrs_ls[[nn]]),
heatmap_legend_side = "bottom", annotation_legend_side = "bottom", merge_legend = TRUE ))
cat("\n\n")
}human_brain
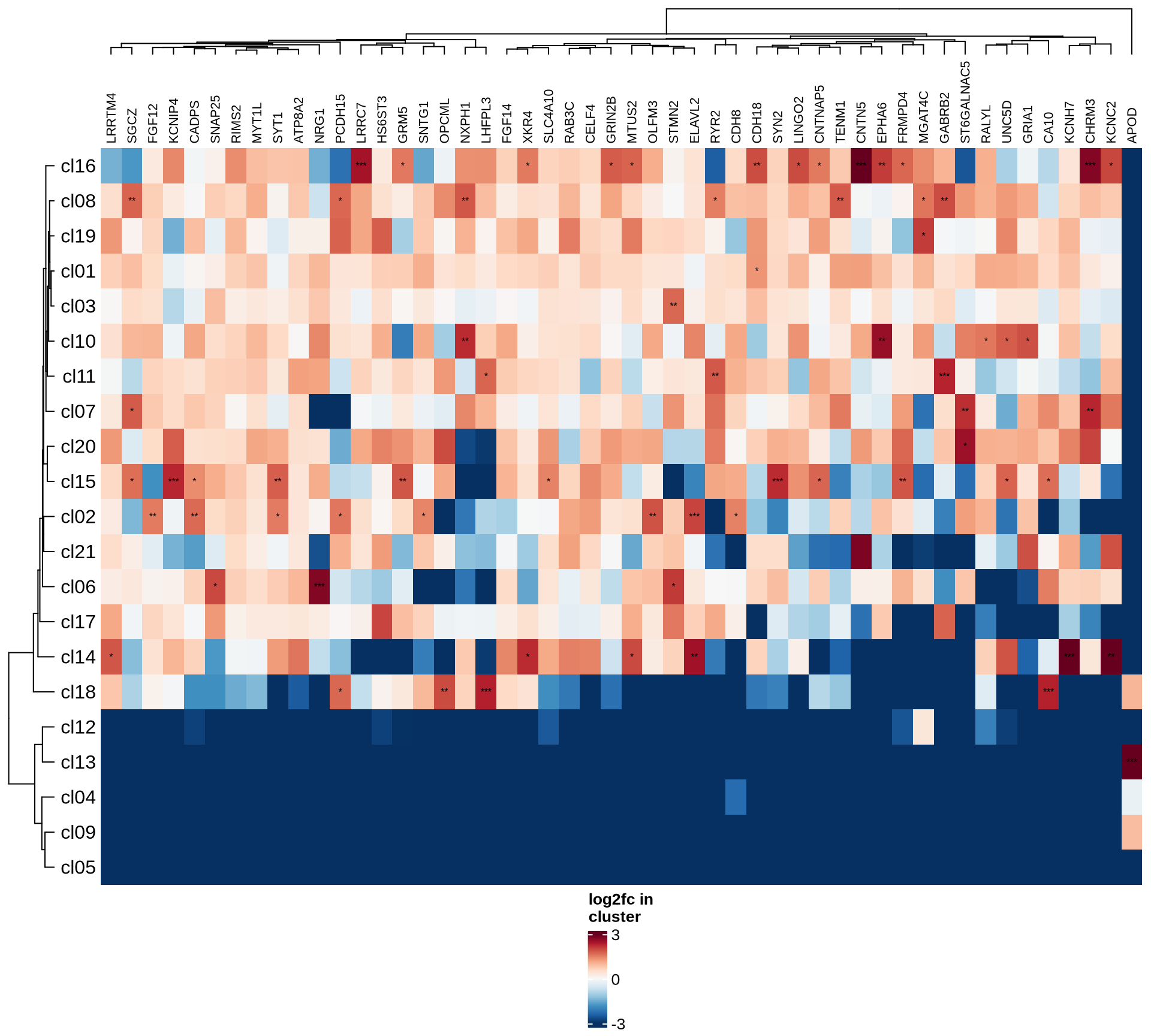
HVGs
ambient
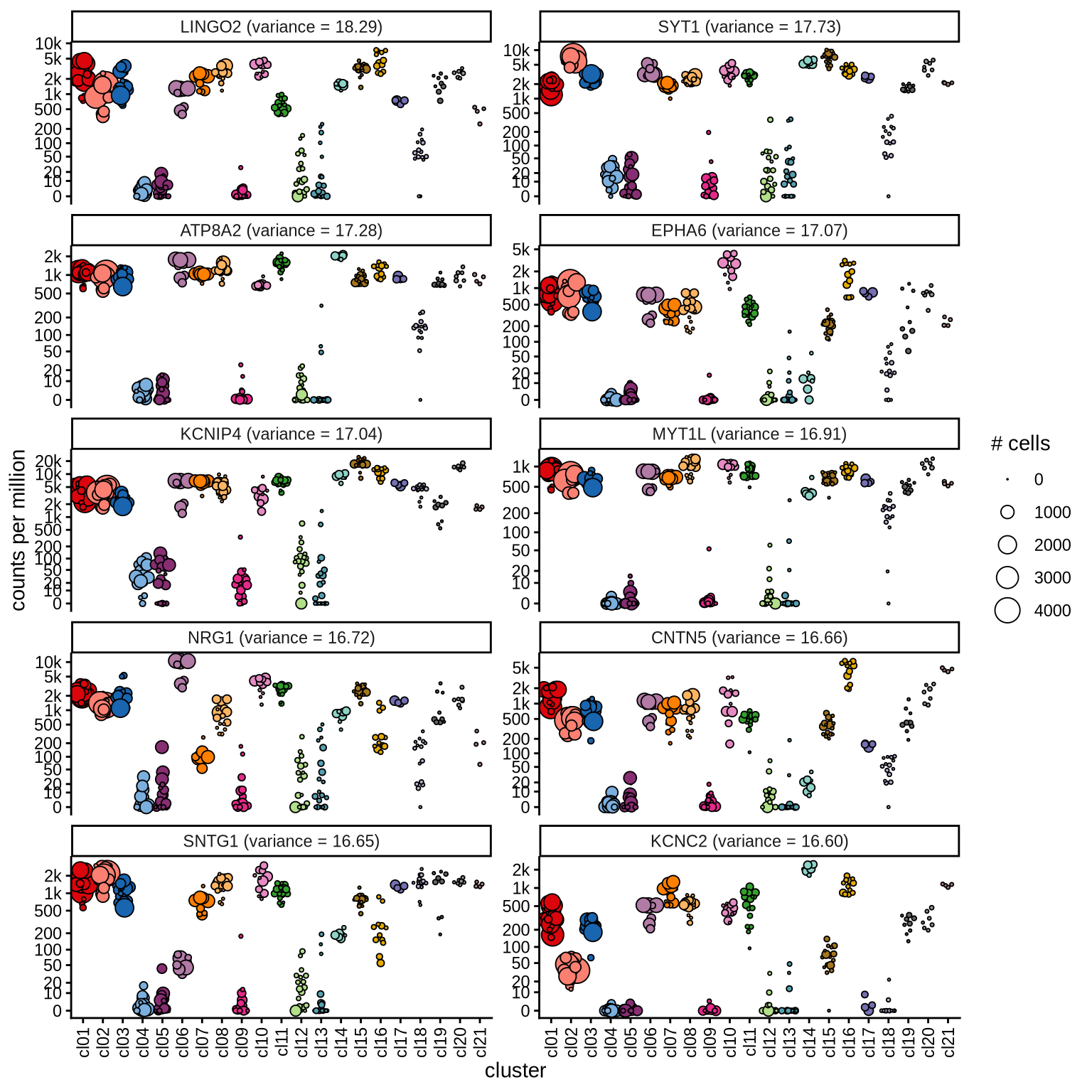
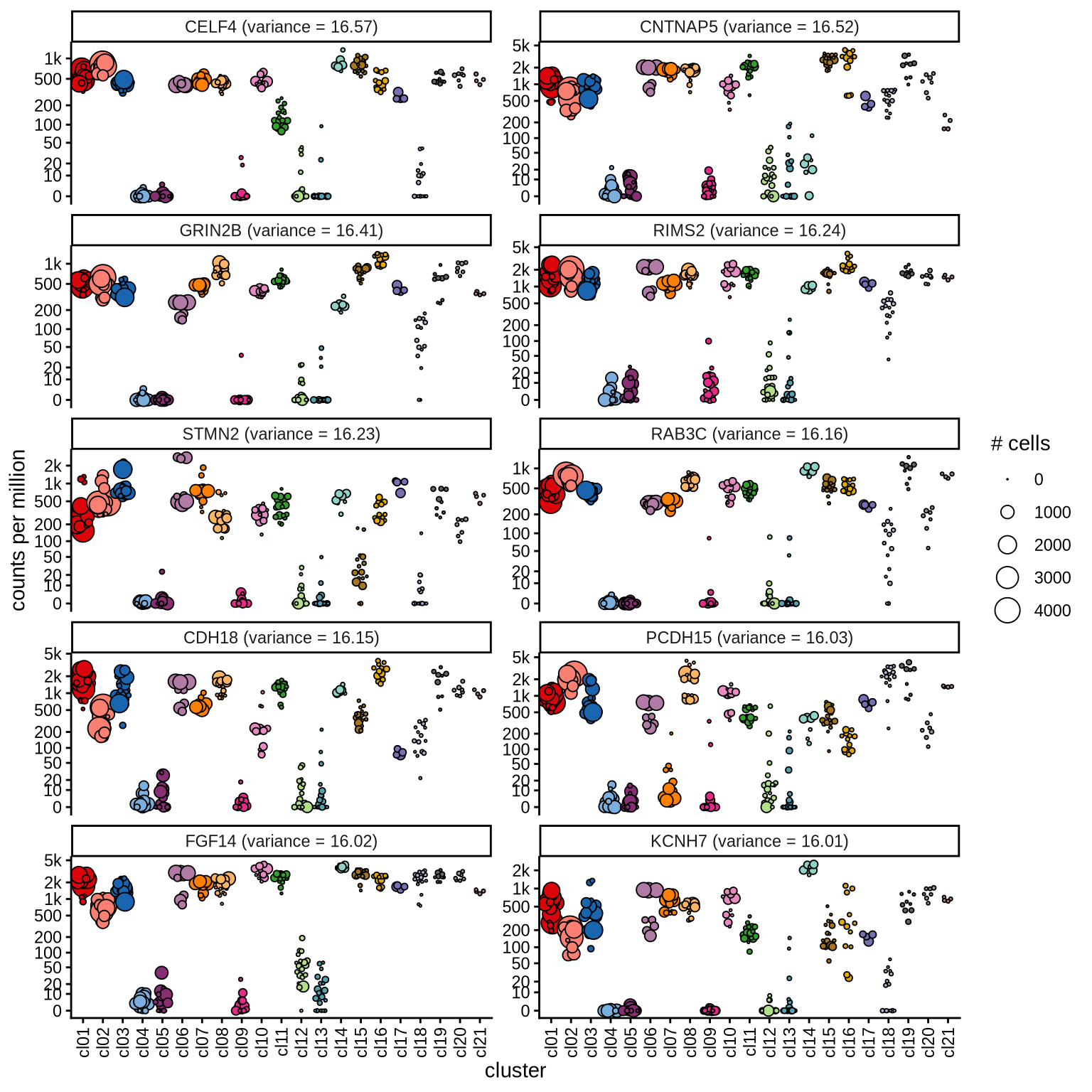
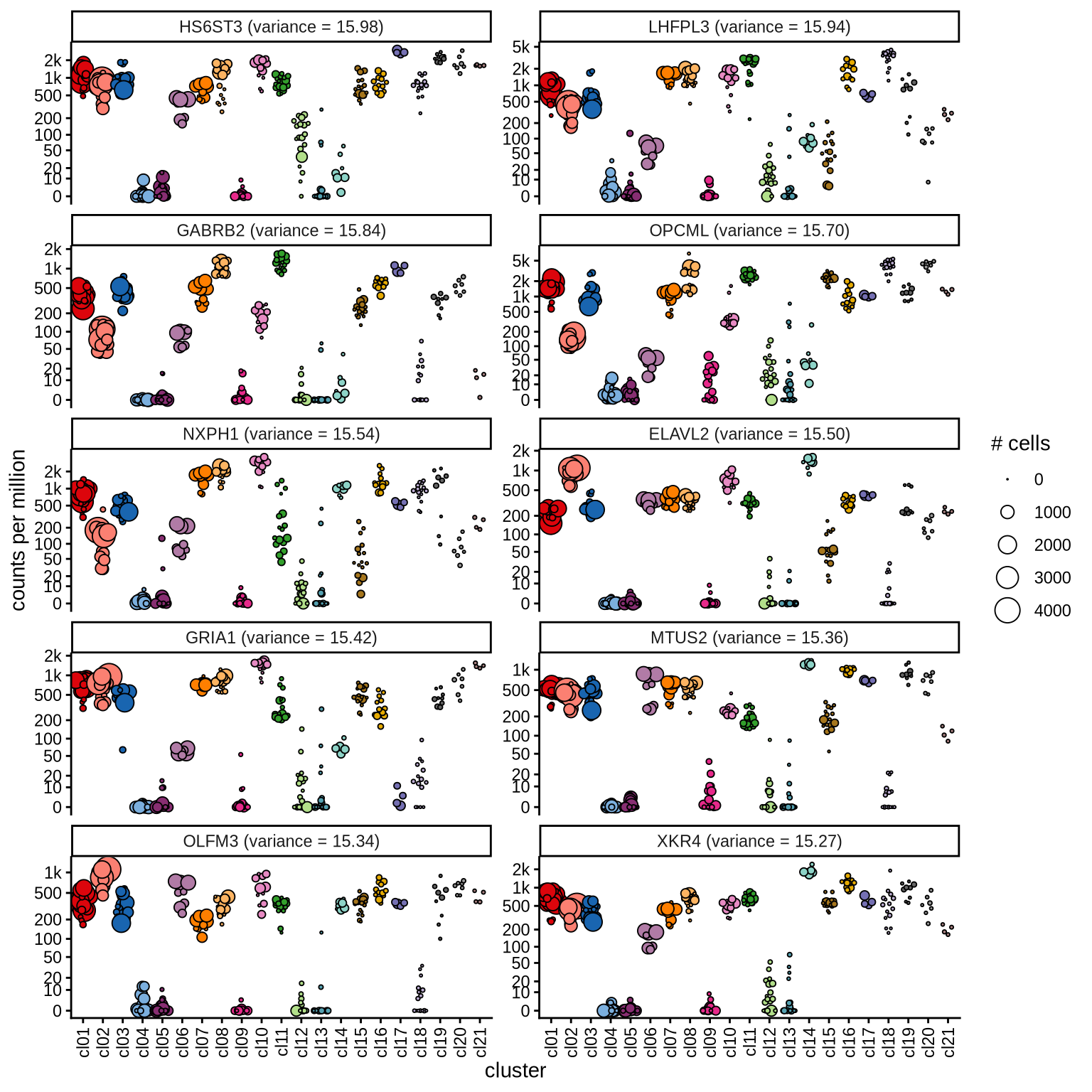
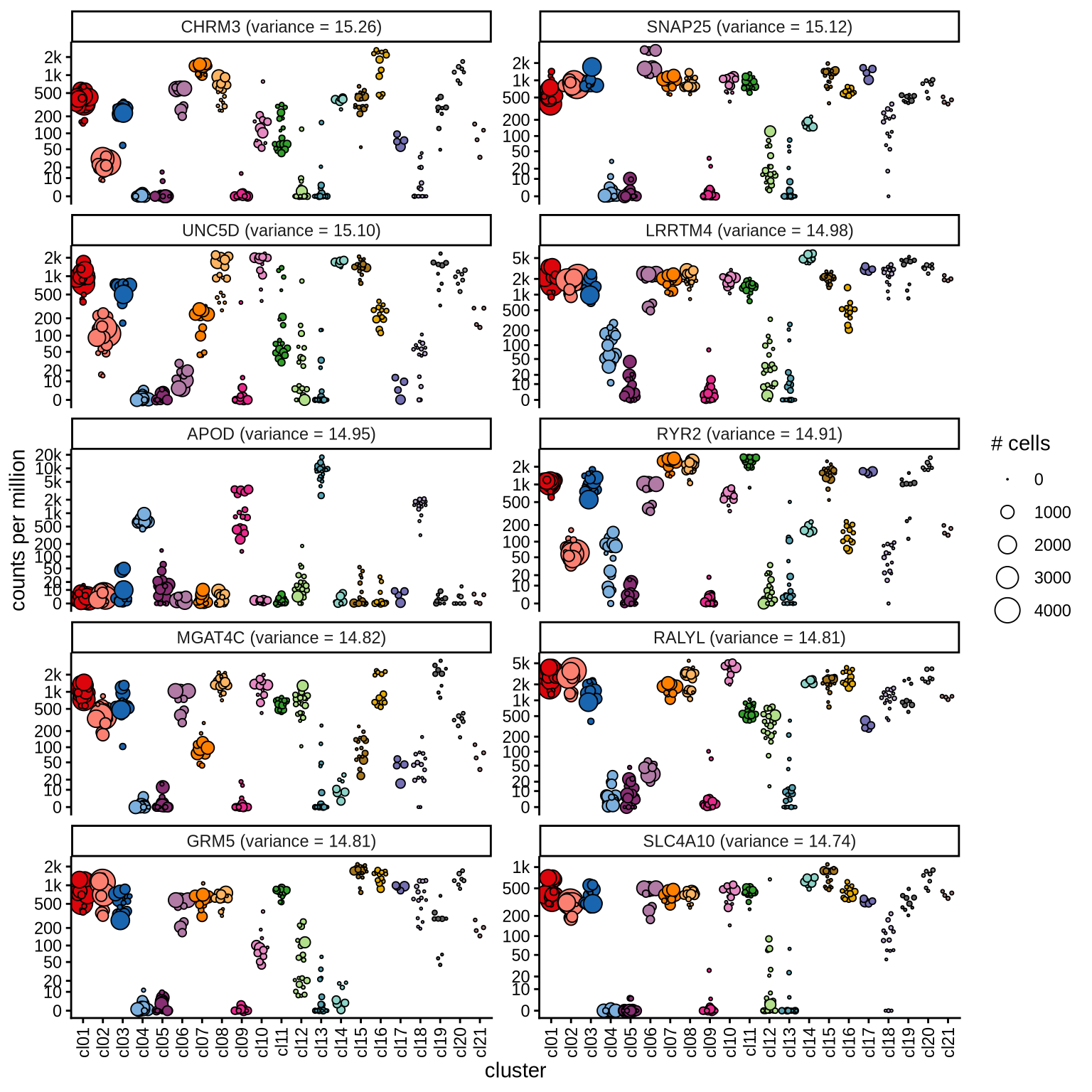
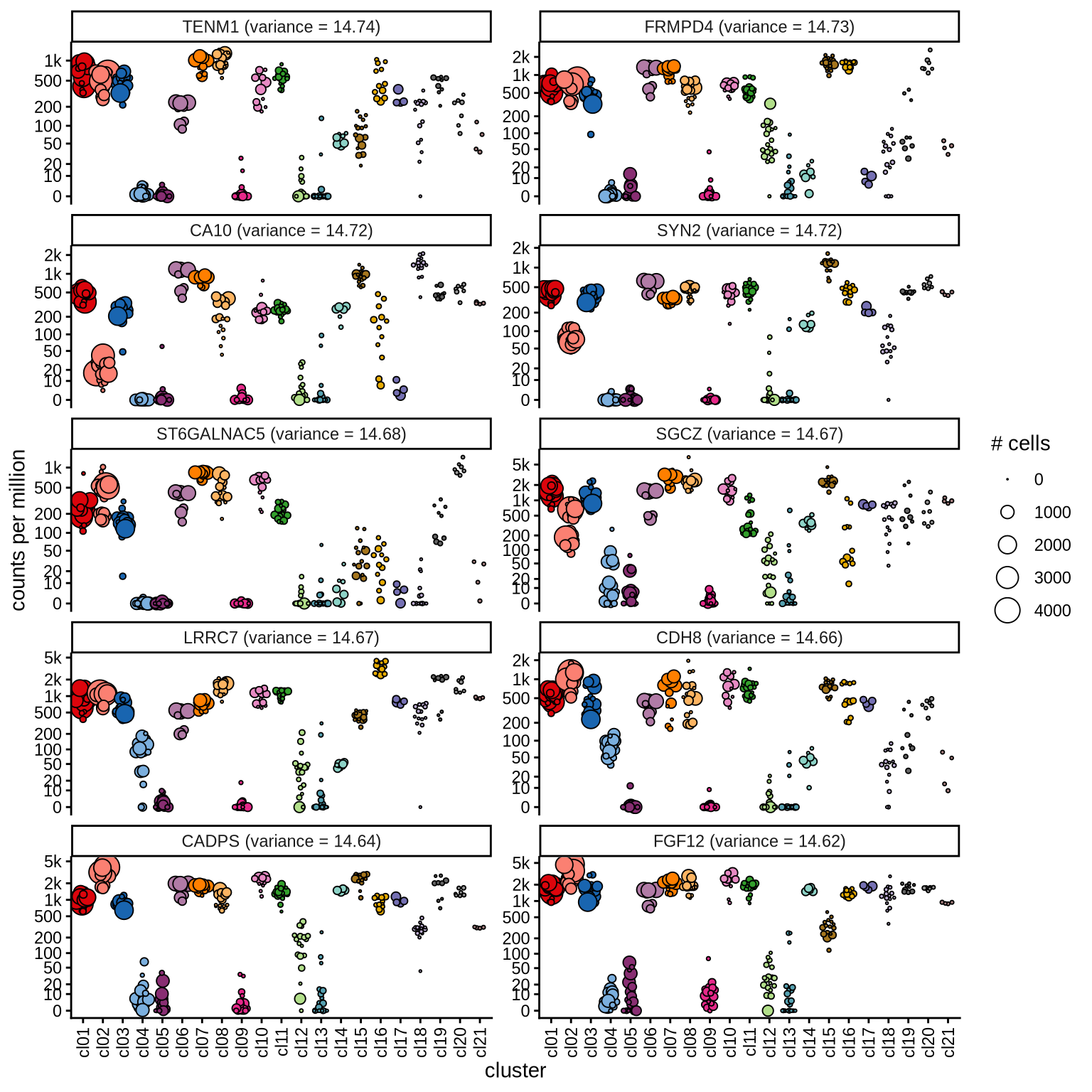
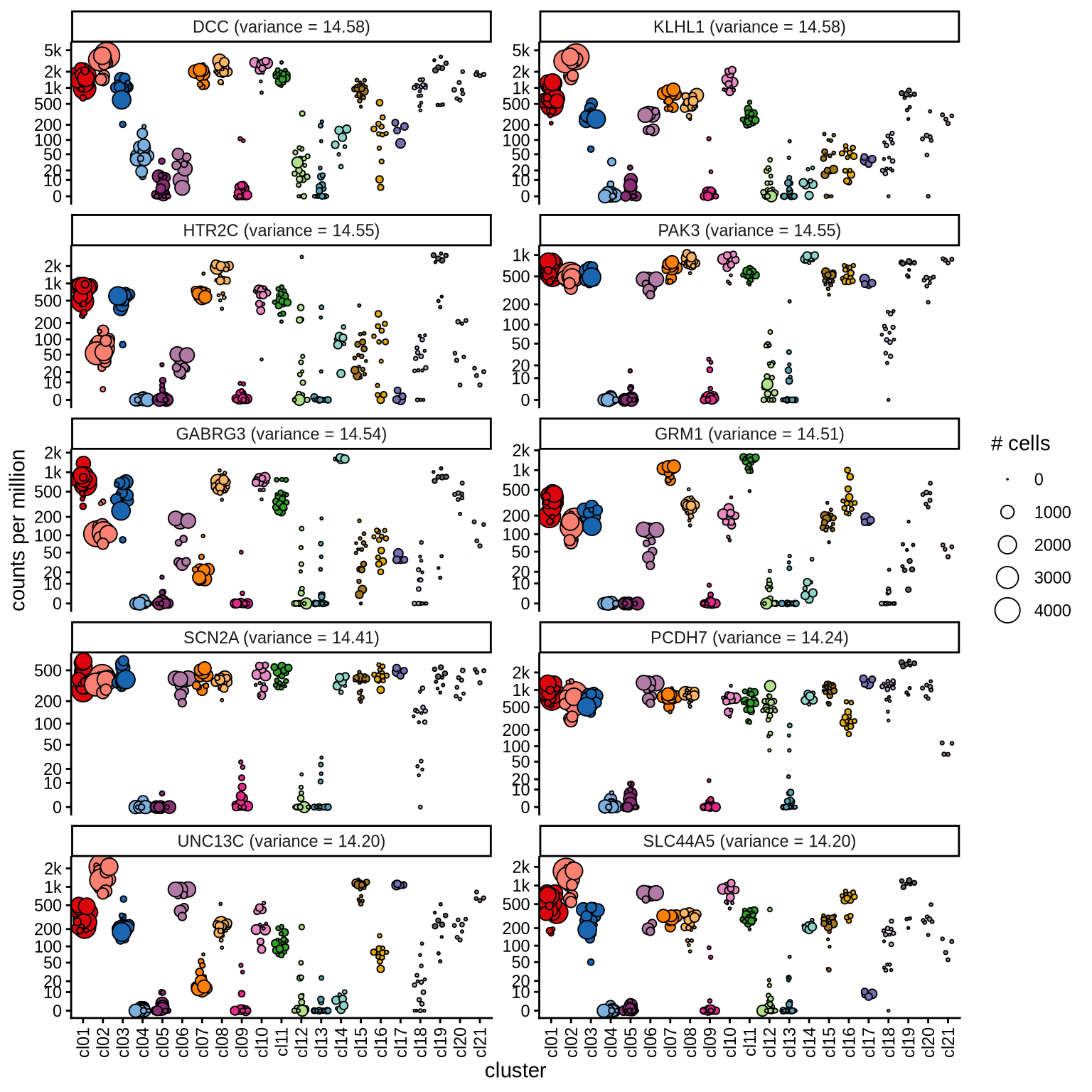
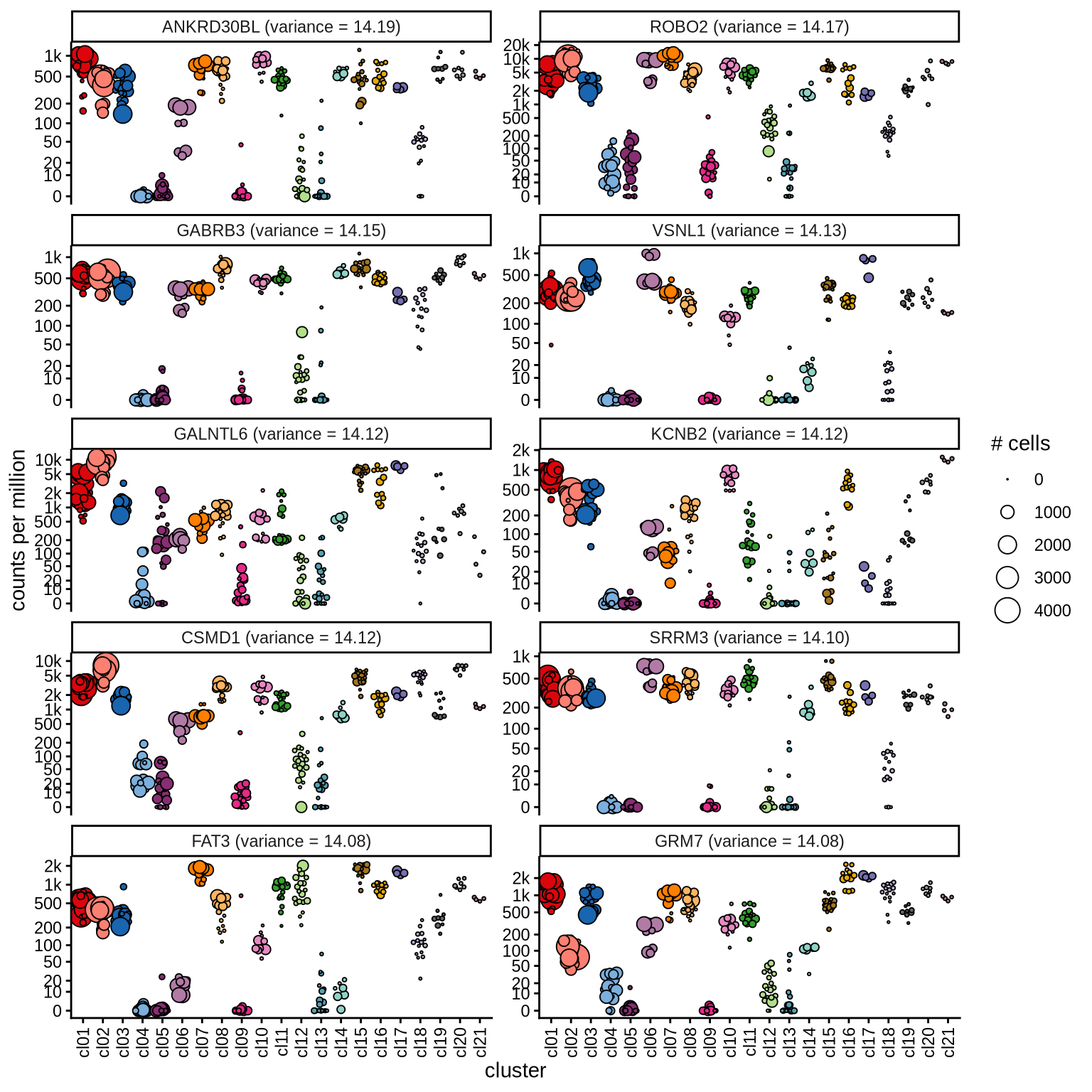
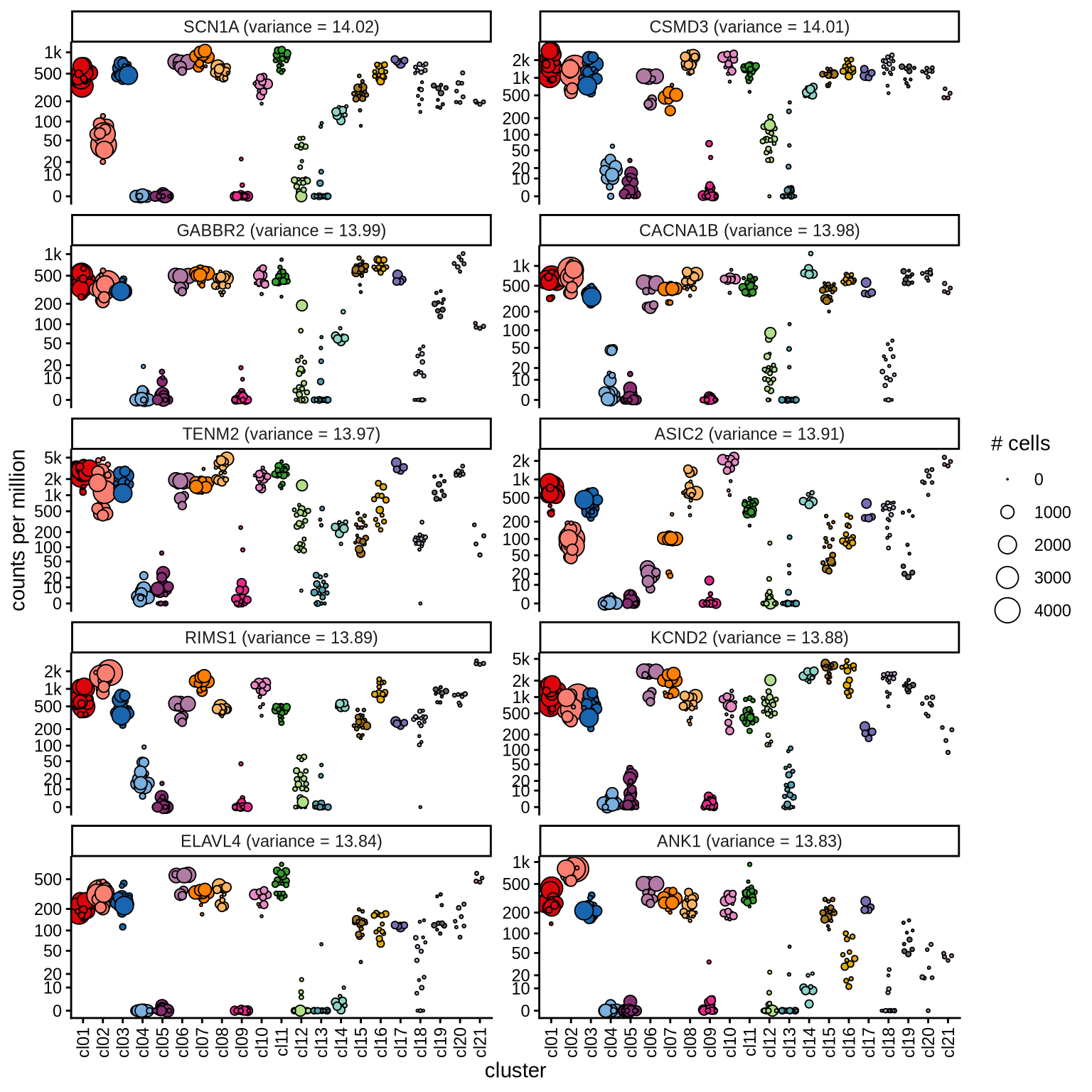
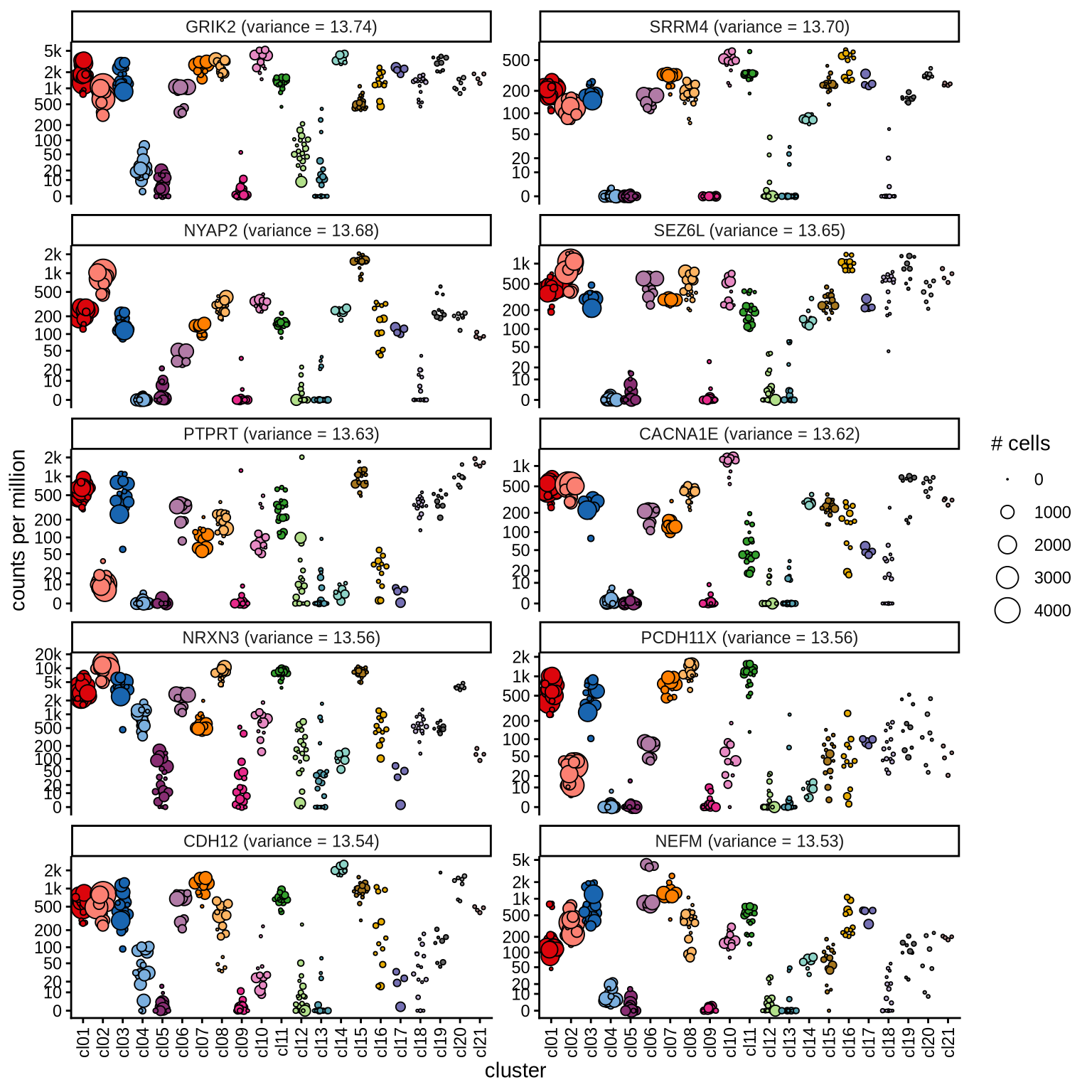
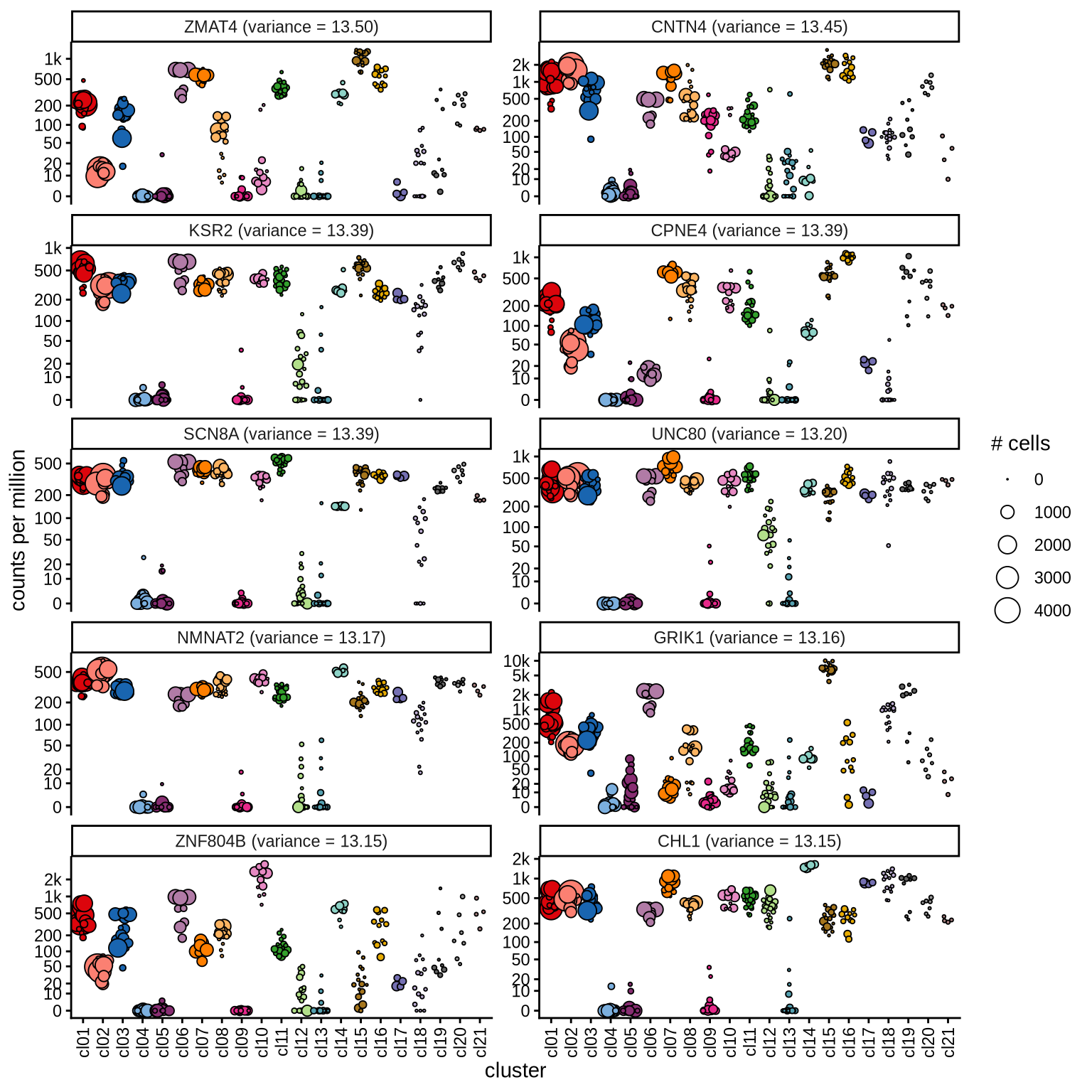
Highly variable genes
Normalized pseudobulk expression values for top 100 highly variable
genes are shown for each cluster, in descending order of variance
(variance calculated using DESeq2::vst).
n_pages = 10
per_p = 10
p = 1
for (i in seq(1, n_pages * per_p, by = per_p)) {
cat('#### page', p, '\n'); p = p + 1
sel_hvgs = hvgs_dt[ i:(i + 9) ]
print(plot_selected_genes(sel_hvgs, cpms_dt, cl_order = cl_ord))
cat('\n\n')
}page 1
page 2
page 3
page 4
page 5
page 6
page 7
page 8
page 9
page 10
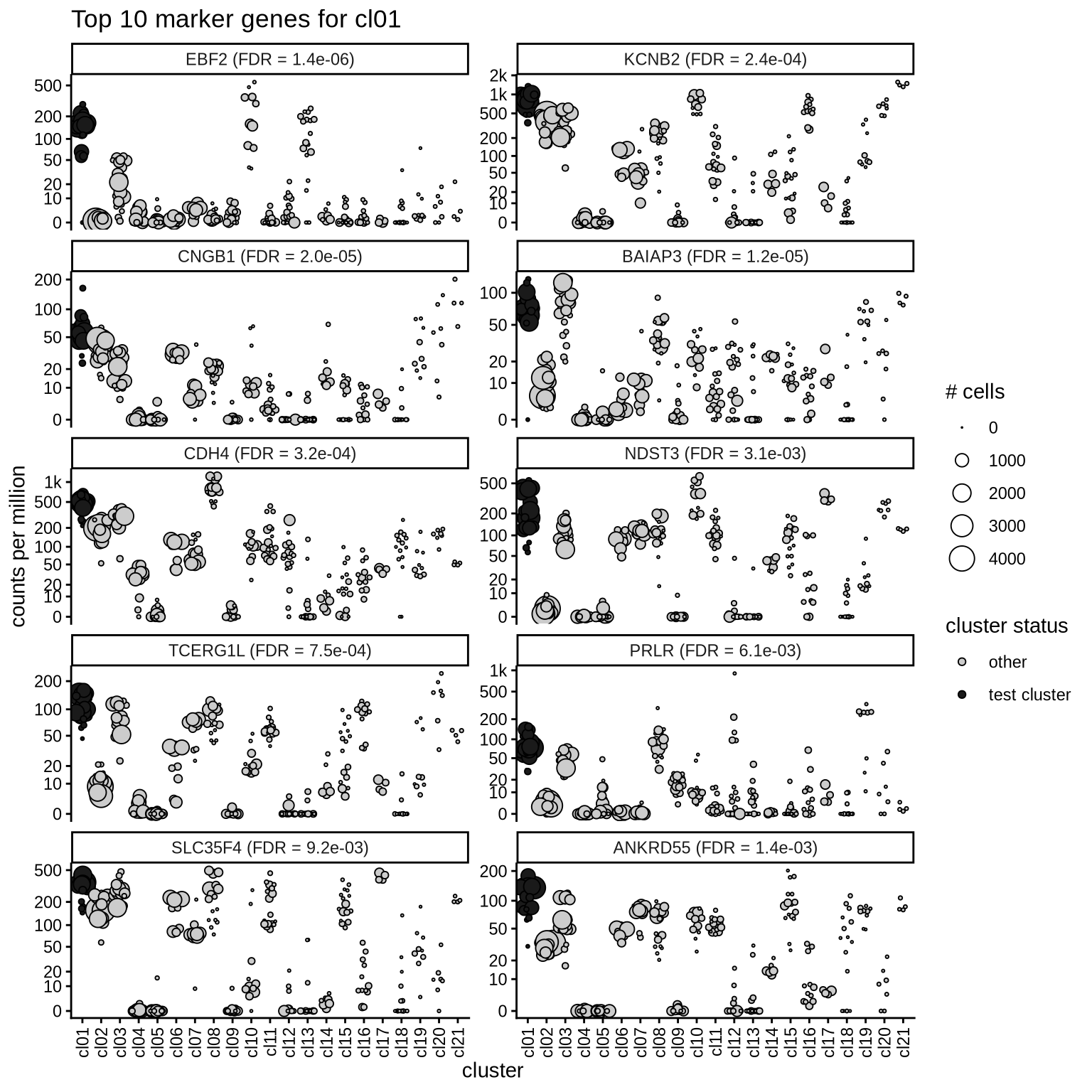
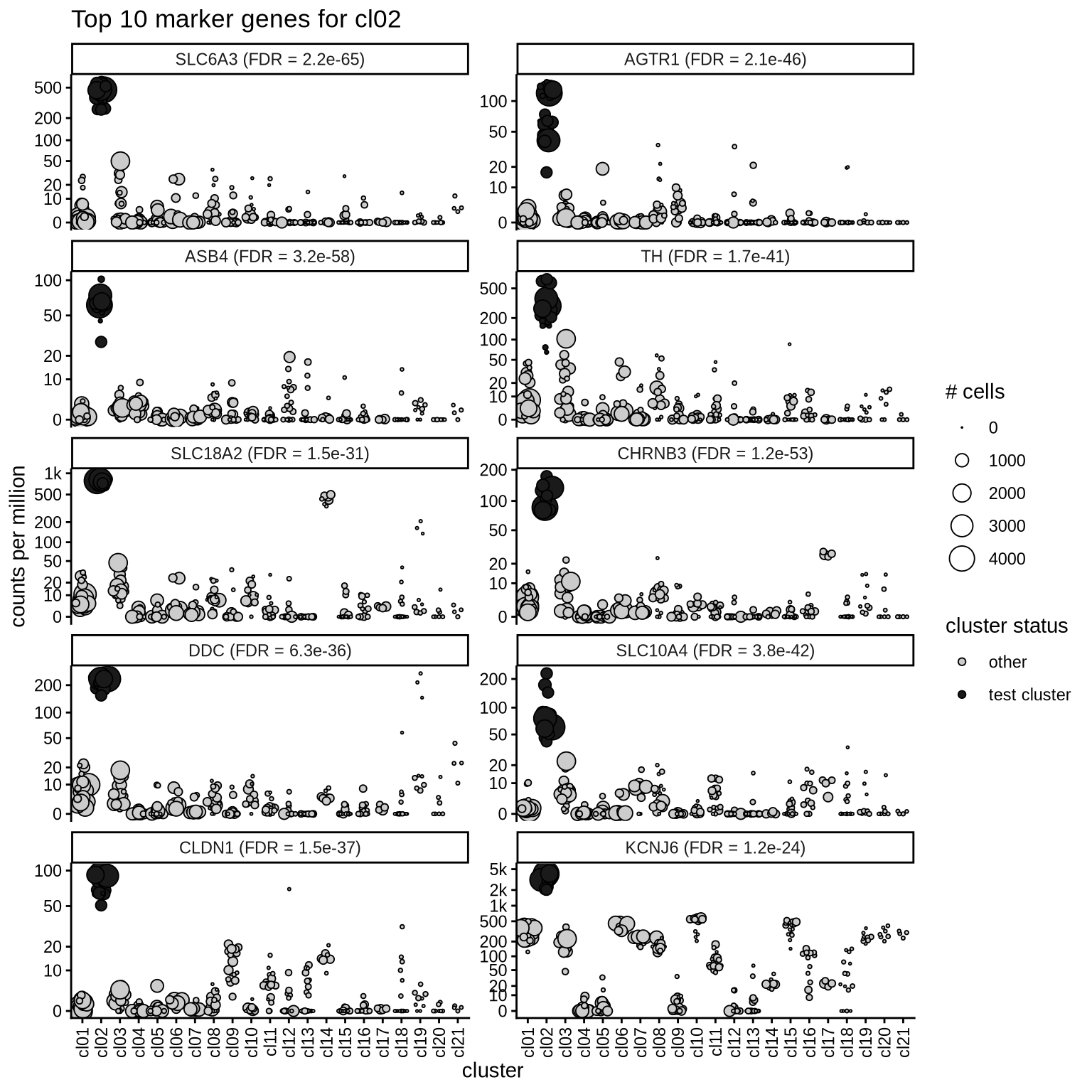
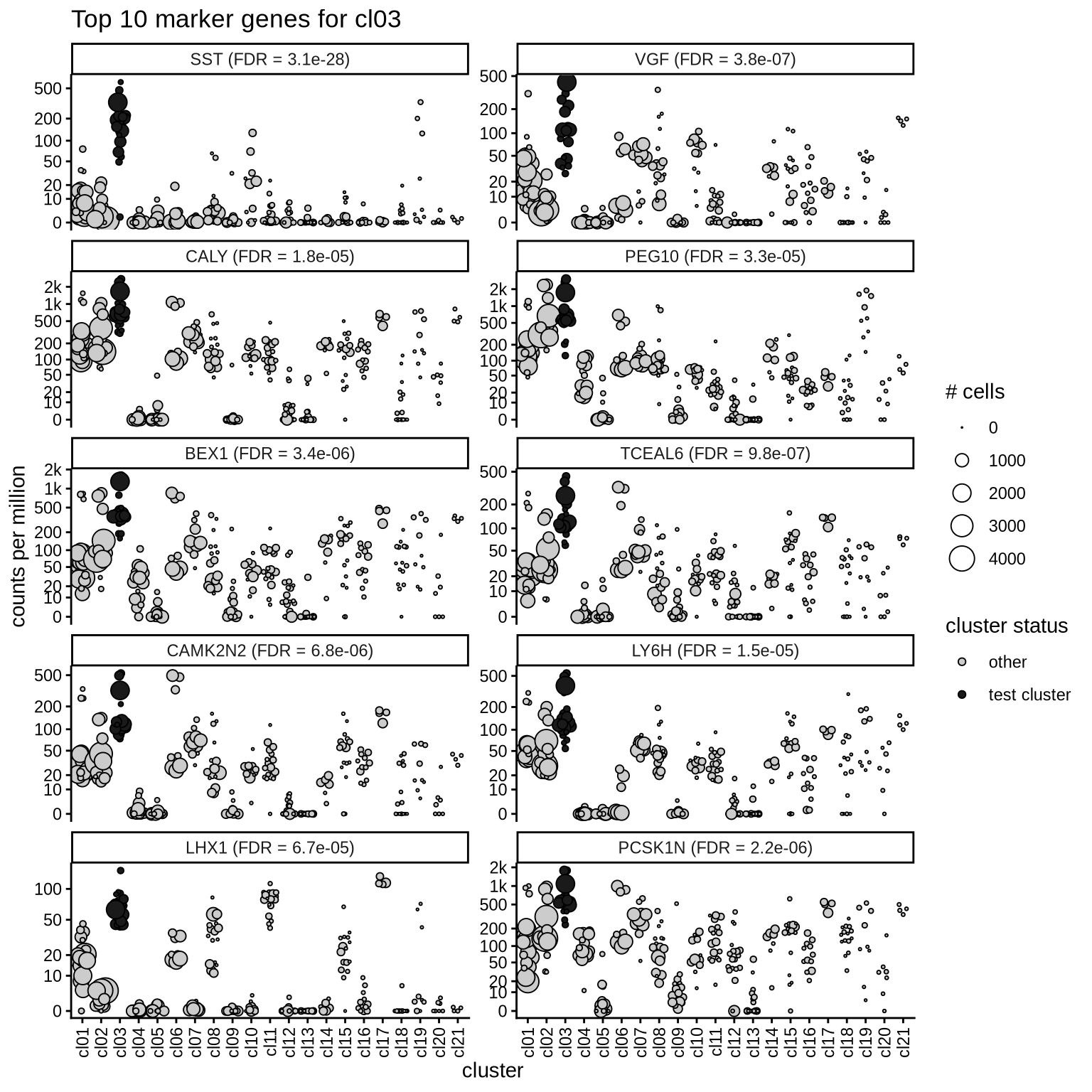
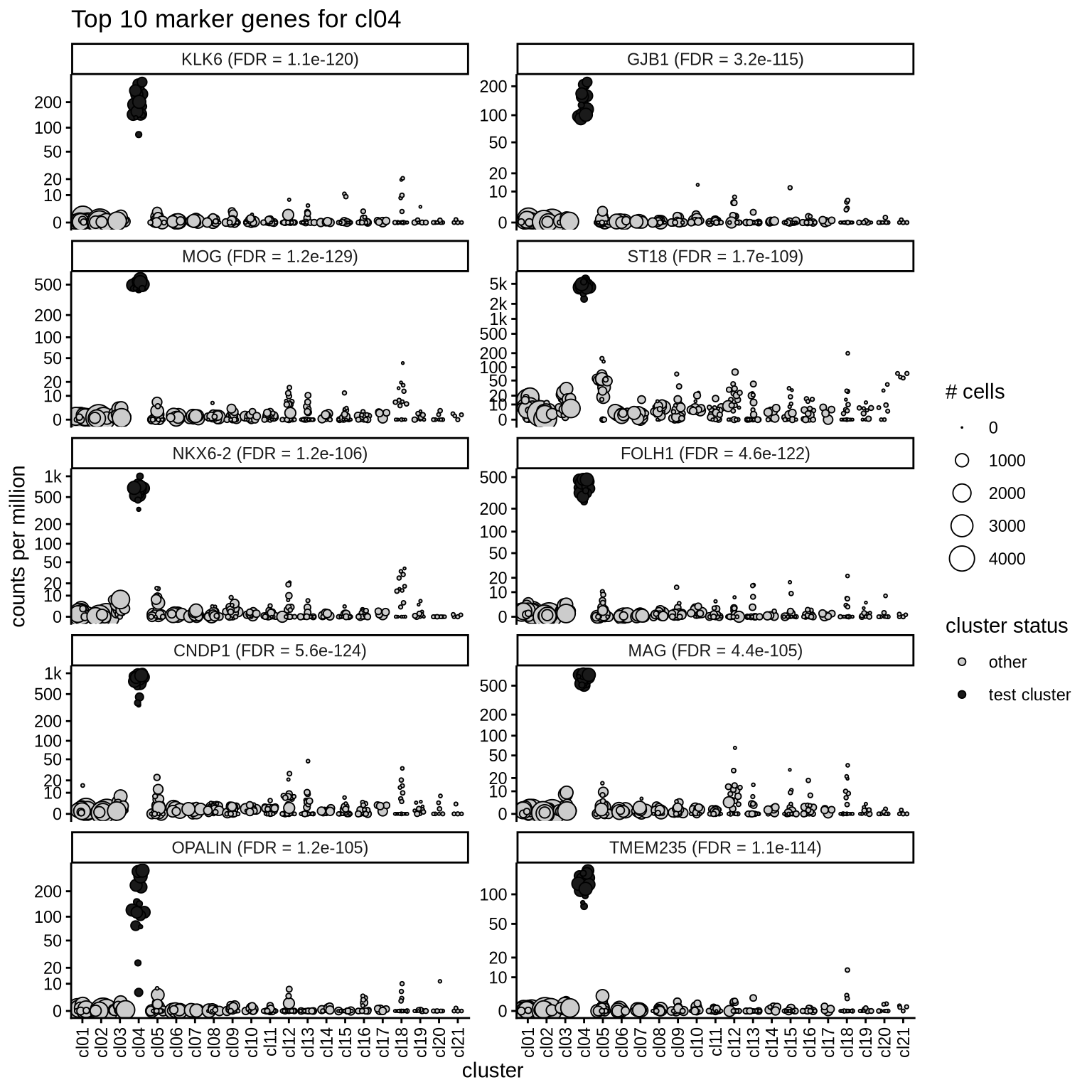
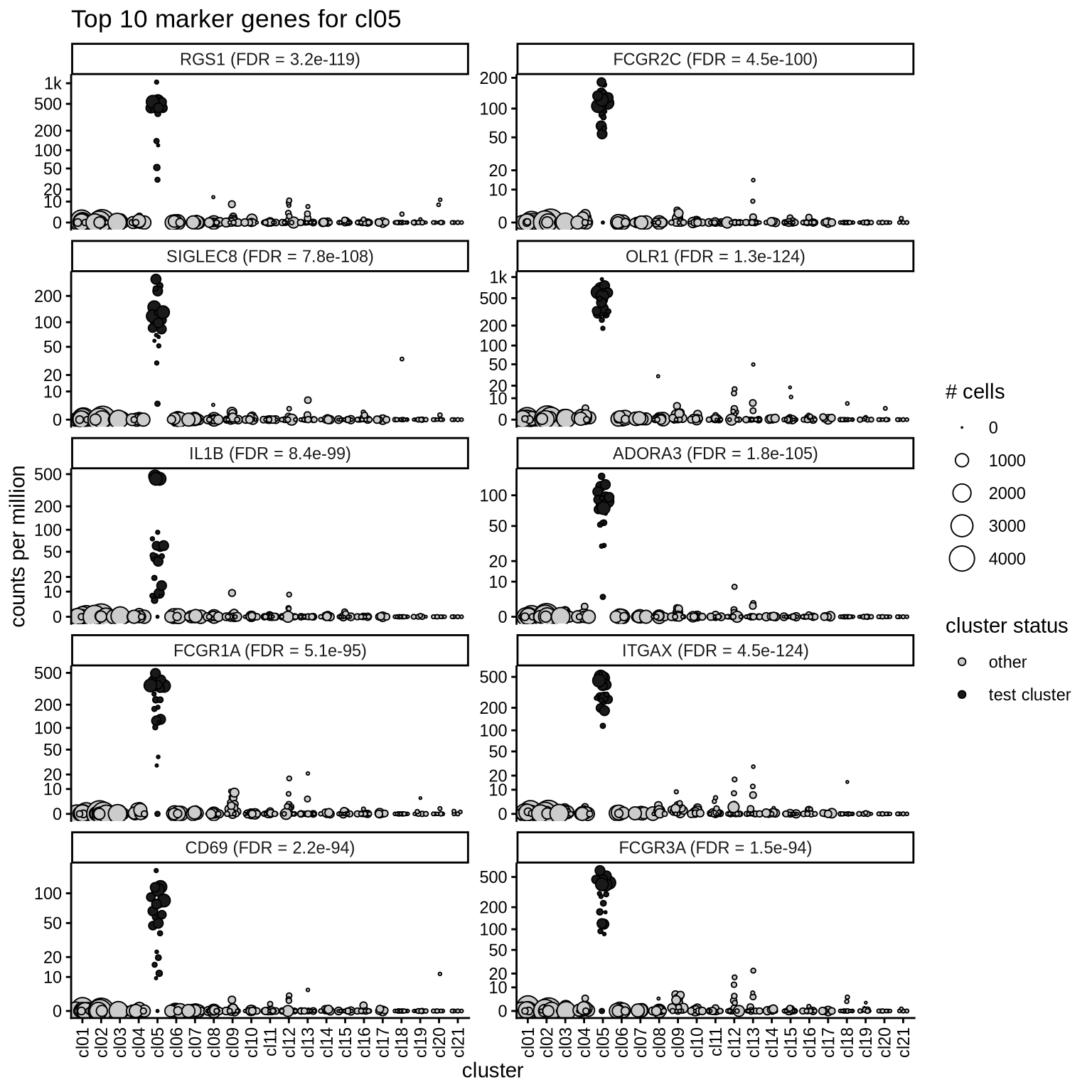
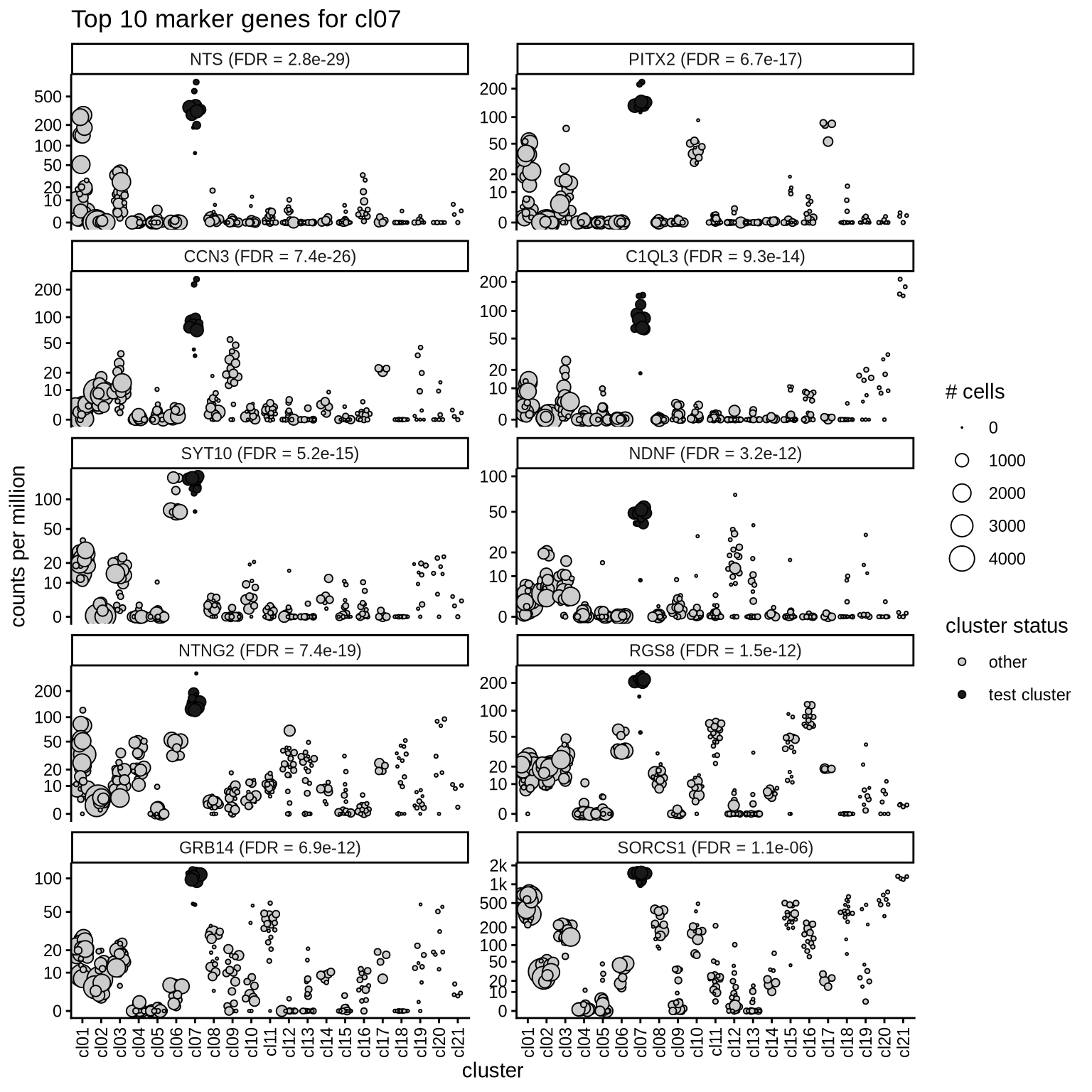
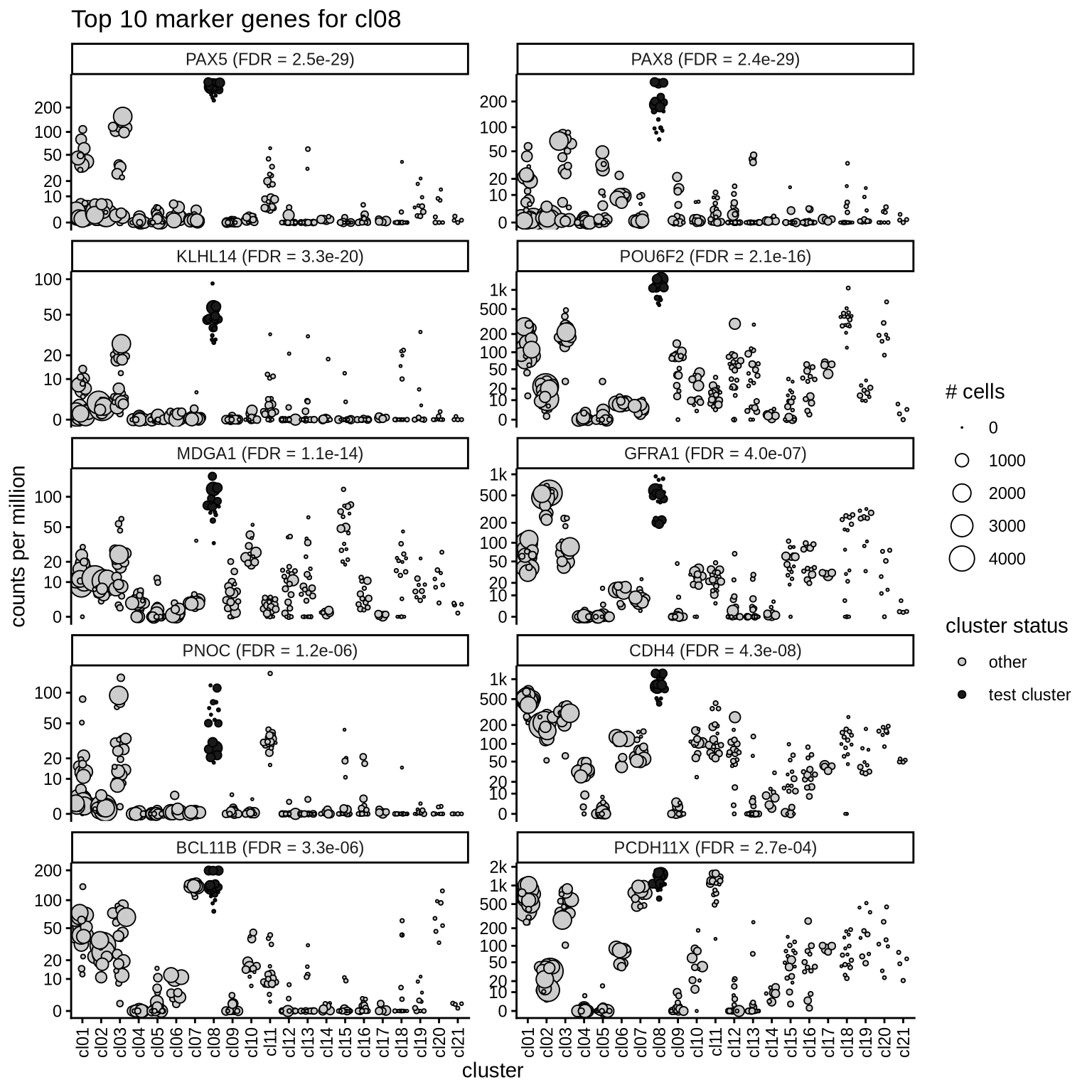
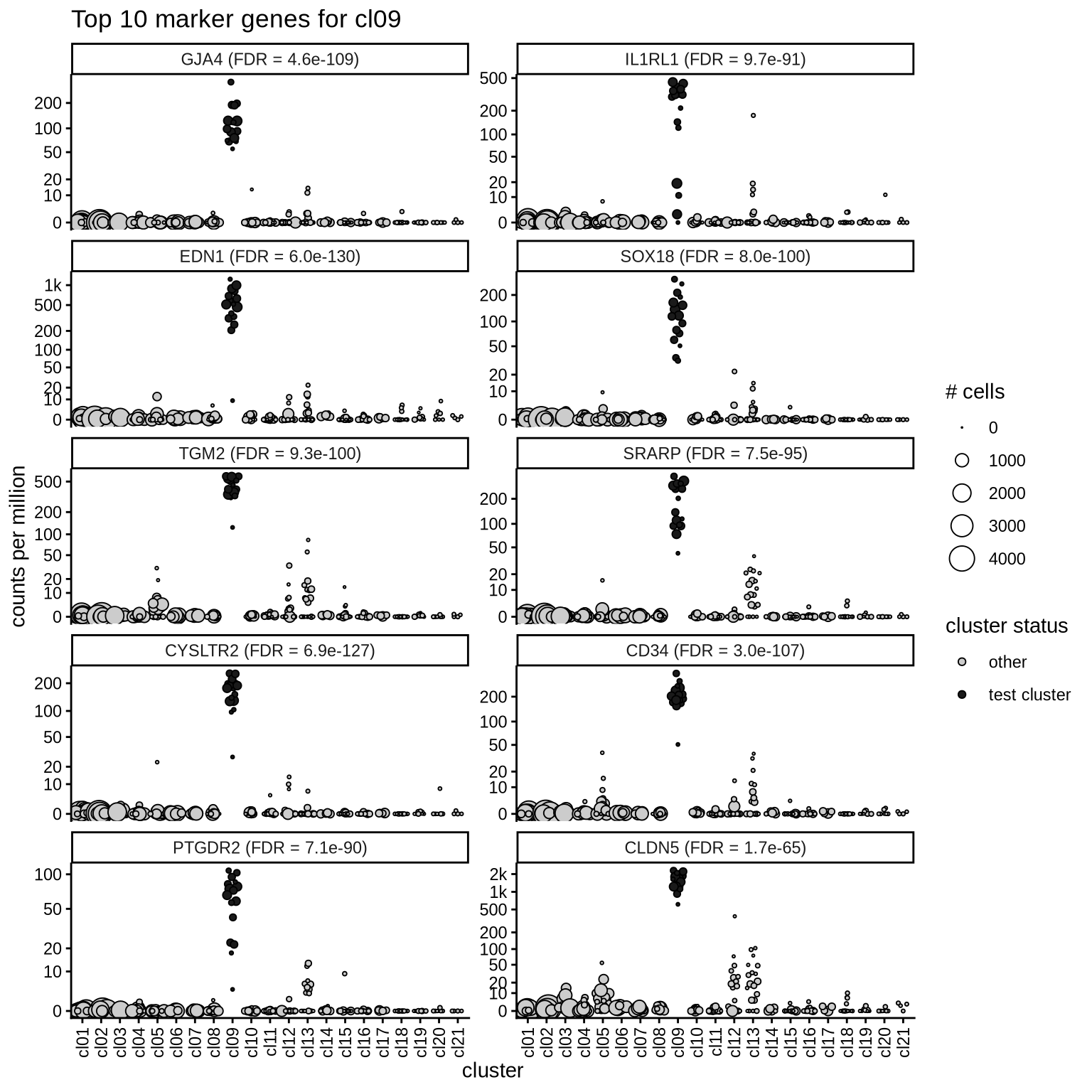
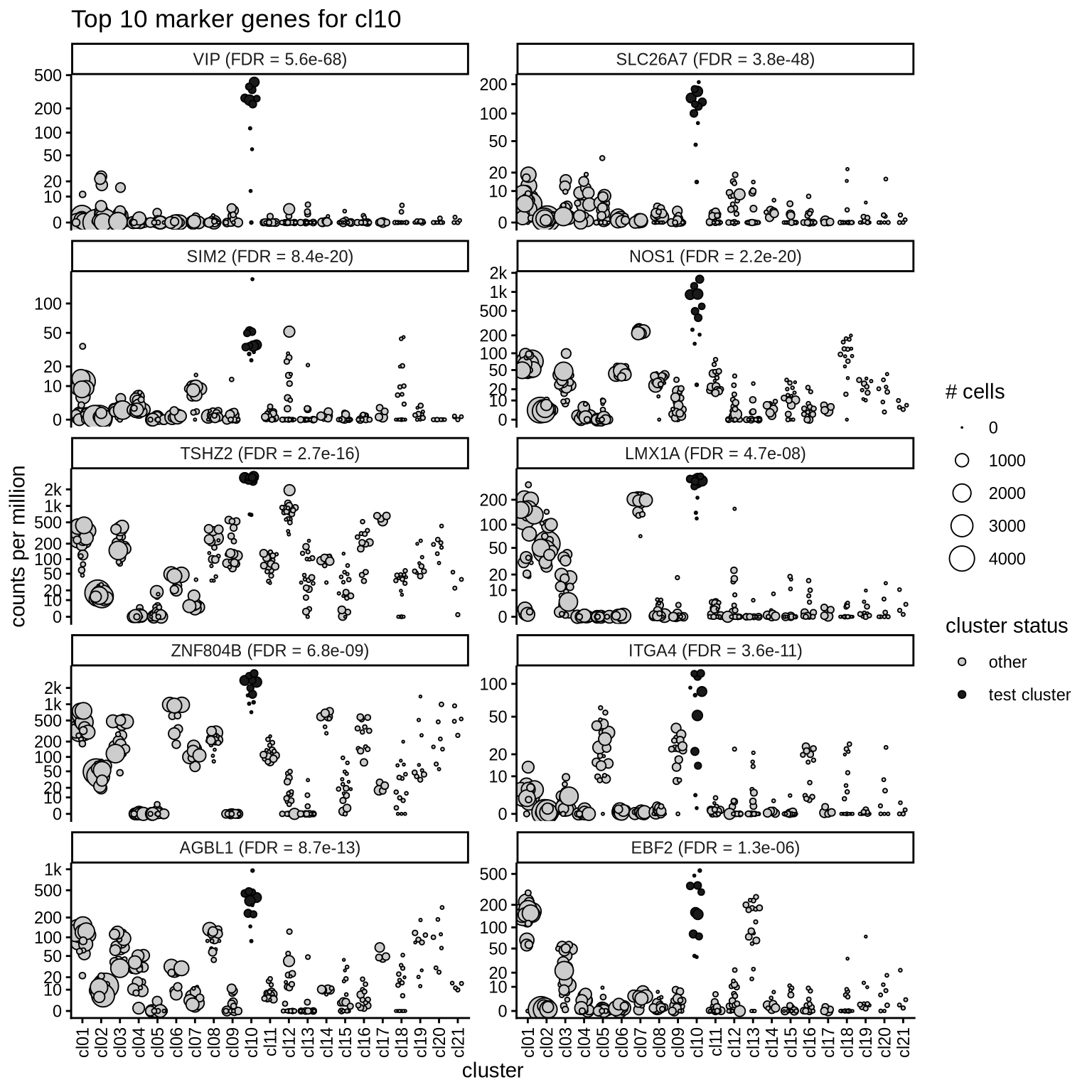
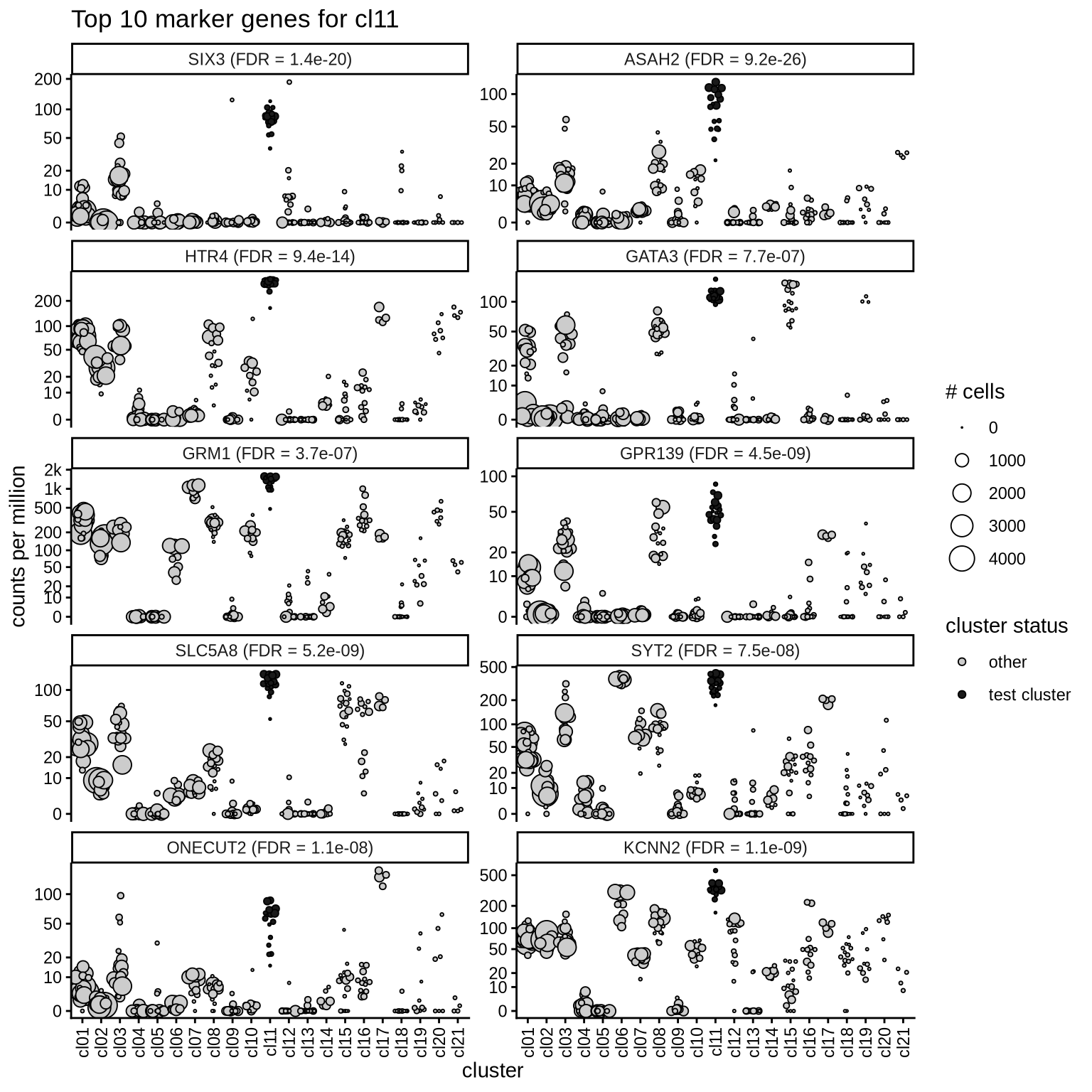
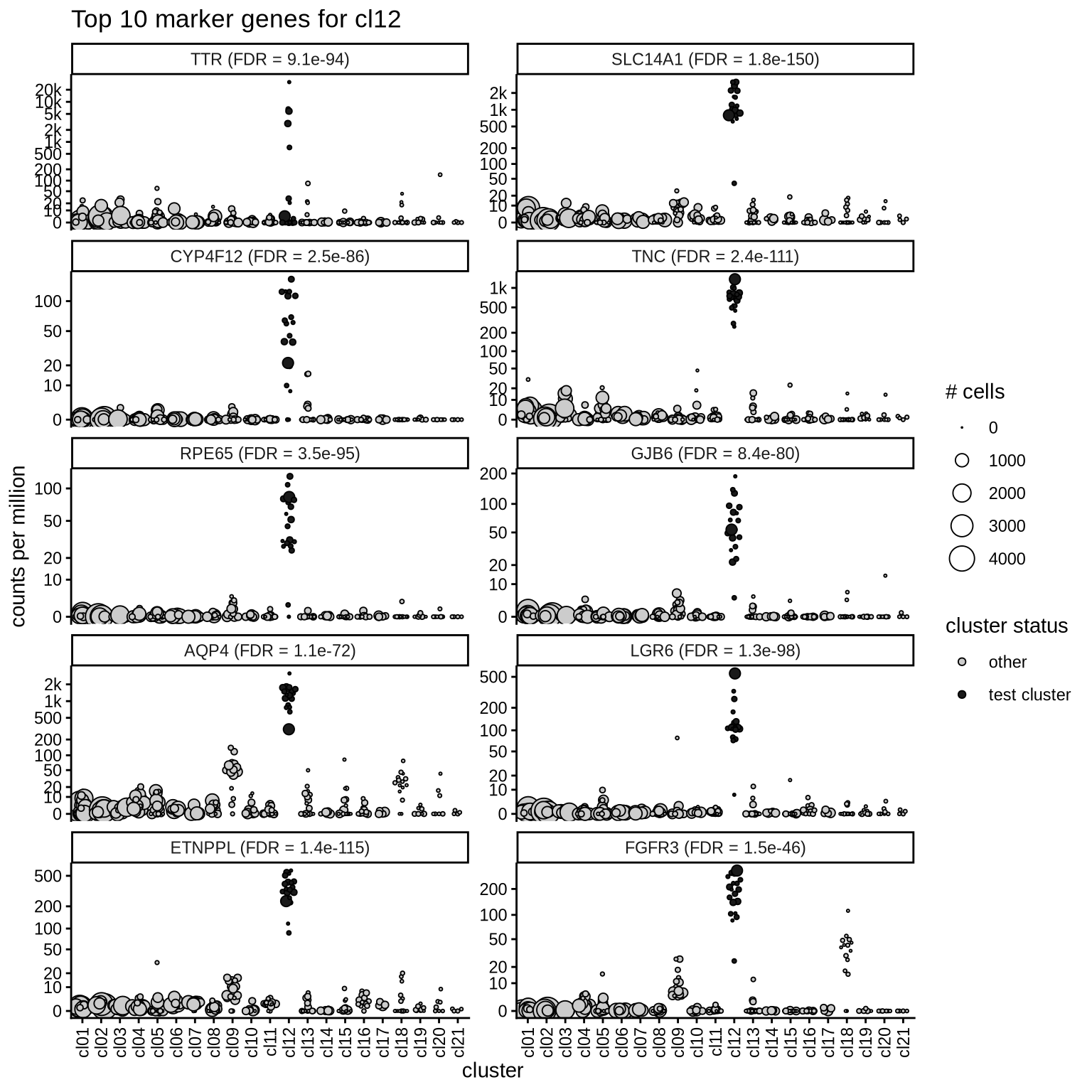
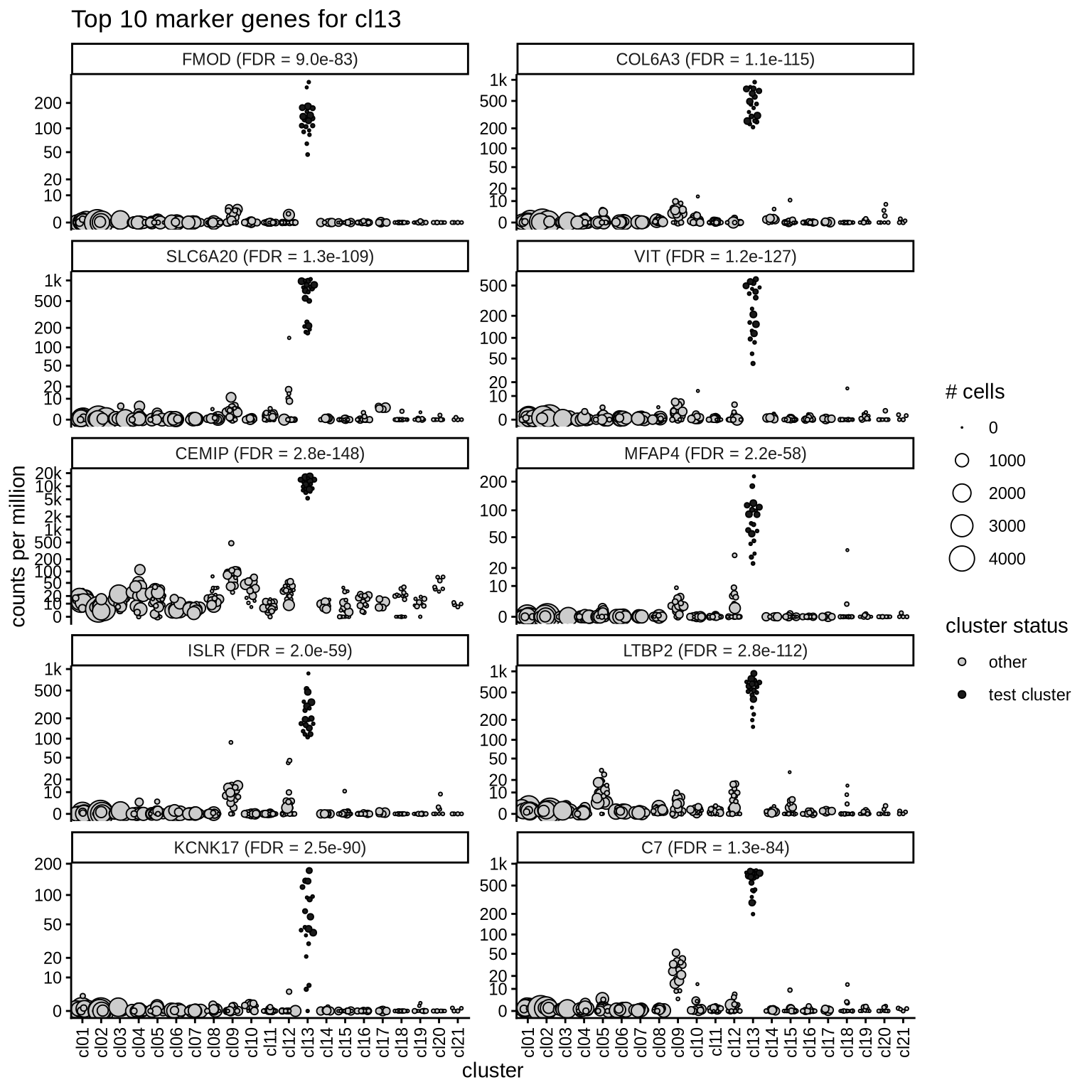
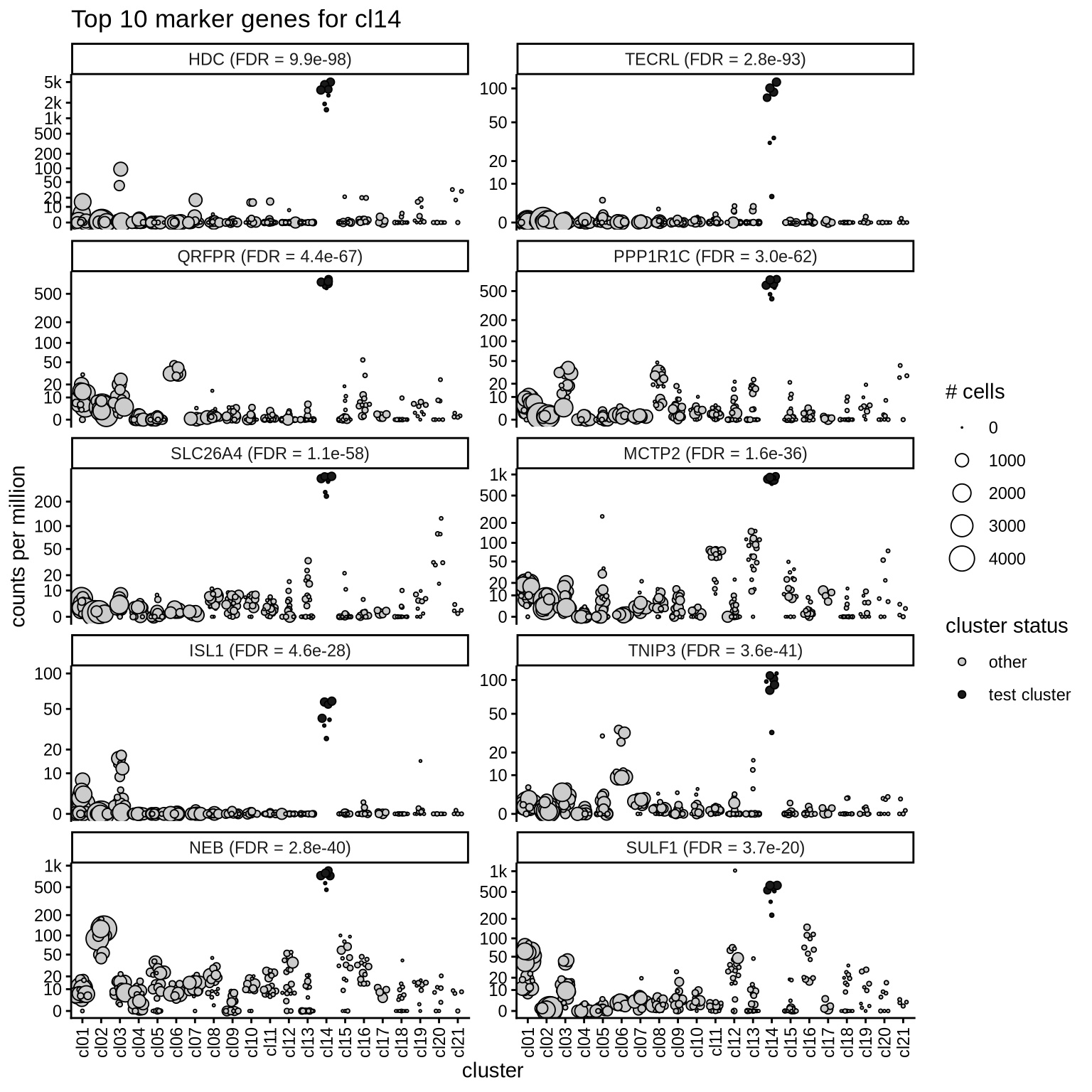
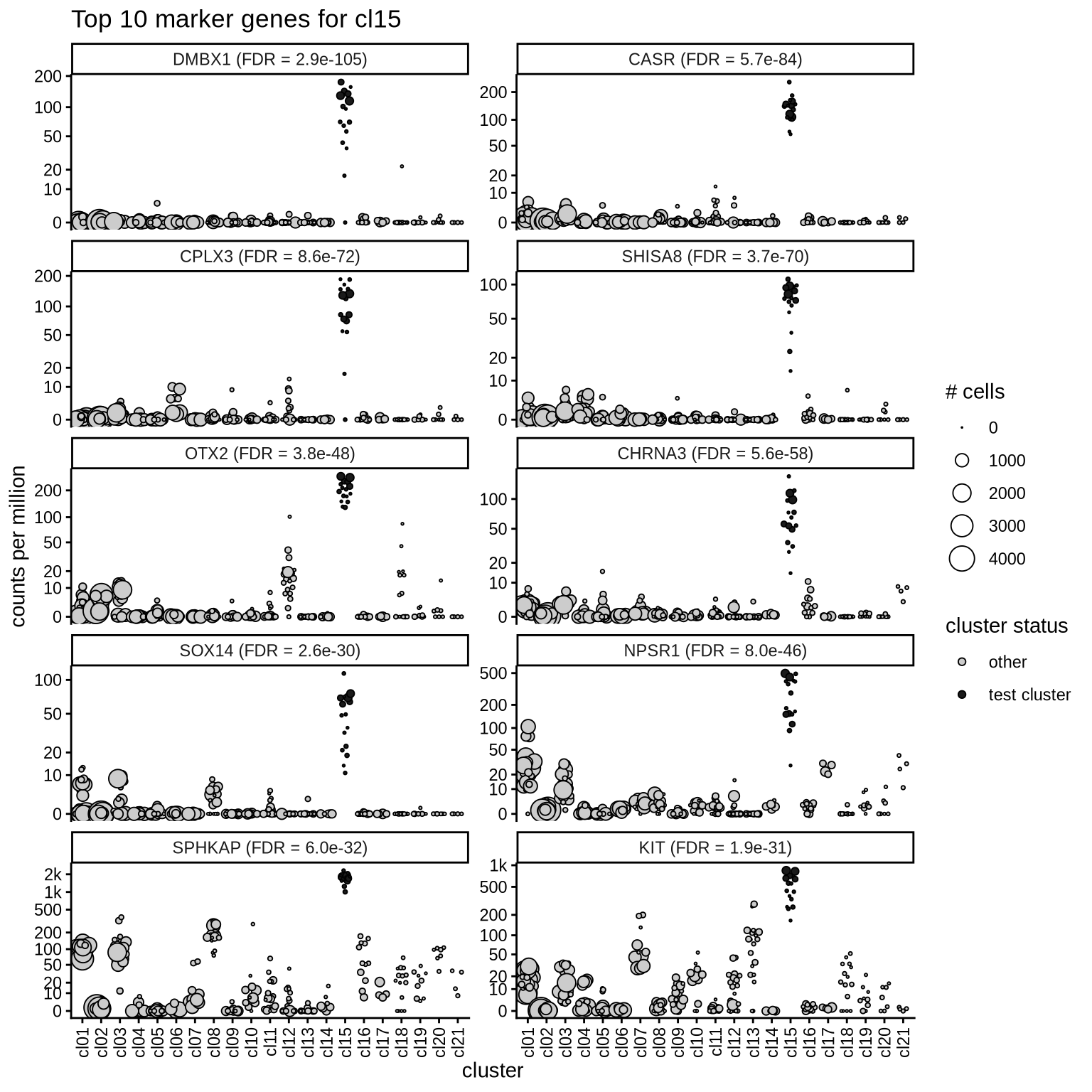
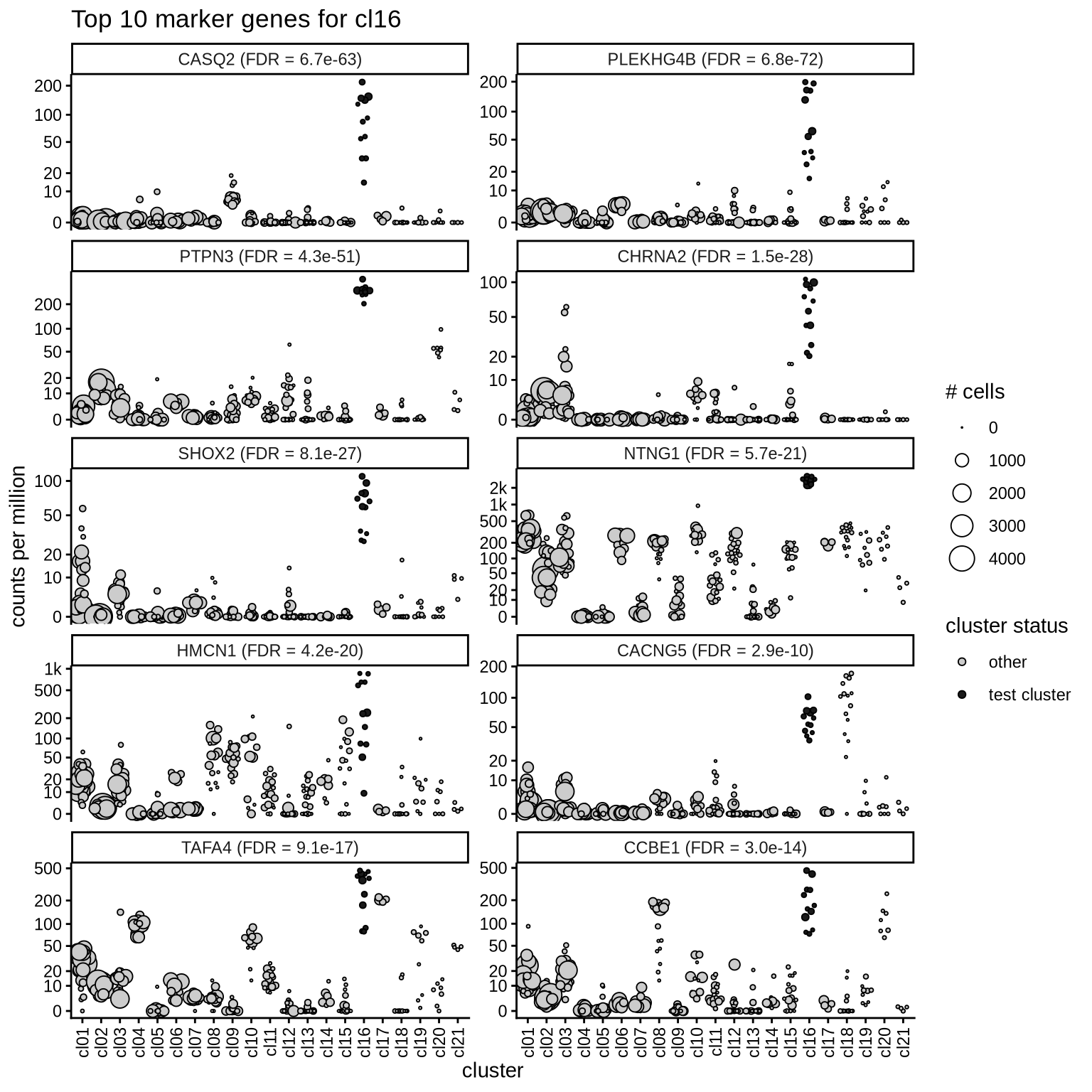
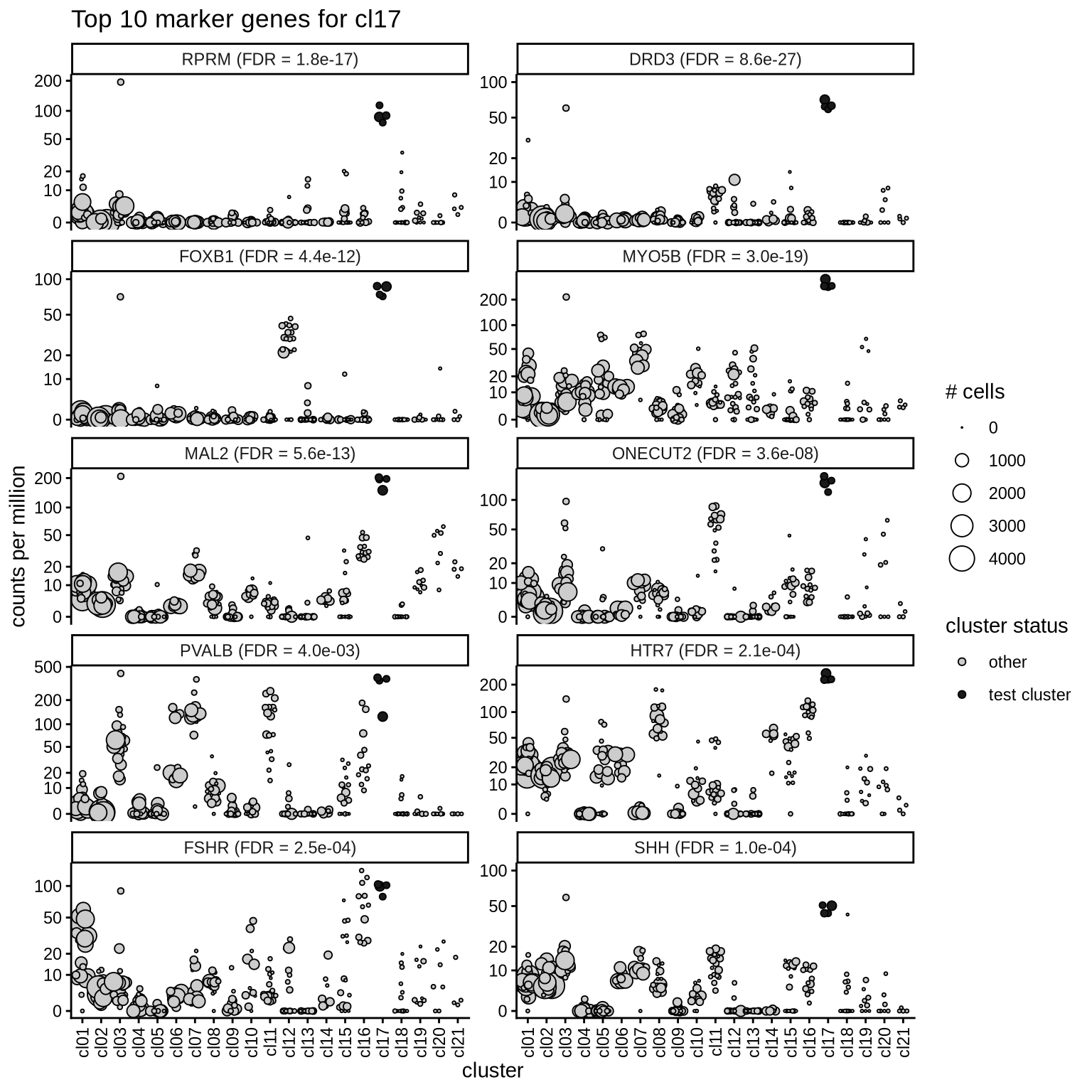
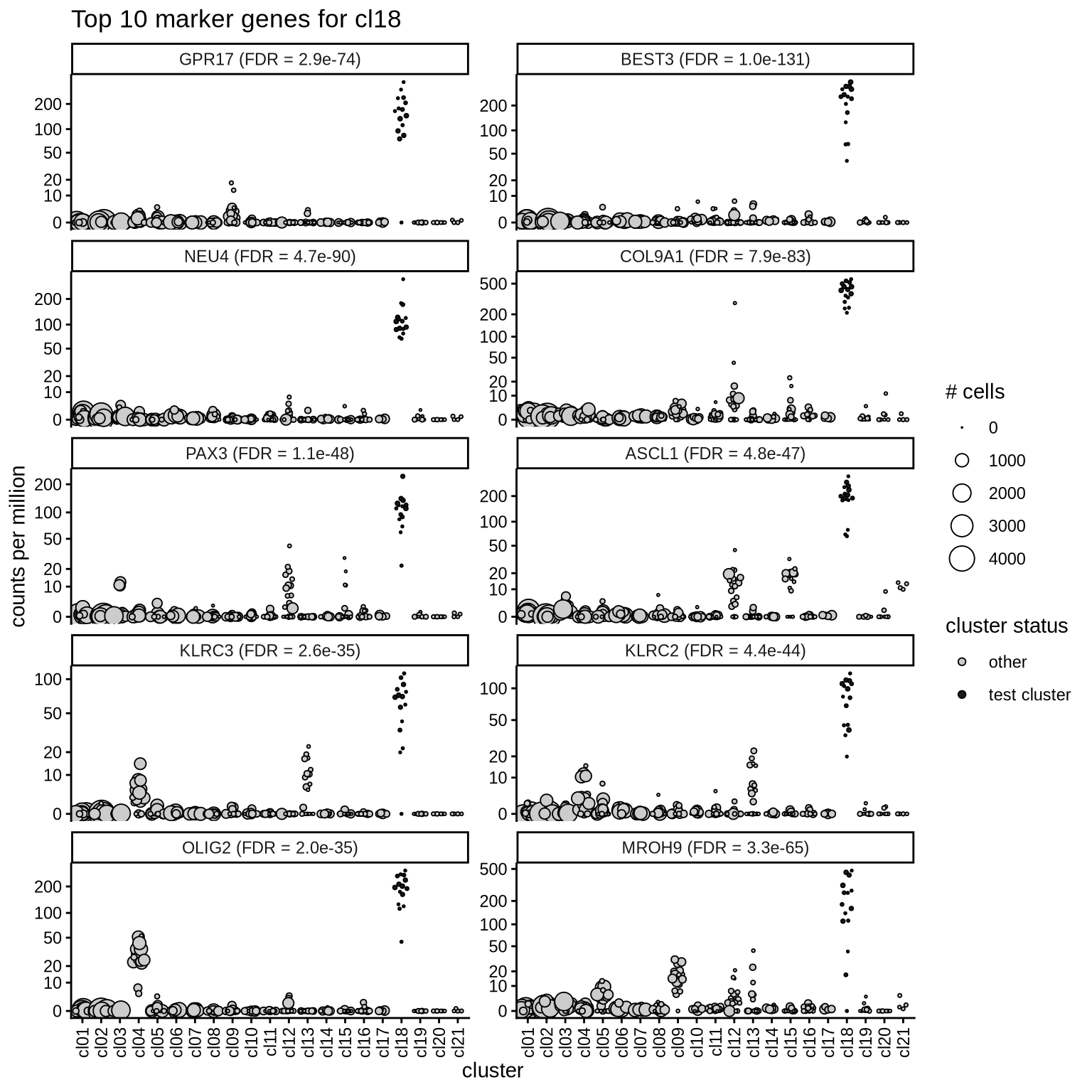
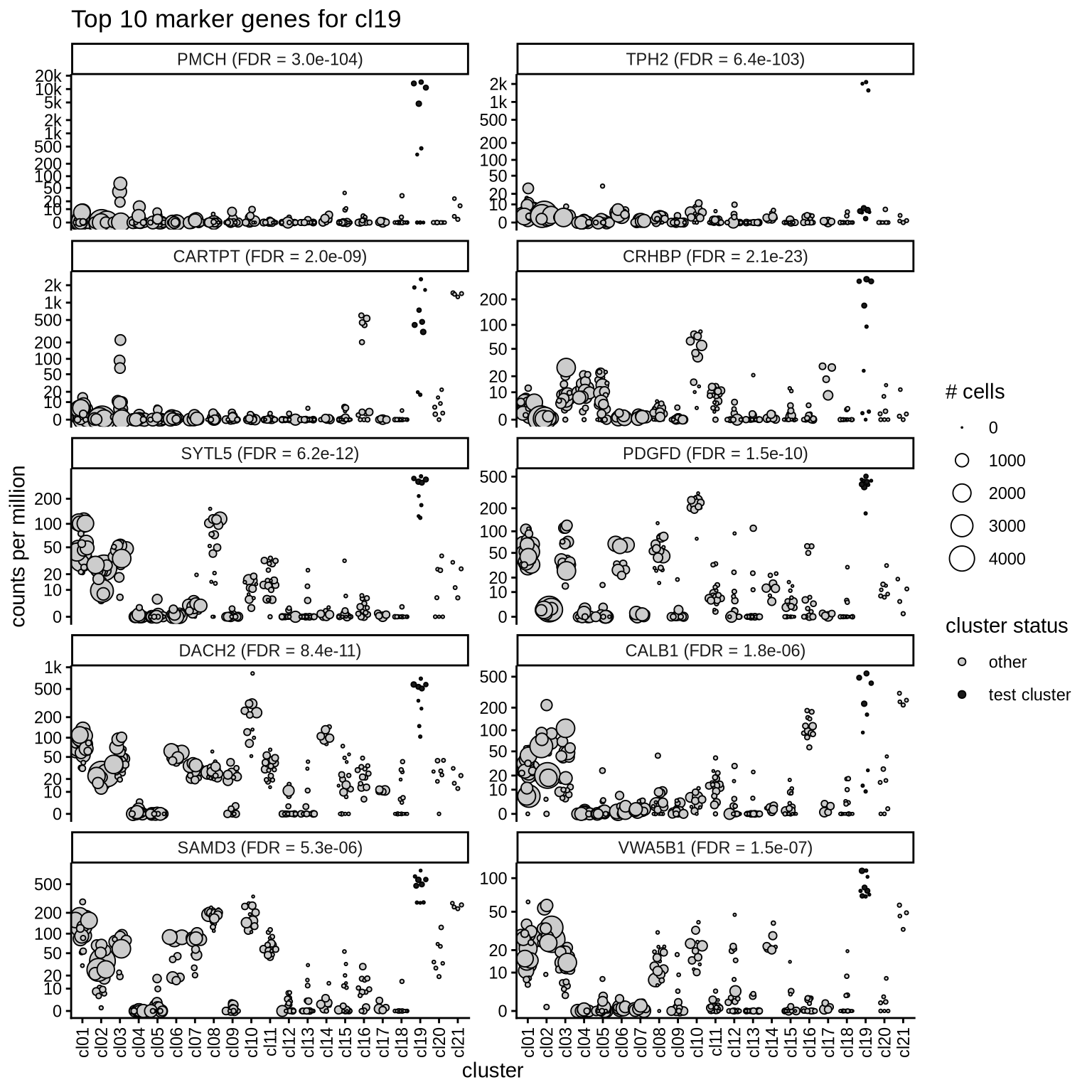
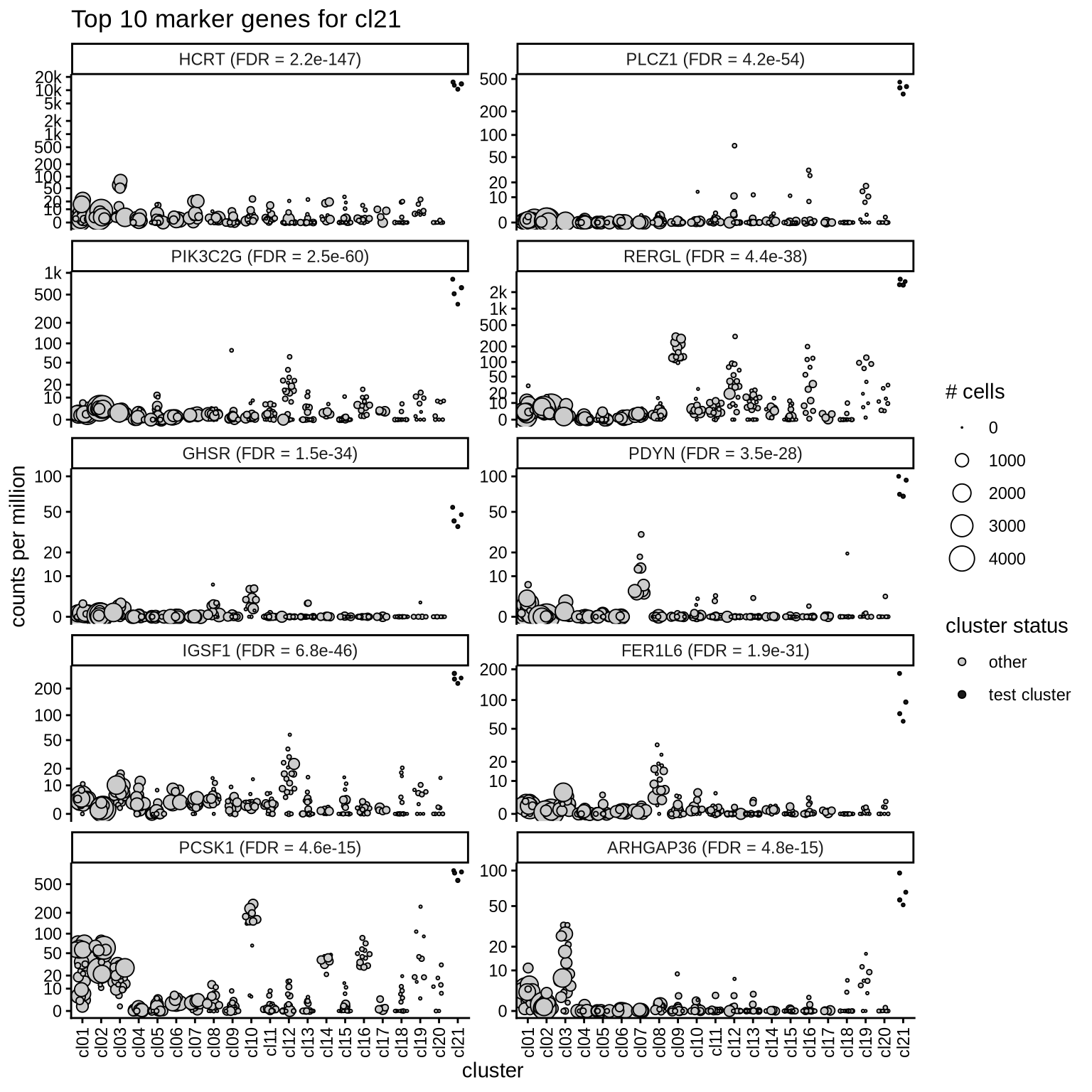
Top marker genes
Normalized pseudobulk expression values of top 10 marker genes with FDR < 0.05 and a minimum expression of 50 CPM are shown for each cluster.
for (sel_cl in cl_ord) {
cat('####', sel_cl, '\n')
print(plot_top_marker_genes(sel_cl, top_min_dt, cpms_dt, cl_order = cl_ord))
cat('\n\n')
}cl01
cl02
cl03
cl04
cl05
cl06

cl07
cl08
cl09
cl10
cl11
cl12
cl13
cl14
cl15
cl16
cl17
cl18
cl19
cl20

cl21
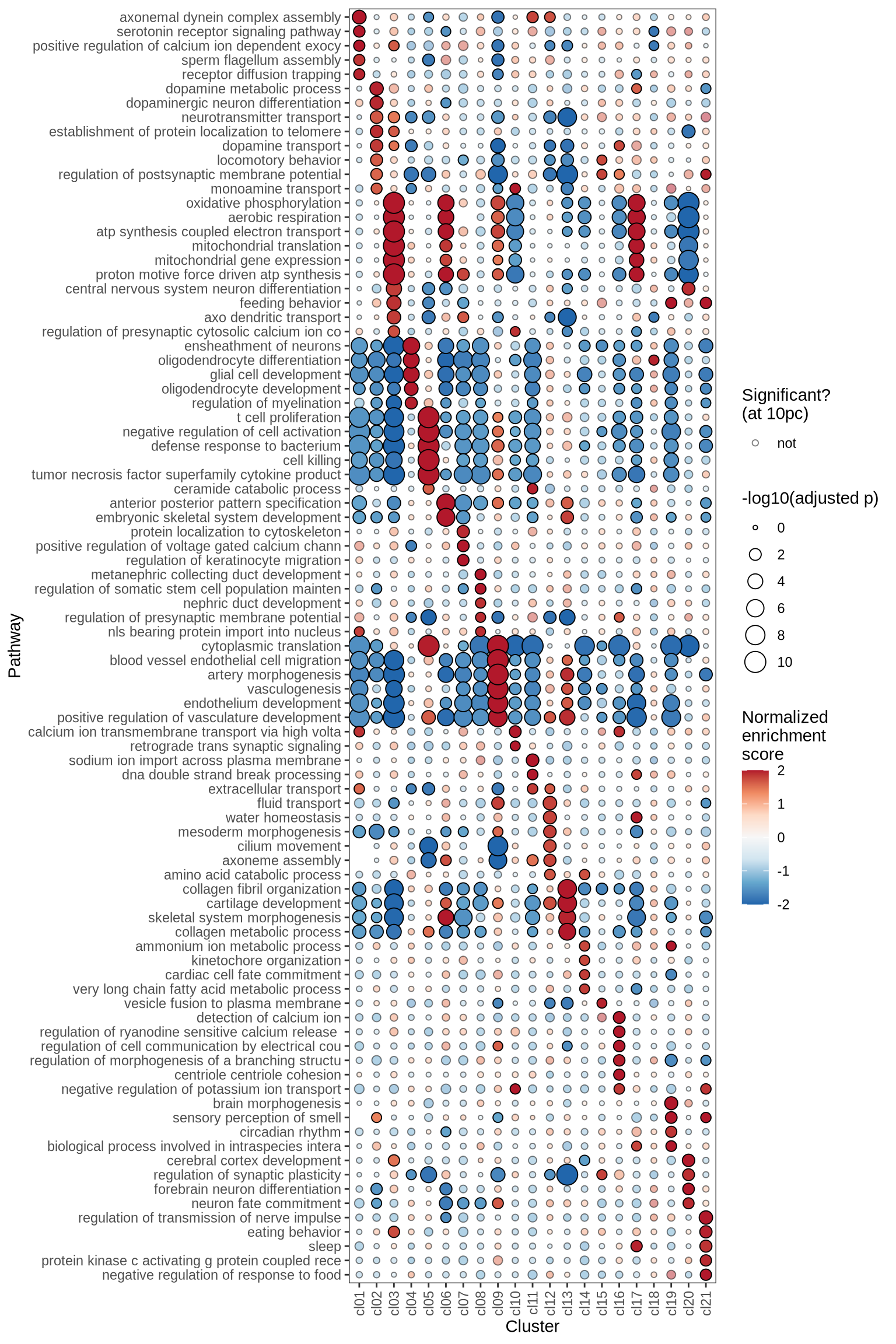
GSEA characterisation of clusters
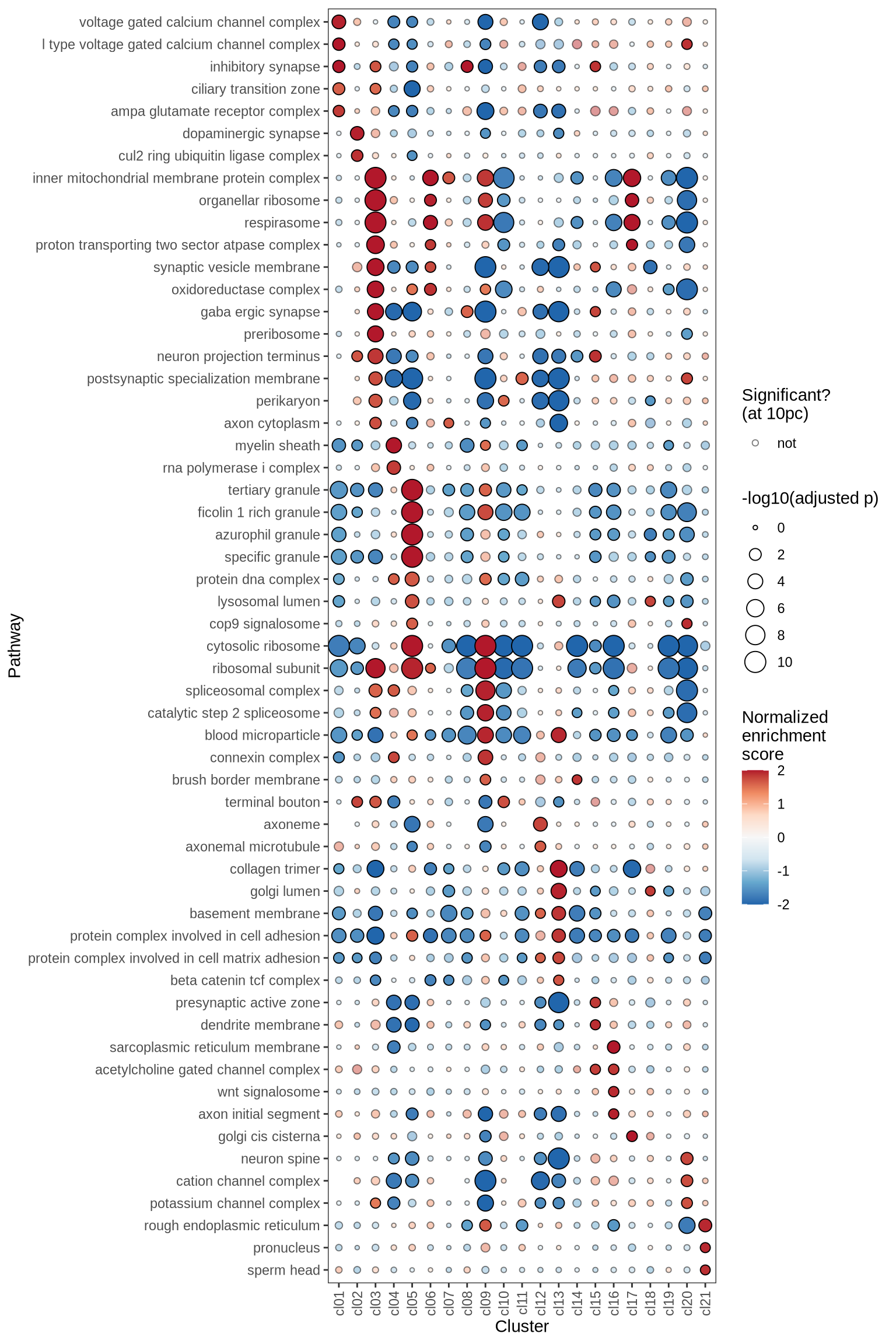
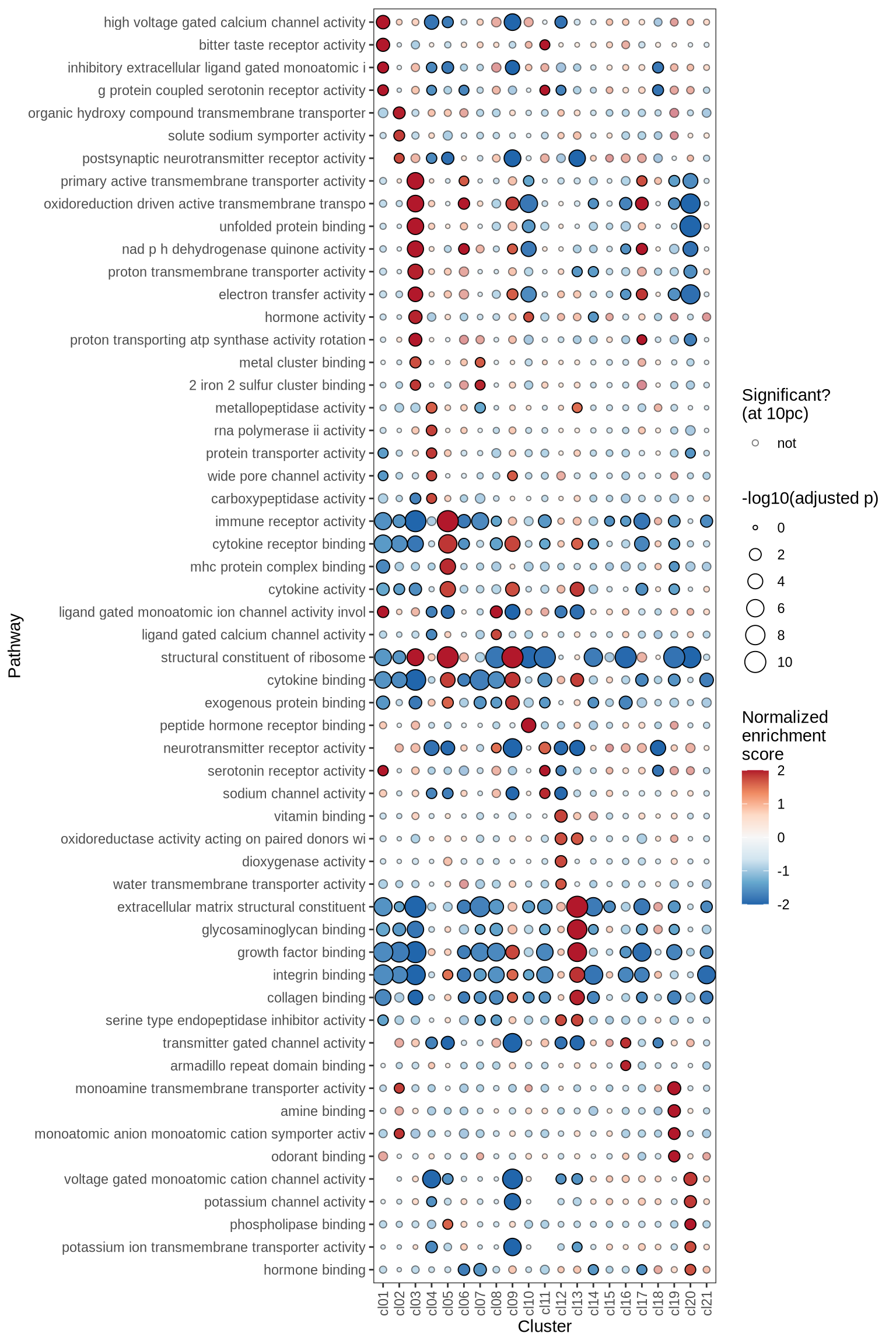
Gene Set Enrichment Analysis (GSEA) was performed on marker genes for each cluster, using log fold change as the ranking variable. The top 10 pathways, grouped into five categories and selected based on a significance threshold of 0.05, are displayed for each cluster.
for (p in names(gsea_ls)) {
if (is.null(gsea_ls[[p]]))
next
cat('### ', p, '\n')
dt = gsea_ls[[p]] %>% .[ gsea_var == gsea_var_sel ]
plot_gsea_dotplot(dt, gsea_cut = gsea_cut, n_top_paths = 5, max_nes = 2,
size_range = c(10, 200), what = "pos_only", cl_order = cl_ord) %>% print
cat('\n\n')
}go_bp
go_cc
go_mf
R session info
Details of the R package versions used are given below.
devtools::session_info()## ─ Session info ───────────────────────────────────────────────────────────────
## setting value
## version R version 4.4.3 (2025-02-28)
## os Red Hat Enterprise Linux 8.10 (Ootpa)
## system x86_64, linux-gnu
## ui X11
## language (EN)
## collate en_US.UTF-8
## ctype en_US.UTF-8
## tz Europe/Zurich
## date 2026-03-25
## pandoc 3.8.2.1 @ /home/macnairw/packages/scprocess/.snakemake/conda/4fef11cadd34f9d2d13a0d6139d09340_/bin/ (via rmarkdown)
## quarto NA
##
## ─ Packages ───────────────────────────────────────────────────────────────────
## package * version date (UTC) lib source
## abind 1.4-8 2024-09-12 [1] CRAN (R 4.4.3)
## assertthat * 0.2.1 2019-03-21 [1] CRAN (R 4.4.3)
## basilisk 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## basilisk.utils 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## beachmat 2.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## beeswarm 0.4.0 2021-06-01 [1] CRAN (R 4.4.3)
## Biobase * 2.66.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocGenerics * 0.52.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocManager 1.30.27 2025-11-14 [1] CRAN (R 4.4.3)
## BiocNeighbors 2.0.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## BiocParallel * 1.40.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocSingular 1.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## BiocStyle * 2.34.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## bluster 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## bookdown 0.45 2025-10-03 [1] CRAN (R 4.4.3)
## bslib 0.9.0 2025-01-30 [1] CRAN (R 4.4.3)
## ca 0.71.1 2020-01-24 [1] CRAN (R 4.4.3)
## cachem 1.1.0 2024-05-16 [1] CRAN (R 4.4.3)
## Cairo 1.7-0 2025-10-29 [1] CRAN (R 4.4.3)
## callr 3.7.6 2024-03-25 [1] CRAN (R 4.4.3)
## cellranger 1.1.0 2016-07-27 [1] CRAN (R 4.4.3)
## circlize * 0.4.16 2024-02-20 [1] CRAN (R 4.4.3)
## cli 3.6.5 2025-04-23 [1] CRAN (R 4.4.3)
## clue 0.3-66 2024-11-13 [1] CRAN (R 4.4.3)
## cluster 2.1.8.1 2025-03-12 [1] CRAN (R 4.4.3)
## codetools 0.2-20 2024-03-31 [1] CRAN (R 4.4.3)
## colorspace 2.1-2 2025-09-22 [1] CRAN (R 4.4.3)
## ComplexHeatmap * 2.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## cowplot 1.2.0 2025-07-07 [1] CRAN (R 4.4.3)
## crayon 1.5.3 2024-06-20 [1] CRAN (R 4.4.3)
## data.table * 1.17.8 2025-07-10 [1] CRAN (R 4.4.3)
## DelayedArray 0.32.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## deldir 2.0-4 2024-02-28 [1] CRAN (R 4.4.3)
## DESeq2 * 1.46.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## devtools 2.4.6 2025-10-03 [1] CRAN (R 4.4.3)
## digest 0.6.39 2025-11-19 [1] CRAN (R 4.4.3)
## dir.expiry 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## doParallel 1.0.17 2022-02-07 [1] CRAN (R 4.4.3)
## dotCall64 1.2 2024-10-04 [1] CRAN (R 4.4.3)
## dplyr 1.1.4 2023-11-17 [1] CRAN (R 4.4.3)
## dqrng 0.3.2 2023-11-29 [1] CRAN (R 4.4.3)
## edgeR * 4.4.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## ellipsis 0.3.2 2021-04-29 [1] CRAN (R 4.4.3)
## evaluate 1.0.5 2025-08-27 [1] CRAN (R 4.4.3)
## farver 2.1.2 2024-05-13 [1] CRAN (R 4.4.3)
## fastDummies 1.7.5 2025-01-20 [1] CRAN (R 4.4.3)
## fastmap 1.2.0 2024-05-15 [1] CRAN (R 4.4.3)
## fastmatch 1.1-6 2024-12-23 [1] CRAN (R 4.4.3)
## fgsea * 1.32.2 2024-12-19 [1] Bioconductor 3.20 (R 4.4.2)
## filelock 1.0.3 2023-12-11 [1] CRAN (R 4.4.3)
## fitdistrplus 1.2-4 2025-07-03 [1] CRAN (R 4.4.3)
## forcats * 1.0.1 2025-09-25 [1] CRAN (R 4.4.3)
## foreach 1.5.2 2022-02-02 [1] CRAN (R 4.4.3)
## fs 1.6.6 2025-04-12 [1] CRAN (R 4.4.3)
## future * 1.68.0 2025-11-17 [1] CRAN (R 4.4.3)
## future.apply 1.20.0 2025-06-06 [1] CRAN (R 4.4.3)
## generics 0.1.4 2025-05-09 [1] CRAN (R 4.4.3)
## GenomeInfoDb * 1.42.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## GenomeInfoDbData 1.2.13 2026-03-05 [1] Bioconductor
## GenomicRanges * 1.58.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## GetoptLong 1.0.5 2020-12-15 [1] CRAN (R 4.4.3)
## getPass 0.2-4 2023-12-10 [1] CRAN (R 4.4.3)
## ggbeeswarm * 0.7.2 2023-04-29 [1] CRAN (R 4.4.3)
## ggh4x * 0.3.1 2025-05-30 [1] CRAN (R 4.4.3)
## ggplot.multistats * 1.0.1 2024-09-25 [1] CRAN (R 4.4.3)
## ggplot2 * 4.0.1 2025-11-14 [1] CRAN (R 4.4.3)
## ggrepel * 0.9.6 2024-09-07 [1] CRAN (R 4.4.3)
## ggridges 0.5.7 2025-08-27 [1] CRAN (R 4.4.3)
## git2r 0.35.0 2024-10-20 [1] CRAN (R 4.4.3)
## GlobalOptions 0.1.2 2020-06-10 [1] CRAN (R 4.4.3)
## globals 0.18.0 2025-05-08 [1] CRAN (R 4.4.3)
## glue 1.8.0 2024-09-30 [1] CRAN (R 4.4.3)
## goftest 1.2-3 2021-10-07 [1] CRAN (R 4.4.3)
## gridExtra 2.3 2017-09-09 [1] CRAN (R 4.4.3)
## gtable 0.3.6 2024-10-25 [1] CRAN (R 4.4.3)
## harmony * 1.2.4 2025-10-10 [1] CRAN (R 4.4.3)
## hexbin 1.28.5 2024-11-13 [1] CRAN (R 4.4.3)
## htmltools 0.5.8.1 2024-04-04 [1] CRAN (R 4.4.3)
## htmlwidgets 1.6.4 2023-12-06 [1] CRAN (R 4.4.3)
## httpuv 1.6.16 2025-04-16 [1] CRAN (R 4.4.3)
## httr 1.4.7 2023-08-15 [1] CRAN (R 4.4.3)
## ica 1.0-3 2022-07-08 [1] CRAN (R 4.4.3)
## igraph 2.1.4 2025-01-23 [1] CRAN (R 4.4.3)
## IRanges * 2.40.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## irlba 2.3.5.1 2022-10-03 [1] CRAN (R 4.4.3)
## iterators 1.0.14 2022-02-05 [1] CRAN (R 4.4.3)
## jquerylib 0.1.4 2021-04-26 [1] CRAN (R 4.4.3)
## jsonlite 2.0.0 2025-03-27 [1] CRAN (R 4.4.3)
## KernSmooth 2.23-26 2025-01-01 [1] CRAN (R 4.4.3)
## knitr 1.50 2025-03-16 [1] CRAN (R 4.4.3)
## labeling 0.4.3 2023-08-29 [1] CRAN (R 4.4.3)
## later 1.4.4 2025-08-27 [1] CRAN (R 4.4.3)
## lattice 0.22-7 2025-04-02 [1] CRAN (R 4.4.3)
## lazyeval 0.2.2 2019-03-15 [1] CRAN (R 4.4.3)
## lifecycle 1.0.4 2023-11-07 [1] CRAN (R 4.4.3)
## limma * 3.62.1 2024-11-03 [1] Bioconductor 3.20 (R 4.4.2)
## listenv 0.10.0 2025-11-02 [1] CRAN (R 4.4.3)
## lmtest 0.9-40 2022-03-21 [1] CRAN (R 4.4.3)
## locfit 1.5-9.12 2025-03-05 [1] CRAN (R 4.4.3)
## magrittr * 2.0.4 2025-09-12 [1] CRAN (R 4.4.3)
## MASS 7.3-65 2025-02-28 [1] CRAN (R 4.4.3)
## Matrix * 1.7-4 2025-08-28 [1] CRAN (R 4.4.3)
## MatrixGenerics * 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## matrixStats * 1.5.0 2025-01-07 [1] CRAN (R 4.4.3)
## memoise 2.0.1 2021-11-26 [1] CRAN (R 4.4.3)
## metapod 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## MetBrewer 0.2.0 2022-03-21 [1] CRAN (R 4.4.3)
## mime 0.13 2025-03-17 [1] CRAN (R 4.4.3)
## miniUI 0.1.2 2025-04-17 [1] CRAN (R 4.4.3)
## nlme 3.1-168 2025-03-31 [1] CRAN (R 4.4.3)
## otel 0.2.0 2025-08-29 [1] CRAN (R 4.4.3)
## parallelly 1.45.1 2025-07-24 [1] CRAN (R 4.4.3)
## patchwork * 1.3.2 2025-08-25 [1] CRAN (R 4.4.3)
## pbapply 1.7-4 2025-07-20 [1] CRAN (R 4.4.3)
## pillar 1.11.1 2025-09-17 [1] CRAN (R 4.4.3)
## pkgbuild 1.4.8 2025-05-26 [1] CRAN (R 4.4.3)
## pkgconfig 2.0.3 2019-09-22 [1] CRAN (R 4.4.3)
## pkgload 1.4.1 2025-09-23 [1] CRAN (R 4.4.3)
## plotly 4.11.0 2025-06-19 [1] CRAN (R 4.4.3)
## plyr 1.8.9 2023-10-02 [1] CRAN (R 4.4.3)
## png 0.1-8 2022-11-29 [1] CRAN (R 4.4.3)
## polyclip 1.10-7 2024-07-23 [1] CRAN (R 4.4.3)
## processx 3.8.6 2025-02-21 [1] CRAN (R 4.4.3)
## progressr 0.18.0 2025-11-06 [1] CRAN (R 4.4.3)
## promises 1.5.0 2025-11-01 [1] CRAN (R 4.4.3)
## ps 1.9.1 2025-04-12 [1] CRAN (R 4.4.3)
## purrr 1.2.0 2025-11-04 [1] CRAN (R 4.4.3)
## R.methodsS3 1.8.2 2022-06-13 [1] CRAN (R 4.4.3)
## R.oo 1.27.1 2025-05-02 [1] CRAN (R 4.4.3)
## R.utils 2.13.0 2025-02-24 [1] CRAN (R 4.4.3)
## R6 2.6.1 2025-02-15 [1] CRAN (R 4.4.3)
## RANN 2.6.2 2024-08-25 [1] CRAN (R 4.4.3)
## RColorBrewer * 1.1-3 2022-04-03 [1] CRAN (R 4.4.3)
## Rcpp * 1.1.0 2025-07-02 [1] CRAN (R 4.4.3)
## RcppAnnoy 0.0.22 2024-01-23 [1] CRAN (R 4.4.3)
## RcppHNSW 0.6.0 2024-02-04 [1] CRAN (R 4.4.3)
## readxl * 1.4.5 2025-03-07 [1] CRAN (R 4.4.3)
## registry 0.5-1 2019-03-05 [1] CRAN (R 4.4.3)
## remotes 2.5.0 2024-03-17 [1] CRAN (R 4.4.3)
## reshape2 1.4.5 2025-11-12 [1] CRAN (R 4.4.3)
## reticulate 1.44.1 2025-11-14 [1] CRAN (R 4.4.3)
## rhdf5 * 2.50.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## rhdf5filters 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## Rhdf5lib 1.28.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## RhpcBLASctl 0.23-42 2023-02-11 [1] CRAN (R 4.4.3)
## rjson 0.2.23 2024-09-16 [1] CRAN (R 4.4.3)
## rlang 1.1.6 2025-04-11 [1] CRAN (R 4.4.3)
## rmarkdown 2.30 2025-09-28 [1] CRAN (R 4.4.3)
## rmdformats 1.0.4 2022-05-17 [1] CRAN (R 4.4.3)
## ROCR 1.0-11 2020-05-02 [1] CRAN (R 4.4.3)
## rprojroot 2.1.1 2025-08-26 [1] CRAN (R 4.4.3)
## RSpectra 0.16-2 2024-07-18 [1] CRAN (R 4.4.3)
## rstudioapi 0.17.1 2024-10-22 [1] CRAN (R 4.4.3)
## rsvd 1.0.5 2021-04-16 [1] CRAN (R 4.4.1)
## Rtsne 0.17 2023-12-07 [1] CRAN (R 4.4.3)
## S4Arrays 1.6.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## S4Vectors * 0.44.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## S7 0.2.1 2025-11-14 [1] CRAN (R 4.4.3)
## sass 0.4.10 2025-04-11 [1] CRAN (R 4.4.3)
## ScaledMatrix 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## scales * 1.4.0 2025-04-24 [1] CRAN (R 4.4.3)
## scater * 1.34.1 2025-03-03 [1] Bioconductor 3.20 (R 4.4.2)
## scattermore 1.2 2023-06-12 [1] CRAN (R 4.4.3)
## scran * 1.34.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## sctransform 0.4.2 2025-04-30 [1] CRAN (R 4.4.3)
## scuttle * 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## seriation * 1.5.8 2025-08-20 [1] CRAN (R 4.4.3)
## sessioninfo 1.2.3 2025-02-05 [1] CRAN (R 4.4.3)
## Seurat * 5.3.1 2025-10-29 [1] CRAN (R 4.4.3)
## SeuratObject * 5.2.0 2025-08-27 [1] CRAN (R 4.4.3)
## shape 1.4.6.1 2024-02-23 [1] CRAN (R 4.4.3)
## shiny 1.11.1 2025-07-03 [1] CRAN (R 4.4.3)
## SingleCellExperiment * 1.28.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## sp * 2.2-0 2025-02-01 [1] CRAN (R 4.4.3)
## spam 2.11-1 2025-01-20 [1] CRAN (R 4.4.3)
## SparseArray 1.6.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## spatstat.data 3.1-9 2025-10-18 [1] CRAN (R 4.4.3)
## spatstat.explore 3.6-0 2025-11-22 [1] CRAN (R 4.4.3)
## spatstat.geom 3.6-1 2025-11-20 [1] CRAN (R 4.4.3)
## spatstat.random 3.4-3 2025-11-21 [1] CRAN (R 4.4.3)
## spatstat.sparse 3.1-0 2024-06-21 [1] CRAN (R 4.4.3)
## spatstat.univar 3.1-5 2025-11-17 [1] CRAN (R 4.4.3)
## spatstat.utils 3.2-0 2025-09-20 [1] CRAN (R 4.4.3)
## statmod 1.5.1 2025-10-09 [1] CRAN (R 4.4.3)
## strex * 2.0.1 2024-10-03 [1] CRAN (R 4.4.3)
## stringi * 1.8.7 2025-03-27 [1] CRAN (R 4.4.3)
## stringr * 1.6.0 2025-11-04 [1] CRAN (R 4.4.3)
## SummarizedExperiment * 1.36.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## survival 3.8-3 2024-12-17 [1] CRAN (R 4.4.3)
## tensor 1.5.1 2025-06-17 [1] CRAN (R 4.4.3)
## tibble 3.3.0 2025-06-08 [1] CRAN (R 4.4.3)
## tidyr 1.3.1 2024-01-24 [1] CRAN (R 4.4.3)
## tidyselect 1.2.1 2024-03-11 [1] CRAN (R 4.4.3)
## TSP 1.2.6 2025-11-27 [1] CRAN (R 4.4.3)
## UCSC.utils 1.2.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## usethis 3.2.1 2025-09-06 [1] CRAN (R 4.4.3)
## uwot * 0.2.4 2025-11-10 [1] CRAN (R 4.4.3)
## vctrs 0.6.5 2023-12-01 [1] CRAN (R 4.4.3)
## vipor 0.4.7 2023-12-18 [1] CRAN (R 4.4.3)
## viridis * 0.6.5 2024-01-29 [1] CRAN (R 4.4.3)
## viridisLite * 0.4.2 2023-05-02 [1] CRAN (R 4.4.3)
## whisker 0.4.1 2022-12-05 [1] CRAN (R 4.4.3)
## withr 3.0.2 2024-10-28 [1] CRAN (R 4.4.3)
## workflowr * 1.7.2 2025-08-18 [1] CRAN (R 4.4.3)
## xfun 0.54 2025-10-30 [1] CRAN (R 4.4.3)
## xtable 1.8-4 2019-04-21 [1] CRAN (R 4.4.3)
## XVector 0.46.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## yaml 2.3.11 2025-11-28 [1] CRAN (R 4.4.3)
## zellkonverter * 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## zlibbioc 1.52.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## zoo 1.8-14 2025-04-10 [1] CRAN (R 4.4.3)
##
## [1] /home/macnairw/packages/scprocess/.snakemake/conda/4fef11cadd34f9d2d13a0d6139d09340_/lib/R/library
## * ── Packages attached to the search path.
##
## ──────────────────────────────────────────────────────────────────────────────