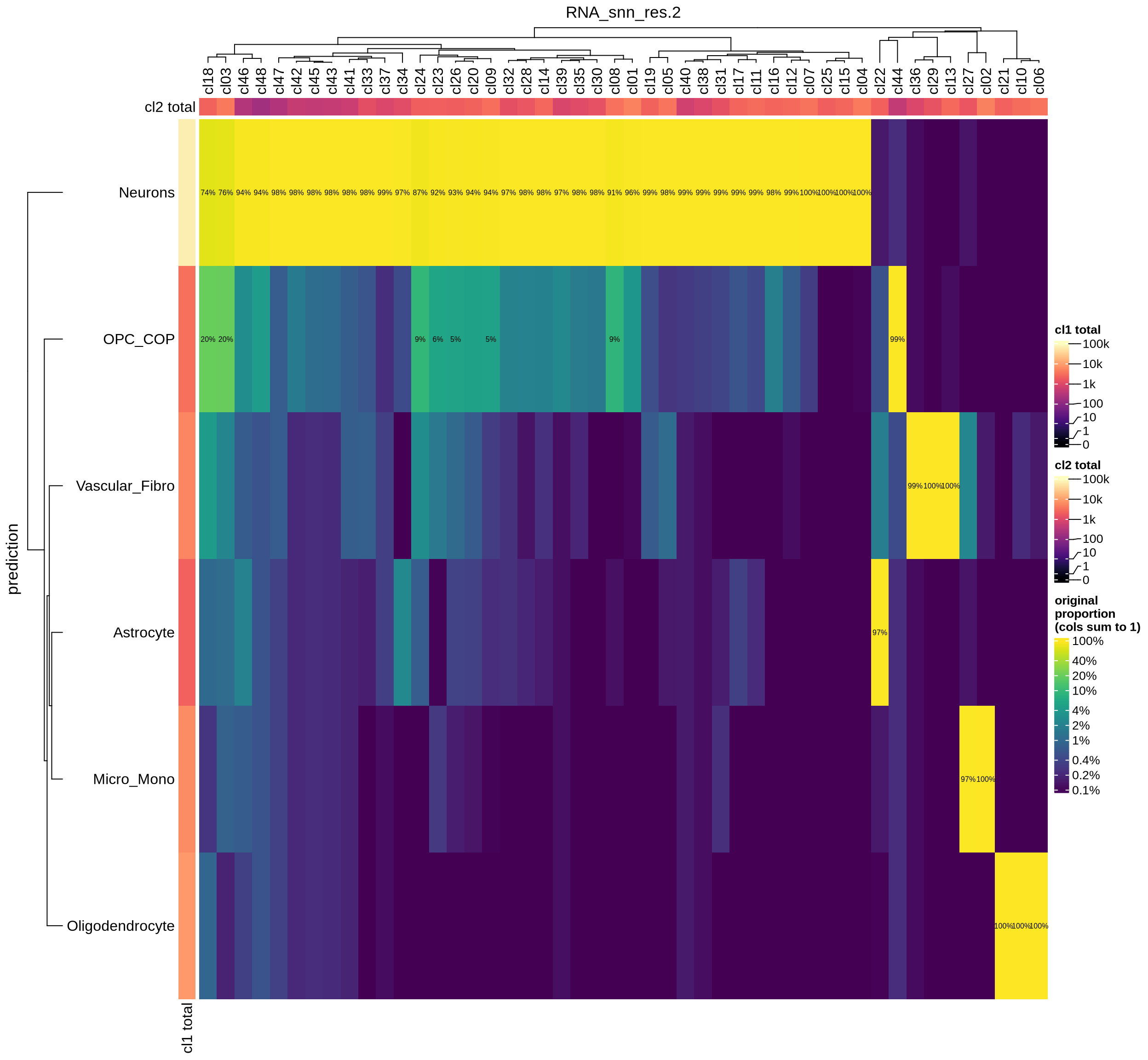
Integration clusters vs predicted cell types
Prediction of cell type labels is a two-step process: The labeller first assigns a predicted cell type to each individual cell. These cell-level predictions are then aggregated at the level of high-resolution clusters (a majority-voting approach is used, where the final, assigned label for a cluster is the most common predicted label among all the cells within it).
The heatmap helps to assess whether this aggregation step seems sensible. Each cell on the heatmap shows the proportion of cells within a cluster (on the x-axis) that are assigned a specific predicted cell type in the first step (on the y-axis).
for (ii in seq_along(guess_f_ls)) {
# unpack
this_labeller = labeller_ls[[ ii ]]
this_model = model_ls[[ ii ]]
this_hi_res_cl = hi_res_cl_ls[[ ii ]]
# make confusion matrix
guesses_sel = guesses_ls[[ ii ]] %>%
.[, .(cell_id, prediction = predicted_label_naive %>% factor)]
int_tmp = int_dt[, .(cell_id, UMAP1, UMAP2, hi_res_cl = get(this_hi_res_cl) %>% fct_infreq)]
confuse_dt = calc_confuse_dt(guesses_sel, int_tmp, "prediction", "hi_res_cl", min_cl2_p = min_cl2_p)
# plot heatmap
cat('### ', this_labeller, ', ', this_model, '\n', sep = '')
draw(plot_cluster_comparison_heatmap(confuse_dt, "prediction", this_hi_res_cl,
plot_var = 'log_p_cl2', do_sort = "hclust"), merge_legend = TRUE)
cat('\n\n')
}scprocess, human_cns
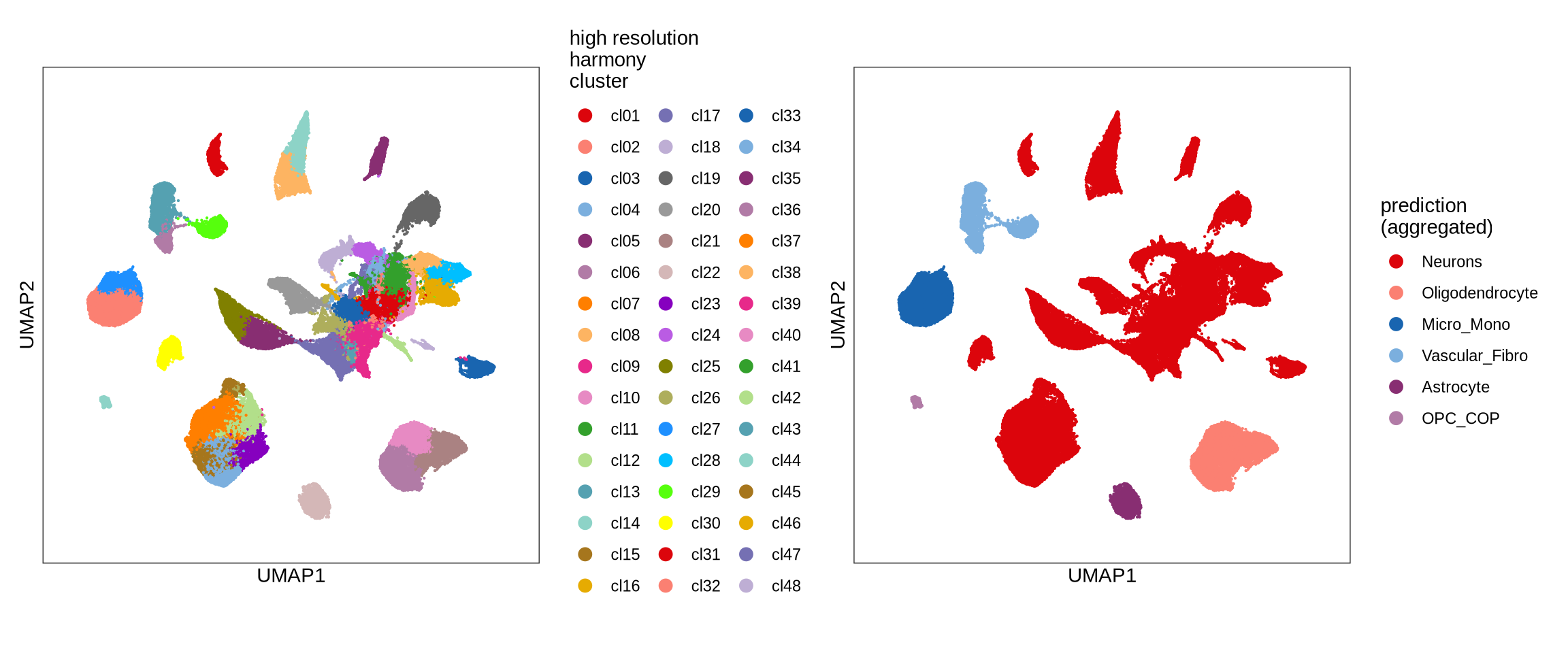
Predicted cell types over UMAP
Clustering of data is performed with different labellers and models. Each of these is shown over a UMAP together with a plot showing predicted cell type labels.
for (ii in seq_along(guess_f_ls)) {
# unpack
this_labeller = labeller_ls[[ ii ]]
this_model = model_ls[[ ii ]]
this_hi_res_cl = hi_res_cl_ls[[ ii ]]
# get data
guesses_sel = guesses_ls[[ ii ]] %>%
.[, .(cell_id, prediction = predicted_label_agg %>% factor)]
int_tmp = int_dt[, .(cell_id, UMAP1, UMAP2, hi_res_cl = get(this_hi_res_cl) %>% fct_infreq)]
# plots
g_cl = plot_umap_cluster(int_dt,
int_tmp[, .(cell_id, cluster = hi_res_cl)], "high resolution\nharmony\ncluster")
g_pred = plot_umap_cluster(int_dt,
guesses_sel[, .(cell_id, cluster = prediction)], "prediction\n(aggregated)")
g = g_cl + g_pred
# plot umaps
cat('### ', this_labeller, ', ', this_model, '\n', sep = '')
print(g)
cat('\n\n')
}scprocess, human_cns
Totals for each predicted cell type
Tables showing how many in total of each celltype are predicted for each label, for each combination of labeller and model.
for (ii in seq_along(guess_f_ls)) {
# unpack
this_labeller = labeller_ls[[ ii ]]
this_model = model_ls[[ ii ]]
# print table
cat('### ', this_labeller, ', ', this_model, '\n', sep = '')
guesses_ls[[ ii ]] %>% calc_labels_table %>% kbl("html") %>% row_spec(row = 0, bold = TRUE) %>%
kable_styling(bootstrap_options = "striped", full_width = FALSE, font_size = 12, position = "center") %>%
scroll_box(width = "60%", height = "500px") %>% print
cat('\n\n')
}scprocess, human_cns
| predicted label | no. cells, aggregated | no. cells, not aggregated |
|---|---|---|
| Neurons | 72214 | 68311 |
| Oligodendrocyte | 8566 | 8579 |
| Micro_Mono | 6407 | 6307 |
| Vascular_Fibro | 5017 | 5491 |
| Astrocyte | 2150 | 2232 |
| OPC_COP | 456 | 3200 |
| T_NK_B_cell | 0 | 562 |
| Ependymal_ChorPlex | 0 | 128 |
R session info
Details of the R package versions used are given below.
devtools::session_info()## ─ Session info ───────────────────────────────────────────────────────────────
## setting value
## version R version 4.4.3 (2025-02-28)
## os Red Hat Enterprise Linux 8.10 (Ootpa)
## system x86_64, linux-gnu
## ui X11
## language (EN)
## collate en_US.UTF-8
## ctype en_US.UTF-8
## tz Europe/Zurich
## date 2026-03-25
## pandoc 3.8.2.1 @ /home/macnairw/packages/scprocess/.snakemake/conda/4fef11cadd34f9d2d13a0d6139d09340_/bin/ (via rmarkdown)
## quarto NA
##
## ─ Packages ───────────────────────────────────────────────────────────────────
## package * version date (UTC) lib source
## abind 1.4-8 2024-09-12 [1] CRAN (R 4.4.3)
## assertthat * 0.2.1 2019-03-21 [1] CRAN (R 4.4.3)
## basilisk 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## basilisk.utils 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## beachmat 2.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## beeswarm 0.4.0 2021-06-01 [1] CRAN (R 4.4.3)
## Biobase * 2.66.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocGenerics * 0.52.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocManager 1.30.27 2025-11-14 [1] CRAN (R 4.4.3)
## BiocNeighbors 2.0.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## BiocParallel * 1.40.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocSingular 1.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## BiocStyle * 2.34.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## bluster 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## bookdown 0.45 2025-10-03 [1] CRAN (R 4.4.3)
## bslib 0.9.0 2025-01-30 [1] CRAN (R 4.4.3)
## ca 0.71.1 2020-01-24 [1] CRAN (R 4.4.3)
## cachem 1.1.0 2024-05-16 [1] CRAN (R 4.4.3)
## Cairo 1.7-0 2025-10-29 [1] CRAN (R 4.4.3)
## callr 3.7.6 2024-03-25 [1] CRAN (R 4.4.3)
## cellranger 1.1.0 2016-07-27 [1] CRAN (R 4.4.3)
## circlize * 0.4.16 2024-02-20 [1] CRAN (R 4.4.3)
## cli 3.6.5 2025-04-23 [1] CRAN (R 4.4.3)
## clue 0.3-66 2024-11-13 [1] CRAN (R 4.4.3)
## cluster 2.1.8.1 2025-03-12 [1] CRAN (R 4.4.3)
## codetools 0.2-20 2024-03-31 [1] CRAN (R 4.4.3)
## colorspace 2.1-2 2025-09-22 [1] CRAN (R 4.4.3)
## ComplexHeatmap * 2.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## cowplot 1.2.0 2025-07-07 [1] CRAN (R 4.4.3)
## crayon 1.5.3 2024-06-20 [1] CRAN (R 4.4.3)
## data.table * 1.17.8 2025-07-10 [1] CRAN (R 4.4.3)
## DelayedArray 0.32.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## deldir 2.0-4 2024-02-28 [1] CRAN (R 4.4.3)
## devtools 2.4.6 2025-10-03 [1] CRAN (R 4.4.3)
## digest 0.6.39 2025-11-19 [1] CRAN (R 4.4.3)
## dir.expiry 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## doParallel 1.0.17 2022-02-07 [1] CRAN (R 4.4.3)
## dotCall64 1.2 2024-10-04 [1] CRAN (R 4.4.3)
## dplyr 1.1.4 2023-11-17 [1] CRAN (R 4.4.3)
## dqrng 0.3.2 2023-11-29 [1] CRAN (R 4.4.3)
## edgeR 4.4.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## ellipsis 0.3.2 2021-04-29 [1] CRAN (R 4.4.3)
## evaluate 1.0.5 2025-08-27 [1] CRAN (R 4.4.3)
## farver 2.1.2 2024-05-13 [1] CRAN (R 4.4.3)
## fastDummies 1.7.5 2025-01-20 [1] CRAN (R 4.4.3)
## fastmap 1.2.0 2024-05-15 [1] CRAN (R 4.4.3)
## filelock 1.0.3 2023-12-11 [1] CRAN (R 4.4.3)
## fitdistrplus 1.2-4 2025-07-03 [1] CRAN (R 4.4.3)
## forcats * 1.0.1 2025-09-25 [1] CRAN (R 4.4.3)
## foreach 1.5.2 2022-02-02 [1] CRAN (R 4.4.3)
## fs 1.6.6 2025-04-12 [1] CRAN (R 4.4.3)
## future * 1.68.0 2025-11-17 [1] CRAN (R 4.4.3)
## future.apply 1.20.0 2025-06-06 [1] CRAN (R 4.4.3)
## generics 0.1.4 2025-05-09 [1] CRAN (R 4.4.3)
## GenomeInfoDb * 1.42.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## GenomeInfoDbData 1.2.13 2026-03-05 [1] Bioconductor
## GenomicRanges * 1.58.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## GetoptLong 1.0.5 2020-12-15 [1] CRAN (R 4.4.3)
## getPass 0.2-4 2023-12-10 [1] CRAN (R 4.4.3)
## ggbeeswarm * 0.7.2 2023-04-29 [1] CRAN (R 4.4.3)
## ggh4x * 0.3.1 2025-05-30 [1] CRAN (R 4.4.3)
## ggplot.multistats * 1.0.1 2024-09-25 [1] CRAN (R 4.4.3)
## ggplot2 * 4.0.1 2025-11-14 [1] CRAN (R 4.4.3)
## ggrepel * 0.9.6 2024-09-07 [1] CRAN (R 4.4.3)
## ggridges 0.5.7 2025-08-27 [1] CRAN (R 4.4.3)
## git2r 0.35.0 2024-10-20 [1] CRAN (R 4.4.3)
## GlobalOptions 0.1.2 2020-06-10 [1] CRAN (R 4.4.3)
## globals 0.18.0 2025-05-08 [1] CRAN (R 4.4.3)
## glue 1.8.0 2024-09-30 [1] CRAN (R 4.4.3)
## goftest 1.2-3 2021-10-07 [1] CRAN (R 4.4.3)
## gridExtra 2.3 2017-09-09 [1] CRAN (R 4.4.3)
## gtable 0.3.6 2024-10-25 [1] CRAN (R 4.4.3)
## harmony * 1.2.4 2025-10-10 [1] CRAN (R 4.4.3)
## hexbin 1.28.5 2024-11-13 [1] CRAN (R 4.4.3)
## htmltools 0.5.8.1 2024-04-04 [1] CRAN (R 4.4.3)
## htmlwidgets 1.6.4 2023-12-06 [1] CRAN (R 4.4.3)
## httpuv 1.6.16 2025-04-16 [1] CRAN (R 4.4.3)
## httr 1.4.7 2023-08-15 [1] CRAN (R 4.4.3)
## ica 1.0-3 2022-07-08 [1] CRAN (R 4.4.3)
## igraph 2.1.4 2025-01-23 [1] CRAN (R 4.4.3)
## IRanges * 2.40.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## irlba 2.3.5.1 2022-10-03 [1] CRAN (R 4.4.3)
## iterators 1.0.14 2022-02-05 [1] CRAN (R 4.4.3)
## jquerylib 0.1.4 2021-04-26 [1] CRAN (R 4.4.3)
## jsonlite 2.0.0 2025-03-27 [1] CRAN (R 4.4.3)
## kableExtra * 1.4.0 2024-01-24 [1] CRAN (R 4.4.3)
## KernSmooth 2.23-26 2025-01-01 [1] CRAN (R 4.4.3)
## knitr 1.50 2025-03-16 [1] CRAN (R 4.4.3)
## later 1.4.4 2025-08-27 [1] CRAN (R 4.4.3)
## lattice 0.22-7 2025-04-02 [1] CRAN (R 4.4.3)
## lazyeval 0.2.2 2019-03-15 [1] CRAN (R 4.4.3)
## lifecycle 1.0.4 2023-11-07 [1] CRAN (R 4.4.3)
## limma 3.62.1 2024-11-03 [1] Bioconductor 3.20 (R 4.4.2)
## listenv 0.10.0 2025-11-02 [1] CRAN (R 4.4.3)
## lmtest 0.9-40 2022-03-21 [1] CRAN (R 4.4.3)
## locfit 1.5-9.12 2025-03-05 [1] CRAN (R 4.4.3)
## magrittr * 2.0.4 2025-09-12 [1] CRAN (R 4.4.3)
## MASS 7.3-65 2025-02-28 [1] CRAN (R 4.4.3)
## Matrix * 1.7-4 2025-08-28 [1] CRAN (R 4.4.3)
## MatrixGenerics * 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## matrixStats * 1.5.0 2025-01-07 [1] CRAN (R 4.4.3)
## memoise 2.0.1 2021-11-26 [1] CRAN (R 4.4.3)
## metapod 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## mime 0.13 2025-03-17 [1] CRAN (R 4.4.3)
## miniUI 0.1.2 2025-04-17 [1] CRAN (R 4.4.3)
## nlme 3.1-168 2025-03-31 [1] CRAN (R 4.4.3)
## otel 0.2.0 2025-08-29 [1] CRAN (R 4.4.3)
## parallelly 1.45.1 2025-07-24 [1] CRAN (R 4.4.3)
## patchwork * 1.3.2 2025-08-25 [1] CRAN (R 4.4.3)
## pbapply 1.7-4 2025-07-20 [1] CRAN (R 4.4.3)
## pillar 1.11.1 2025-09-17 [1] CRAN (R 4.4.3)
## pkgbuild 1.4.8 2025-05-26 [1] CRAN (R 4.4.3)
## pkgconfig 2.0.3 2019-09-22 [1] CRAN (R 4.4.3)
## pkgload 1.4.1 2025-09-23 [1] CRAN (R 4.4.3)
## plotly 4.11.0 2025-06-19 [1] CRAN (R 4.4.3)
## plyr 1.8.9 2023-10-02 [1] CRAN (R 4.4.3)
## png 0.1-8 2022-11-29 [1] CRAN (R 4.4.3)
## polyclip 1.10-7 2024-07-23 [1] CRAN (R 4.4.3)
## processx 3.8.6 2025-02-21 [1] CRAN (R 4.4.3)
## progressr 0.18.0 2025-11-06 [1] CRAN (R 4.4.3)
## promises 1.5.0 2025-11-01 [1] CRAN (R 4.4.3)
## ps 1.9.1 2025-04-12 [1] CRAN (R 4.4.3)
## purrr * 1.2.0 2025-11-04 [1] CRAN (R 4.4.3)
## R.methodsS3 1.8.2 2022-06-13 [1] CRAN (R 4.4.3)
## R.oo 1.27.1 2025-05-02 [1] CRAN (R 4.4.3)
## R.utils 2.13.0 2025-02-24 [1] CRAN (R 4.4.3)
## R6 2.6.1 2025-02-15 [1] CRAN (R 4.4.3)
## RANN 2.6.2 2024-08-25 [1] CRAN (R 4.4.3)
## RColorBrewer * 1.1-3 2022-04-03 [1] CRAN (R 4.4.3)
## Rcpp * 1.1.0 2025-07-02 [1] CRAN (R 4.4.3)
## RcppAnnoy 0.0.22 2024-01-23 [1] CRAN (R 4.4.3)
## RcppHNSW 0.6.0 2024-02-04 [1] CRAN (R 4.4.3)
## readxl * 1.4.5 2025-03-07 [1] CRAN (R 4.4.3)
## registry 0.5-1 2019-03-05 [1] CRAN (R 4.4.3)
## remotes 2.5.0 2024-03-17 [1] CRAN (R 4.4.3)
## reshape2 1.4.5 2025-11-12 [1] CRAN (R 4.4.3)
## reticulate 1.44.1 2025-11-14 [1] CRAN (R 4.4.3)
## rhdf5 * 2.50.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## rhdf5filters 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## Rhdf5lib 1.28.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## rjson 0.2.23 2024-09-16 [1] CRAN (R 4.4.3)
## rlang 1.1.6 2025-04-11 [1] CRAN (R 4.4.3)
## rmarkdown 2.30 2025-09-28 [1] CRAN (R 4.4.3)
## rmdformats 1.0.4 2022-05-17 [1] CRAN (R 4.4.3)
## ROCR 1.0-11 2020-05-02 [1] CRAN (R 4.4.3)
## rprojroot 2.1.1 2025-08-26 [1] CRAN (R 4.4.3)
## RSpectra 0.16-2 2024-07-18 [1] CRAN (R 4.4.3)
## rstudioapi 0.17.1 2024-10-22 [1] CRAN (R 4.4.3)
## rsvd 1.0.5 2021-04-16 [1] CRAN (R 4.4.1)
## Rtsne 0.17 2023-12-07 [1] CRAN (R 4.4.3)
## S4Arrays 1.6.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## S4Vectors * 0.44.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## S7 0.2.1 2025-11-14 [1] CRAN (R 4.4.3)
## sass 0.4.10 2025-04-11 [1] CRAN (R 4.4.3)
## ScaledMatrix 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## scales * 1.4.0 2025-04-24 [1] CRAN (R 4.4.3)
## scater * 1.34.1 2025-03-03 [1] Bioconductor 3.20 (R 4.4.2)
## scattermore 1.2 2023-06-12 [1] CRAN (R 4.4.3)
## scran * 1.34.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## sctransform 0.4.2 2025-04-30 [1] CRAN (R 4.4.3)
## scuttle * 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## seriation * 1.5.8 2025-08-20 [1] CRAN (R 4.4.3)
## sessioninfo 1.2.3 2025-02-05 [1] CRAN (R 4.4.3)
## Seurat * 5.3.1 2025-10-29 [1] CRAN (R 4.4.3)
## SeuratObject * 5.2.0 2025-08-27 [1] CRAN (R 4.4.3)
## shape 1.4.6.1 2024-02-23 [1] CRAN (R 4.4.3)
## shiny 1.11.1 2025-07-03 [1] CRAN (R 4.4.3)
## SingleCellExperiment * 1.28.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## sp * 2.2-0 2025-02-01 [1] CRAN (R 4.4.3)
## spam 2.11-1 2025-01-20 [1] CRAN (R 4.4.3)
## SparseArray 1.6.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## spatstat.data 3.1-9 2025-10-18 [1] CRAN (R 4.4.3)
## spatstat.explore 3.6-0 2025-11-22 [1] CRAN (R 4.4.3)
## spatstat.geom 3.6-1 2025-11-20 [1] CRAN (R 4.4.3)
## spatstat.random 3.4-3 2025-11-21 [1] CRAN (R 4.4.3)
## spatstat.sparse 3.1-0 2024-06-21 [1] CRAN (R 4.4.3)
## spatstat.univar 3.1-5 2025-11-17 [1] CRAN (R 4.4.3)
## spatstat.utils 3.2-0 2025-09-20 [1] CRAN (R 4.4.3)
## statmod 1.5.1 2025-10-09 [1] CRAN (R 4.4.3)
## strex * 2.0.1 2024-10-03 [1] CRAN (R 4.4.3)
## stringi 1.8.7 2025-03-27 [1] CRAN (R 4.4.3)
## stringr * 1.6.0 2025-11-04 [1] CRAN (R 4.4.3)
## SummarizedExperiment * 1.36.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## survival 3.8-3 2024-12-17 [1] CRAN (R 4.4.3)
## svglite 2.2.2 2025-10-21 [1] CRAN (R 4.4.3)
## systemfonts 1.3.1 2025-10-01 [1] CRAN (R 4.4.3)
## tensor 1.5.1 2025-06-17 [1] CRAN (R 4.4.3)
## textshaping 1.0.4 2025-10-10 [1] CRAN (R 4.4.3)
## tibble 3.3.0 2025-06-08 [1] CRAN (R 4.4.3)
## tidyr 1.3.1 2024-01-24 [1] CRAN (R 4.4.3)
## tidyselect 1.2.1 2024-03-11 [1] CRAN (R 4.4.3)
## TSP 1.2.6 2025-11-27 [1] CRAN (R 4.4.3)
## UCSC.utils 1.2.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## usethis 3.2.1 2025-09-06 [1] CRAN (R 4.4.3)
## uwot * 0.2.4 2025-11-10 [1] CRAN (R 4.4.3)
## vctrs 0.6.5 2023-12-01 [1] CRAN (R 4.4.3)
## vipor 0.4.7 2023-12-18 [1] CRAN (R 4.4.3)
## viridis * 0.6.5 2024-01-29 [1] CRAN (R 4.4.3)
## viridisLite * 0.4.2 2023-05-02 [1] CRAN (R 4.4.3)
## whisker 0.4.1 2022-12-05 [1] CRAN (R 4.4.3)
## withr 3.0.2 2024-10-28 [1] CRAN (R 4.4.3)
## workflowr * 1.7.2 2025-08-18 [1] CRAN (R 4.4.3)
## xfun 0.54 2025-10-30 [1] CRAN (R 4.4.3)
## xgboost * 3.1.2.1 2026-01-06 [1] local
## xml2 1.5.0 2025-11-17 [1] CRAN (R 4.4.3)
## xtable 1.8-4 2019-04-21 [1] CRAN (R 4.4.3)
## XVector 0.46.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## yaml * 2.3.11 2025-11-28 [1] CRAN (R 4.4.3)
## zellkonverter * 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## zlibbioc 1.52.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## zoo 1.8-14 2025-04-10 [1] CRAN (R 4.4.3)
##
## [1] /home/macnairw/packages/scprocess/.snakemake/conda/4fef11cadd34f9d2d13a0d6139d09340_/lib/R/library
## * ── Packages attached to the search path.
##
## ──────────────────────────────────────────────────────────────────────────────