Cells that passed quality control filtering were integrated with
Harmony together with doublets identified using
scDblfinder. Clustering of cells at a high resolution is
then performed. Cells that appear in clusters which are enriched in
doublets are excluded from further analysis. Integration of cells and
clustering is repeated after removing doublet enriched clusters.
Doublets over UMAP
The plot displays a binned UMAP with the proportion of doublets as well as the number of doublets in each bin.
g_dbl = plot_umap_doublets(int_dbl)
g_dens = plot_umap_density(int_dbl[, .(cell_id, UMAP1, UMAP2) ])
g = g_dbl + g_dens
print(g)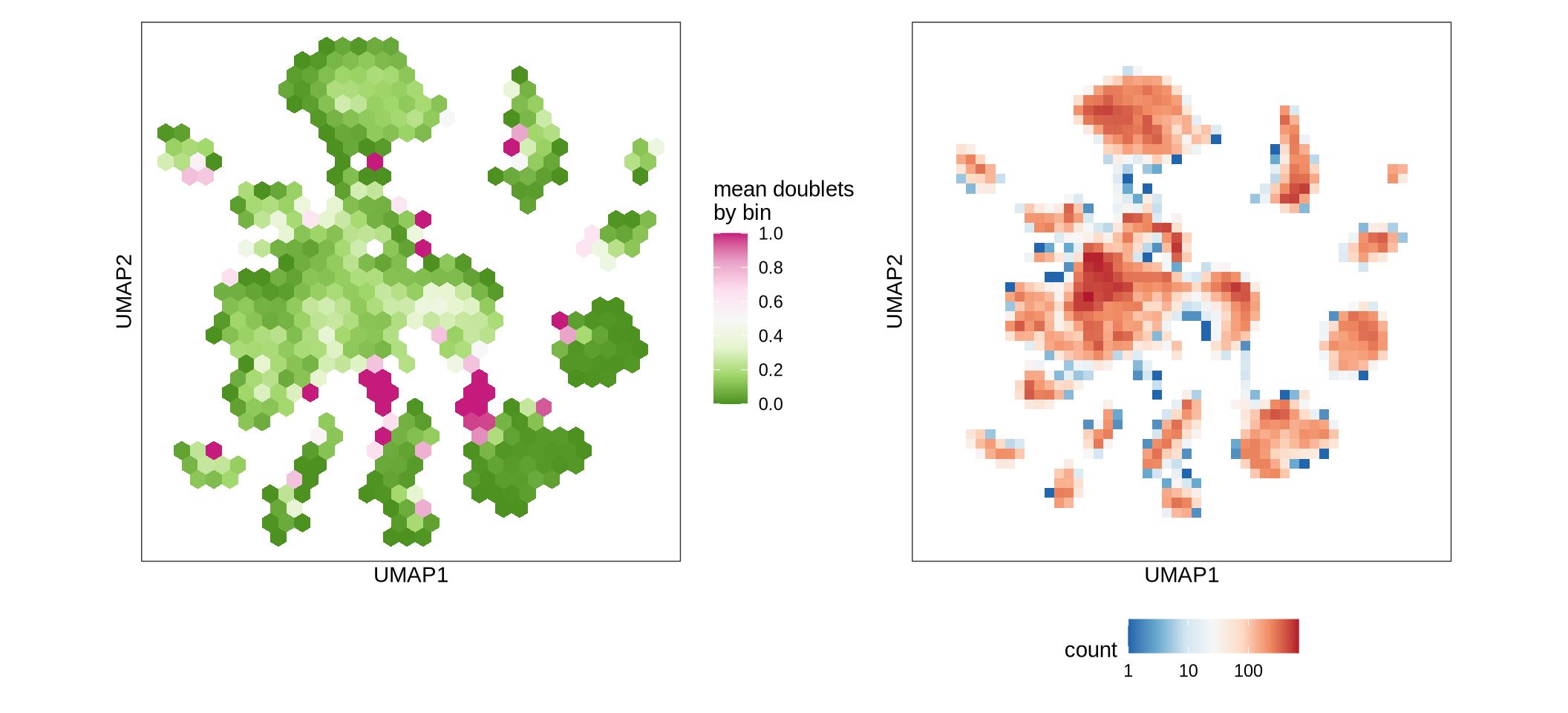
Doublet proportions in clusters
The plot shows the proportion of doublets for each cluster in relation to the total number of cells in that cluster. Clusters with a doublet proportion exceeding 50% are excluded from further analysis.
( plot_doublet_clusters(int_dbl, dbl_cl_prop) )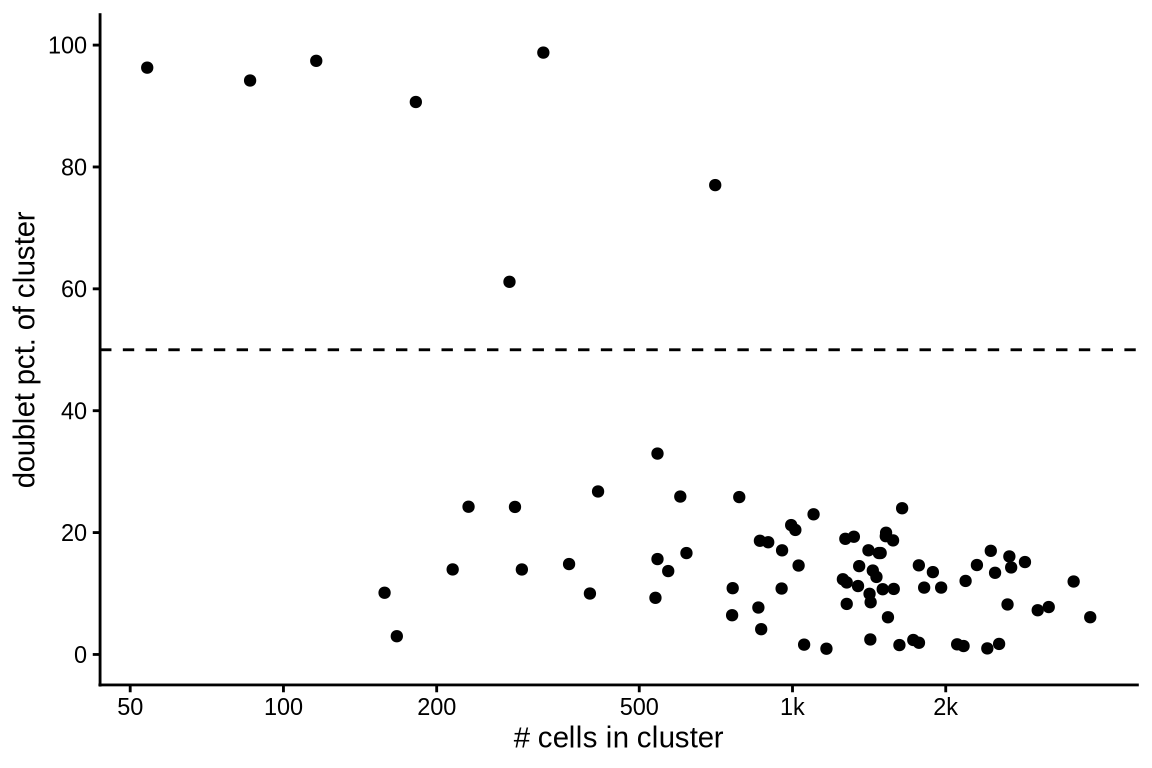
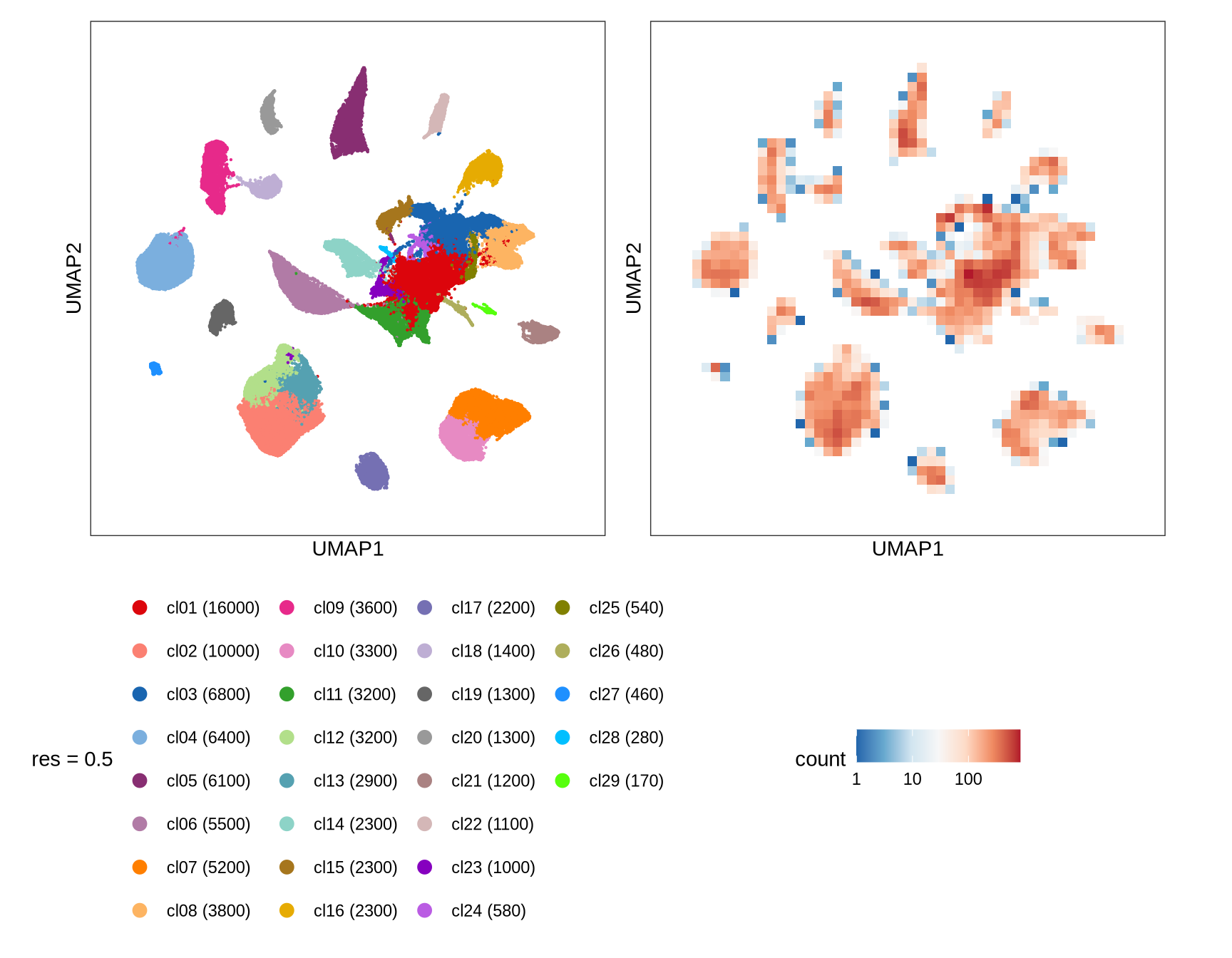
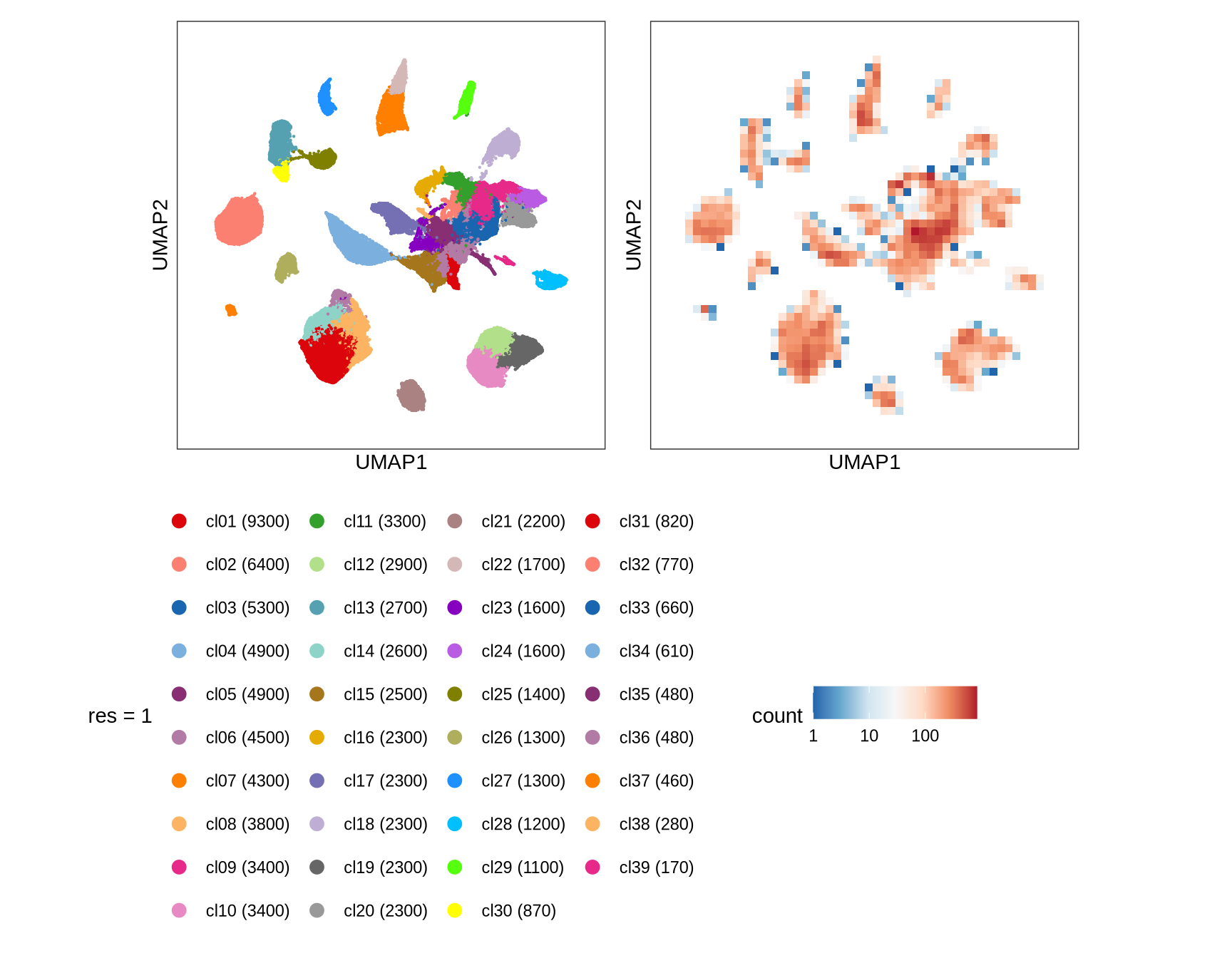
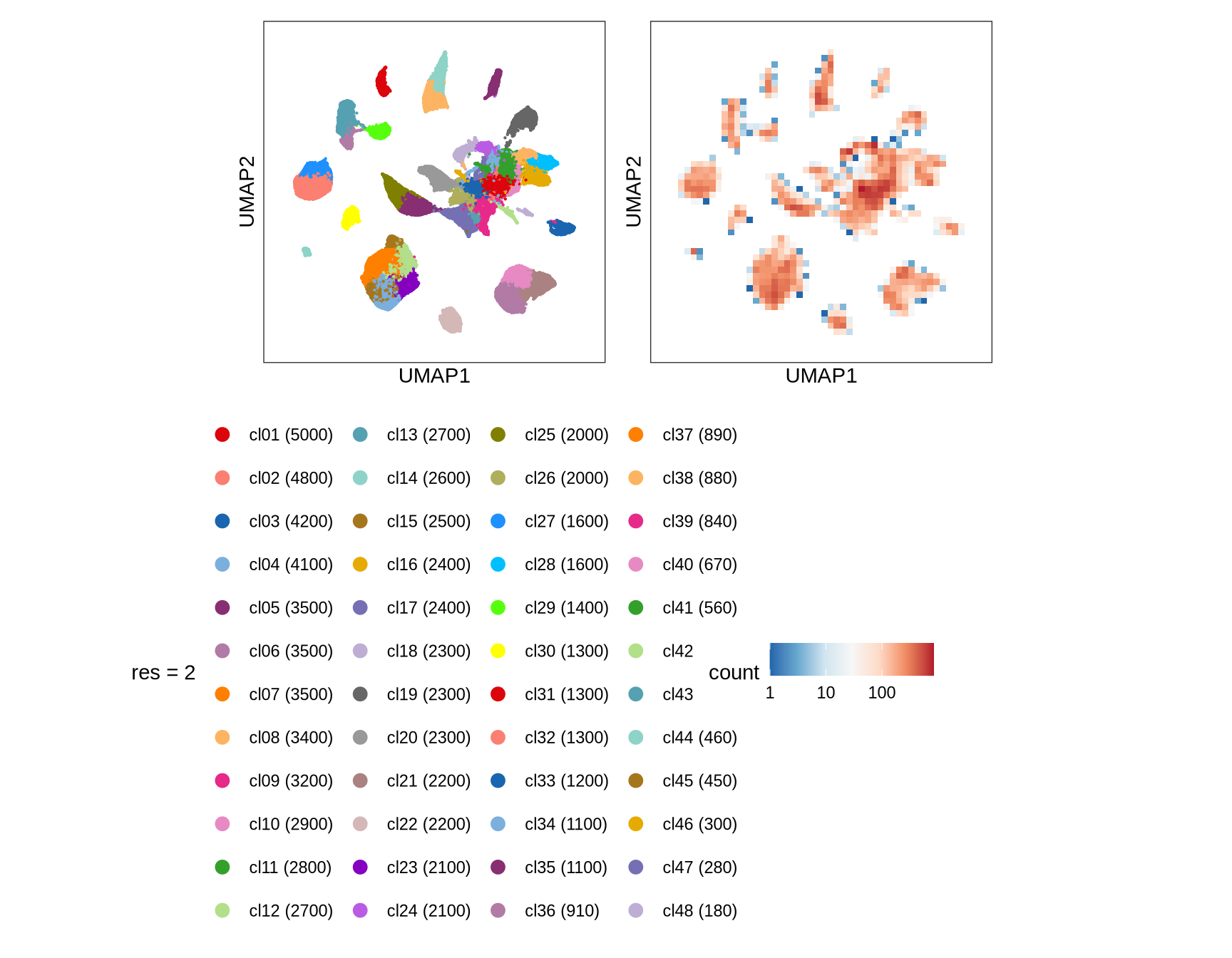
Clusters over UMAP
Clustering of data is performed at different resolution values. For each value the clusters are displayed over a UMAP together with a plot showing the density of cells.
After removing doublets, there were 94810 QC-ed cells used for integration.
for (res in res_ls) {
cat('### ', res, '\n')
g_cl = plot_umap_cluster(
umap_dt = int_dt[, .(cell_id, UMAP1, UMAP2) ],
clust_dt = int_dt[, .(cell_id, cluster = get(paste0('RNA_snn_res.', res))) ],
name = sprintf('res = %s', res))
g_dens = plot_umap_density(int_dt[, .(cell_id, UMAP1, UMAP2) ])
g = g_cl + g_dens
print(g)
cat('\n\n')
}0.1
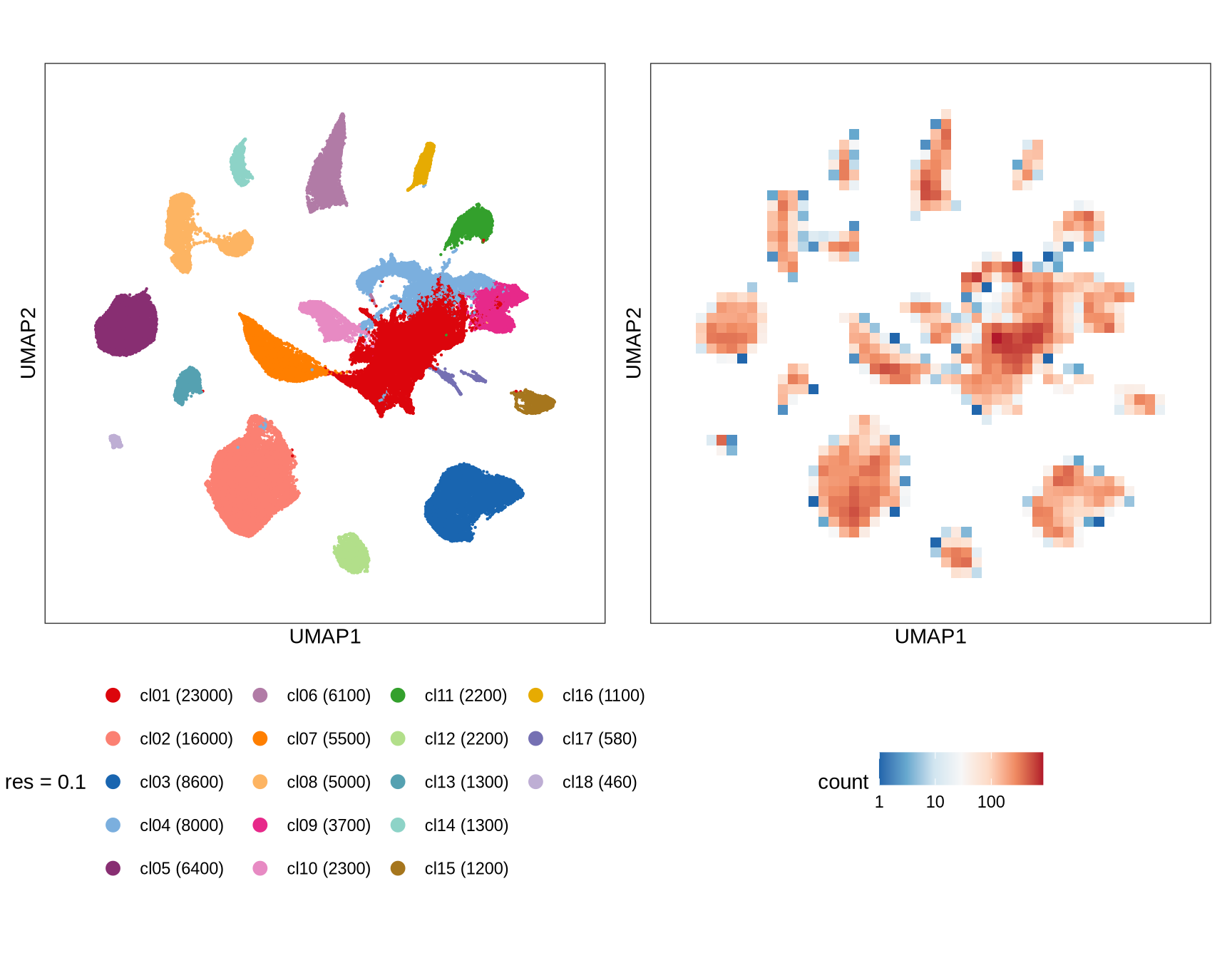
0.2
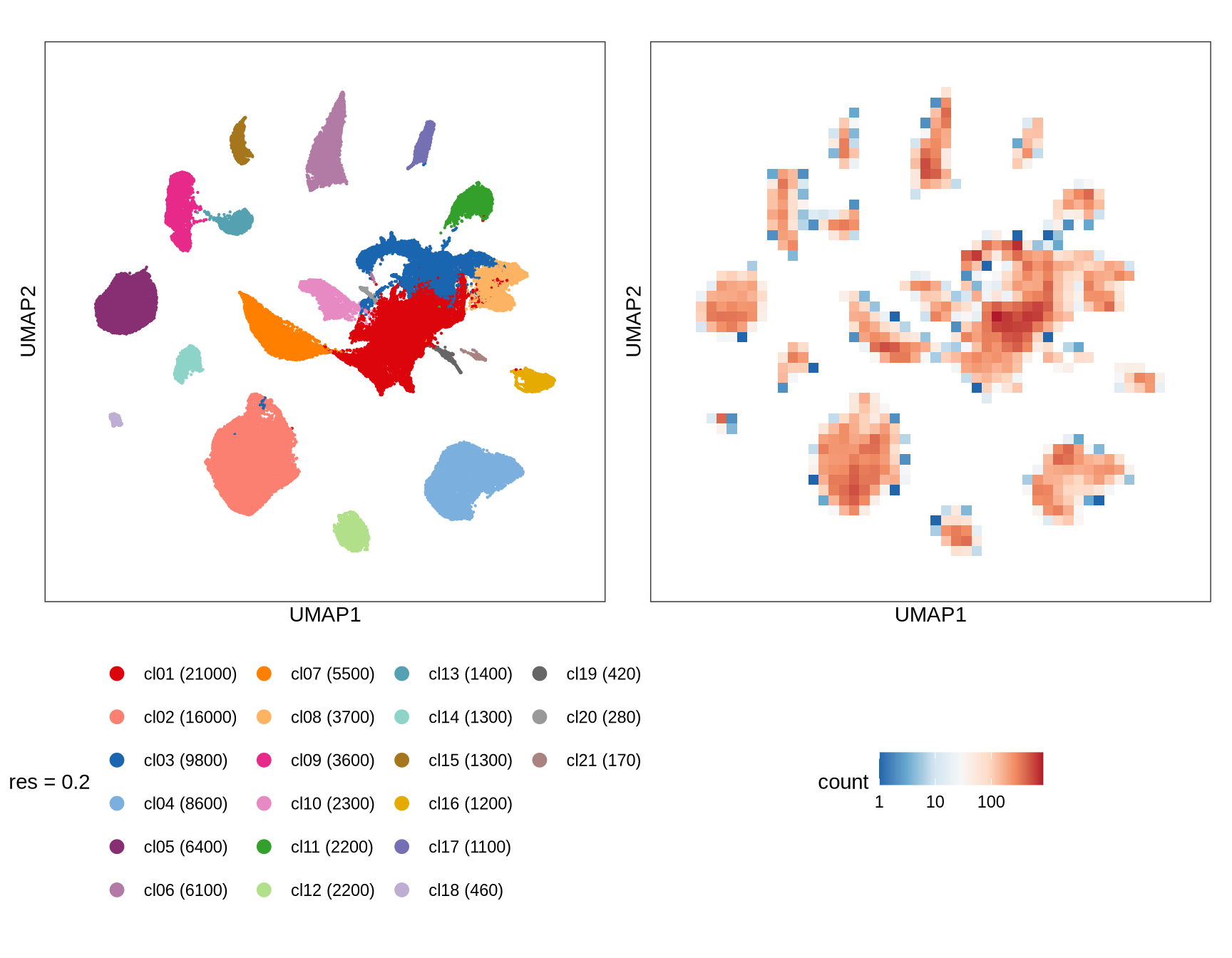
0.5
1
2
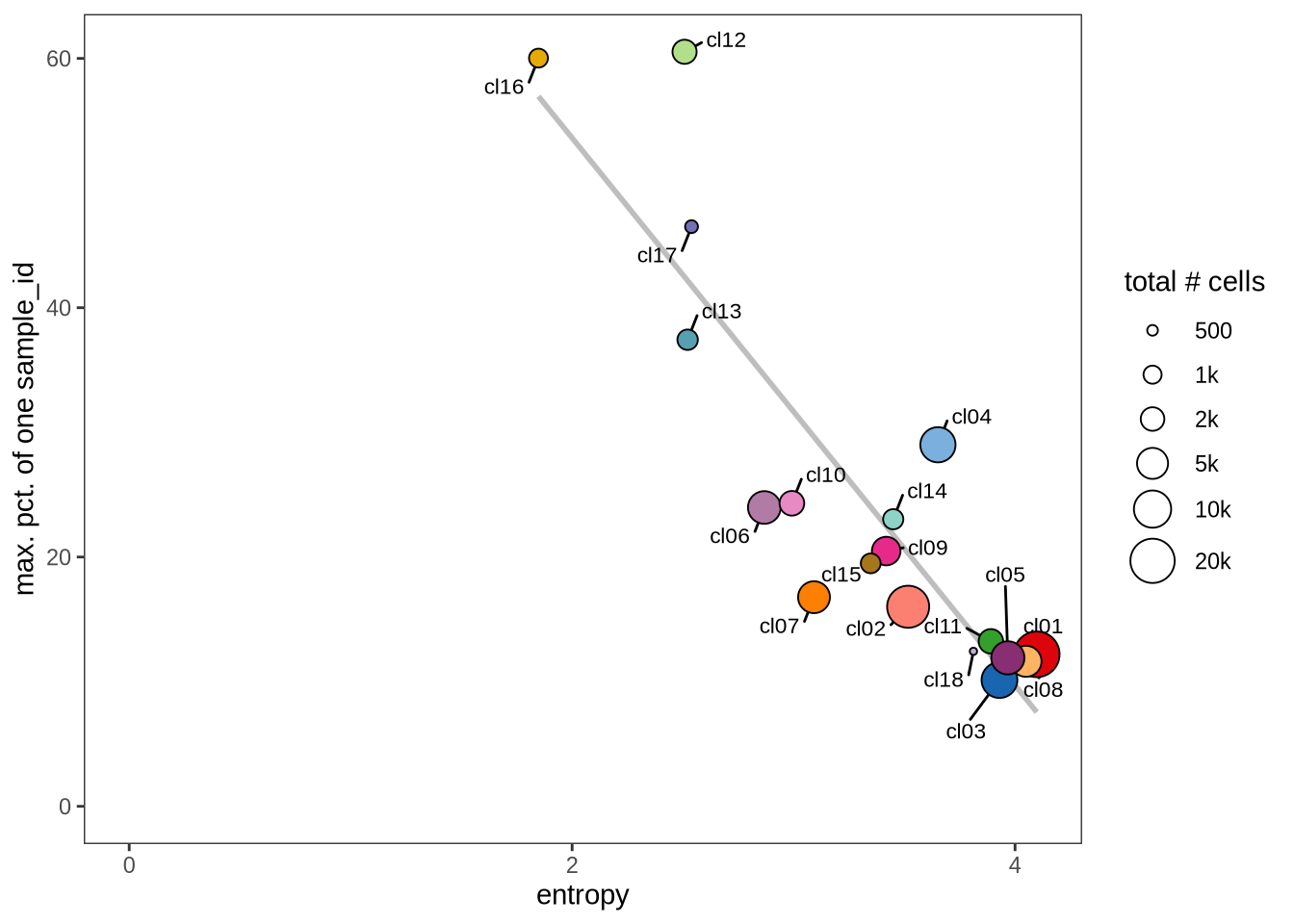
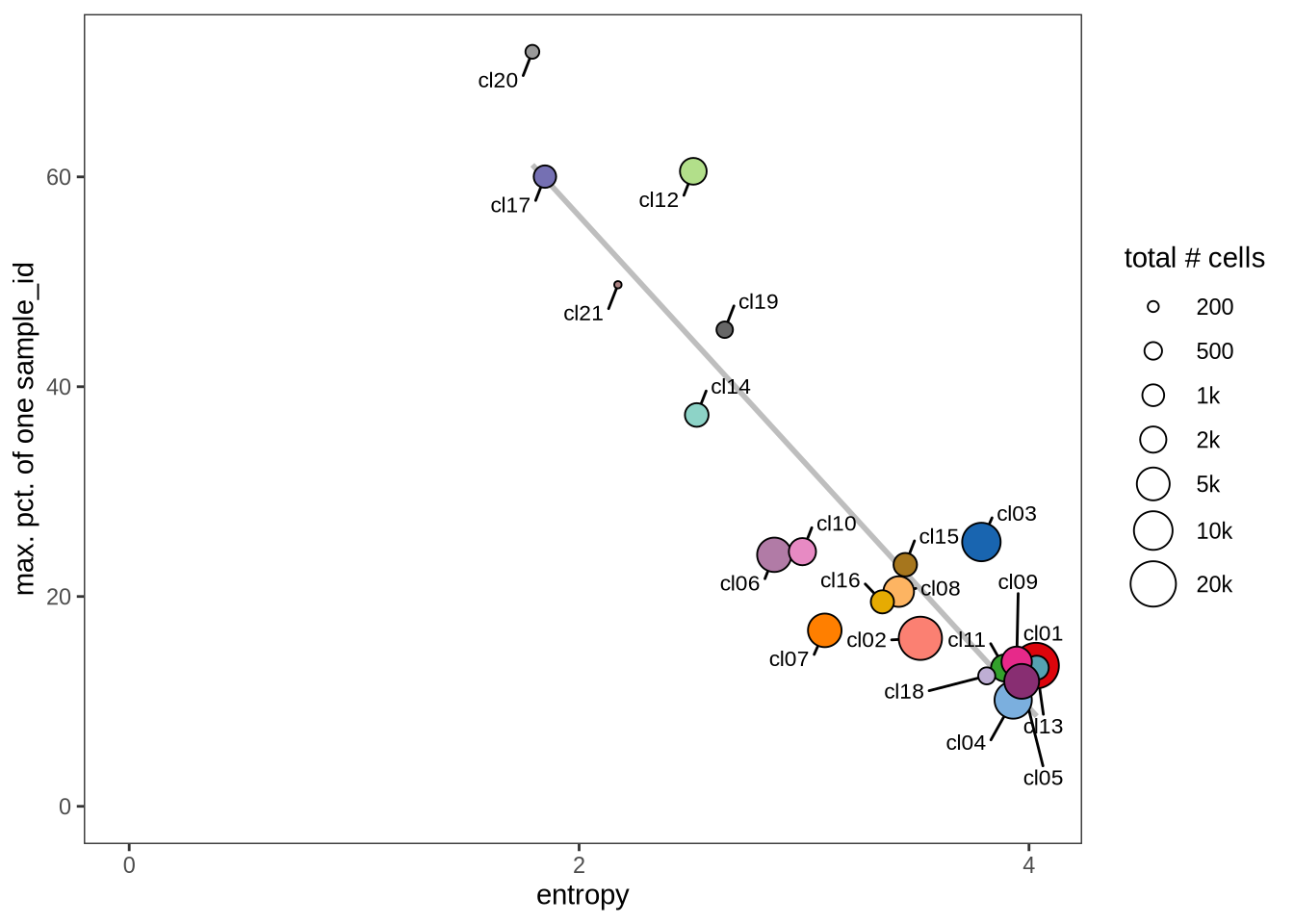
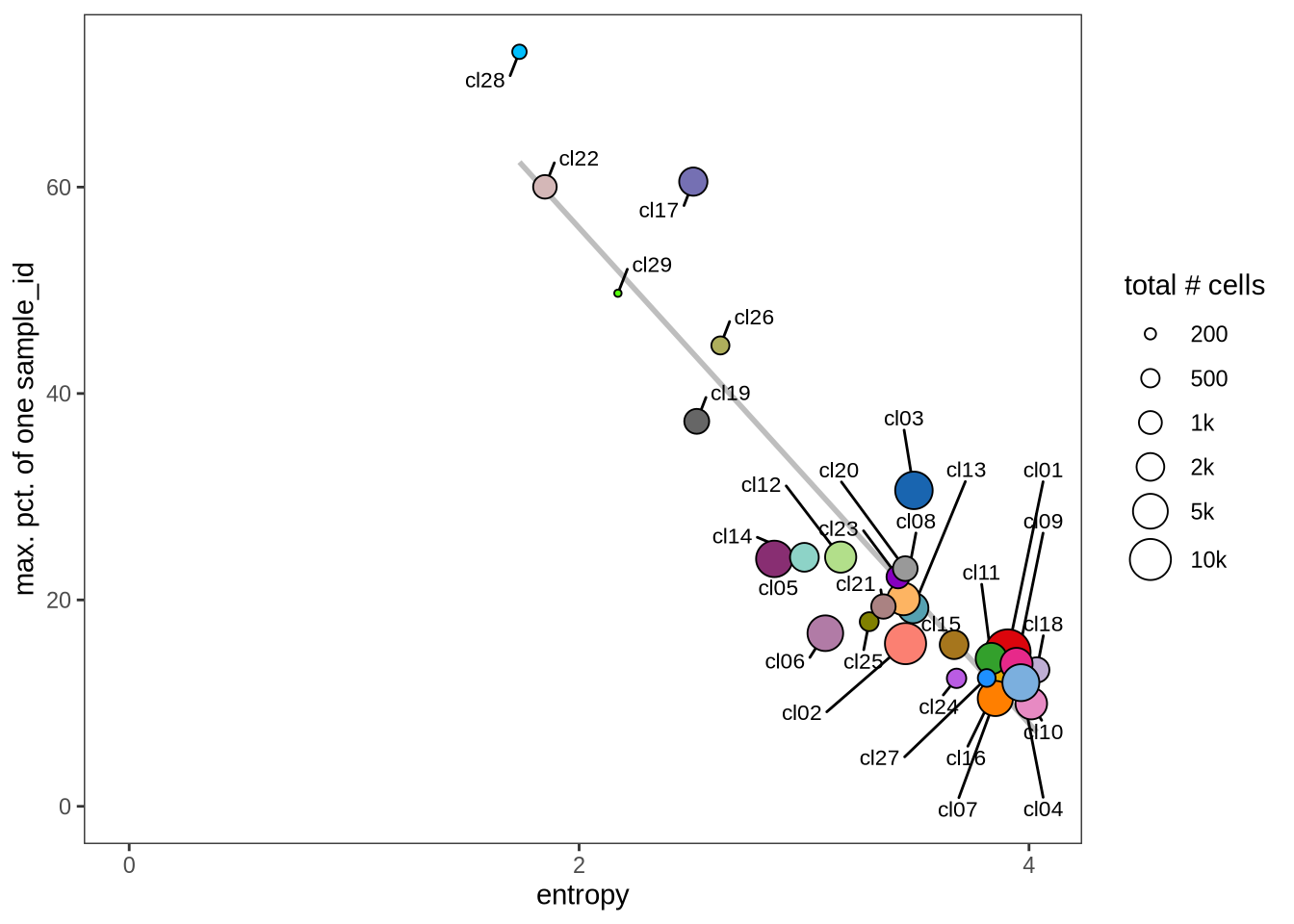
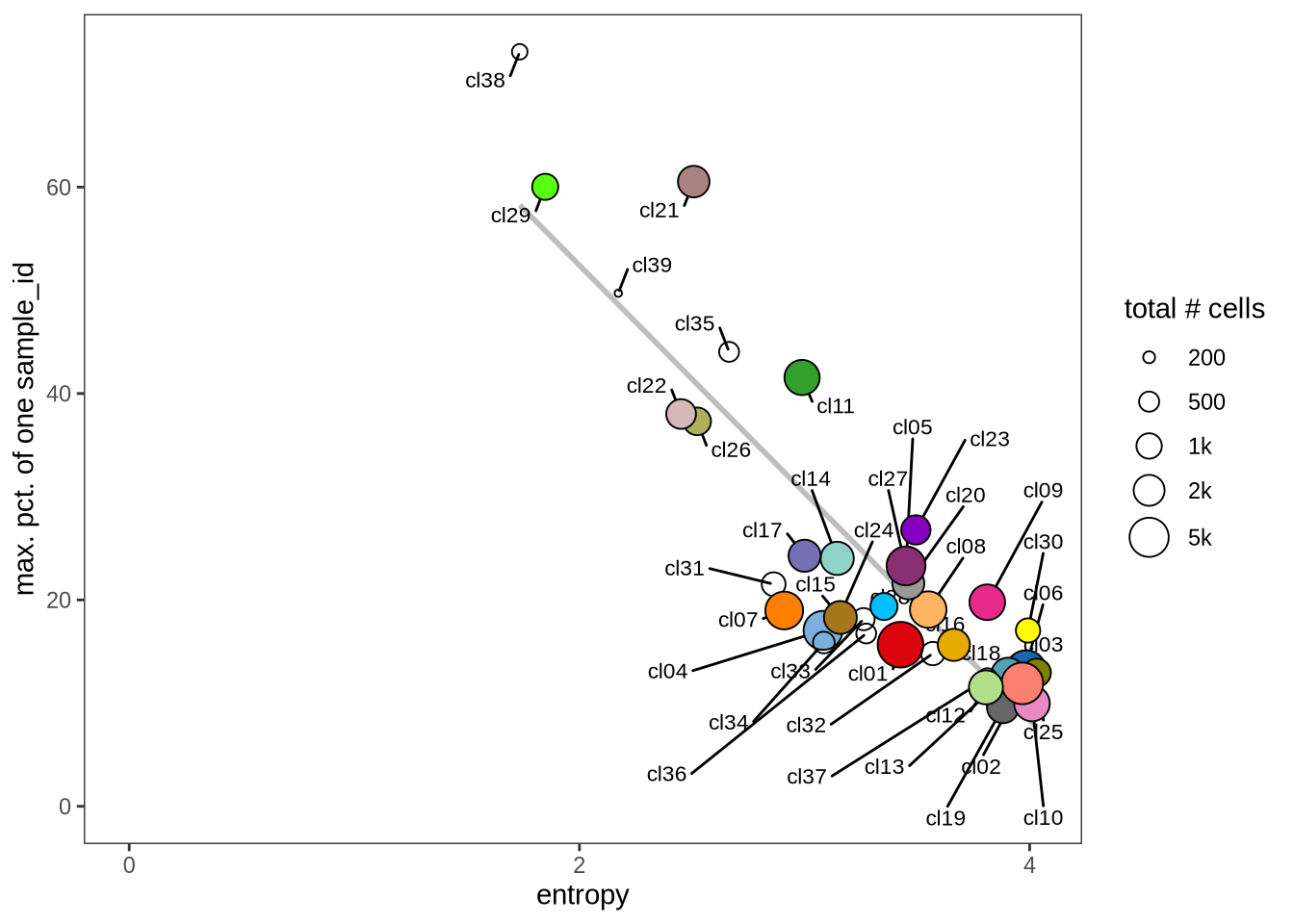
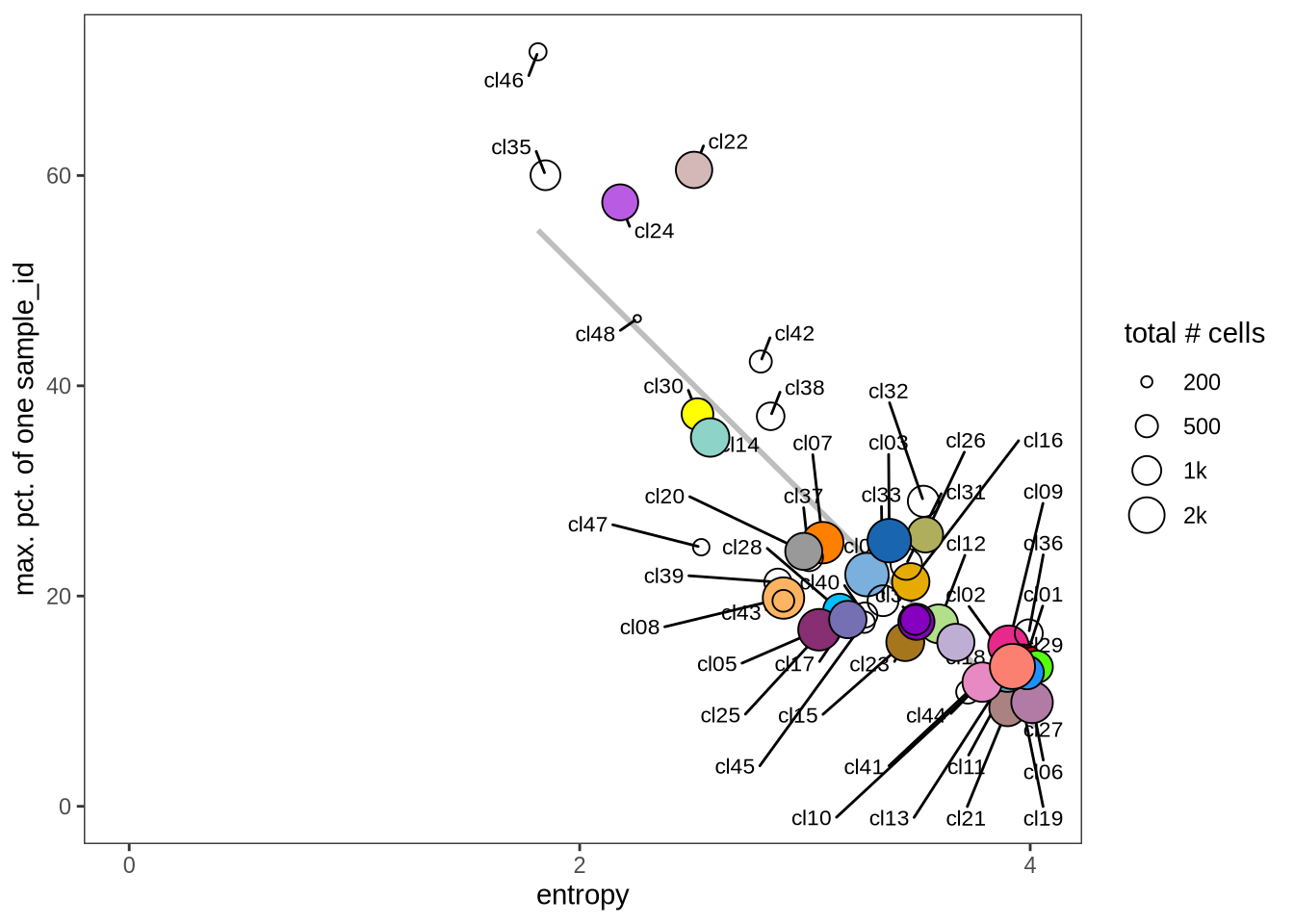
Evaluating cluster distribution across samples
This plot visualizes the relationship between the entropy of clusters and the maximum proportion of cells from a single sample within each cluster (higher values suggest that a cluster predominantly contains cells from a single sample). Entropy measures how evenly distributed cells are across samples within each cluster—higher entropy indicates that cells from different samples are more evenly distributed, while lower entropy suggests that a cluster is dominated by cells from a small number of samples.
n_clusts = res_ls %>% lapply(function(res){
tmp_dt = int_dt[, .(cluster = get(paste0('RNA_snn_res.', res)))]
length(unique(tmp_dt$cluster))
})
res_ls_pl = res_ls[n_clusts > 1]
for (res in res_ls_pl) {
cat('### ', res, '\n')
input_dt = int_dt[, .(batch_var = get(batch_var), cell_id,
cluster = get(paste0('RNA_snn_res.', res)))]
suppressWarnings(print(plot_cluster_entropies(input_dt, batch_var, what = "norm")))
cat('\n\n')
}0.1
0.2
0.5
1
2
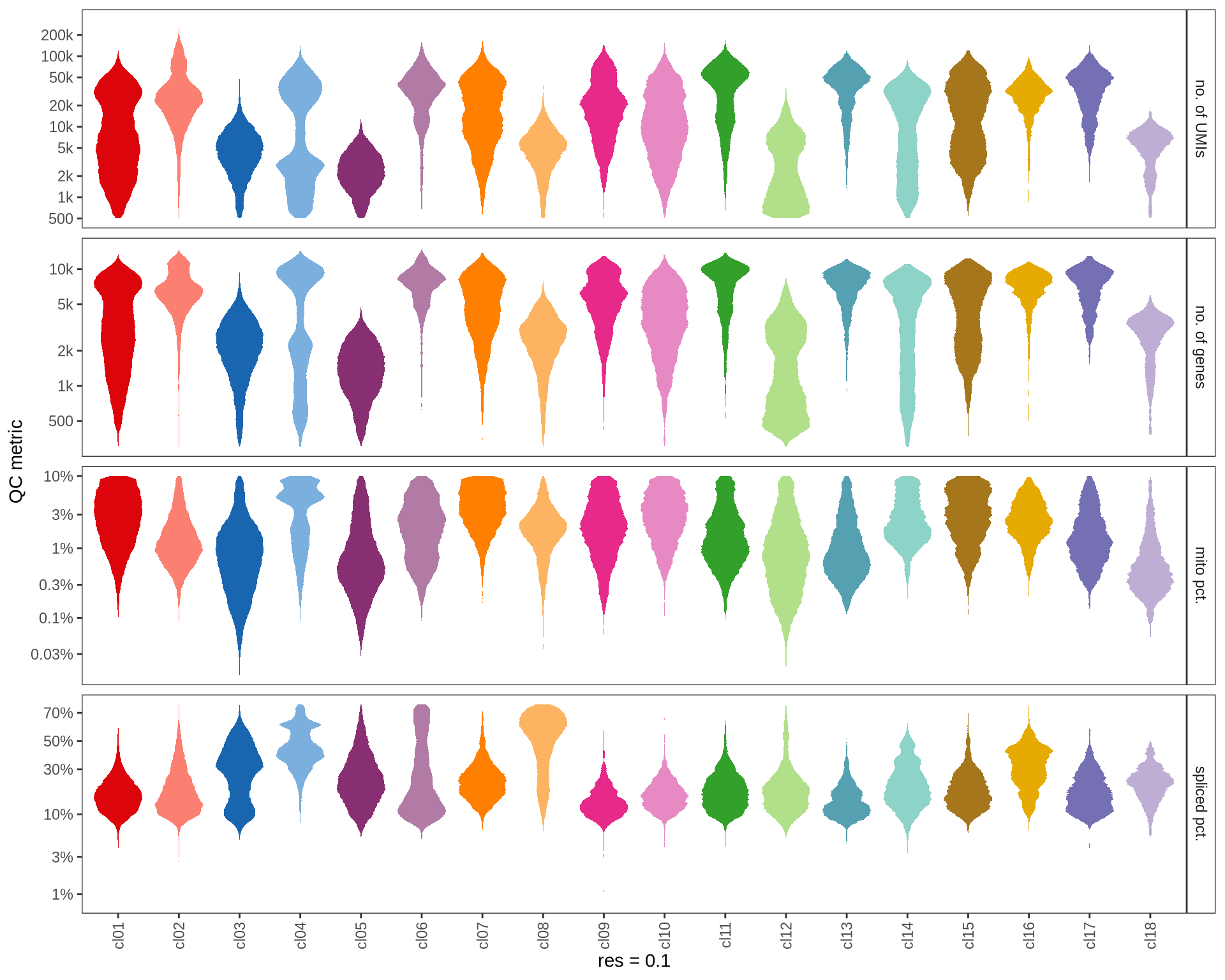
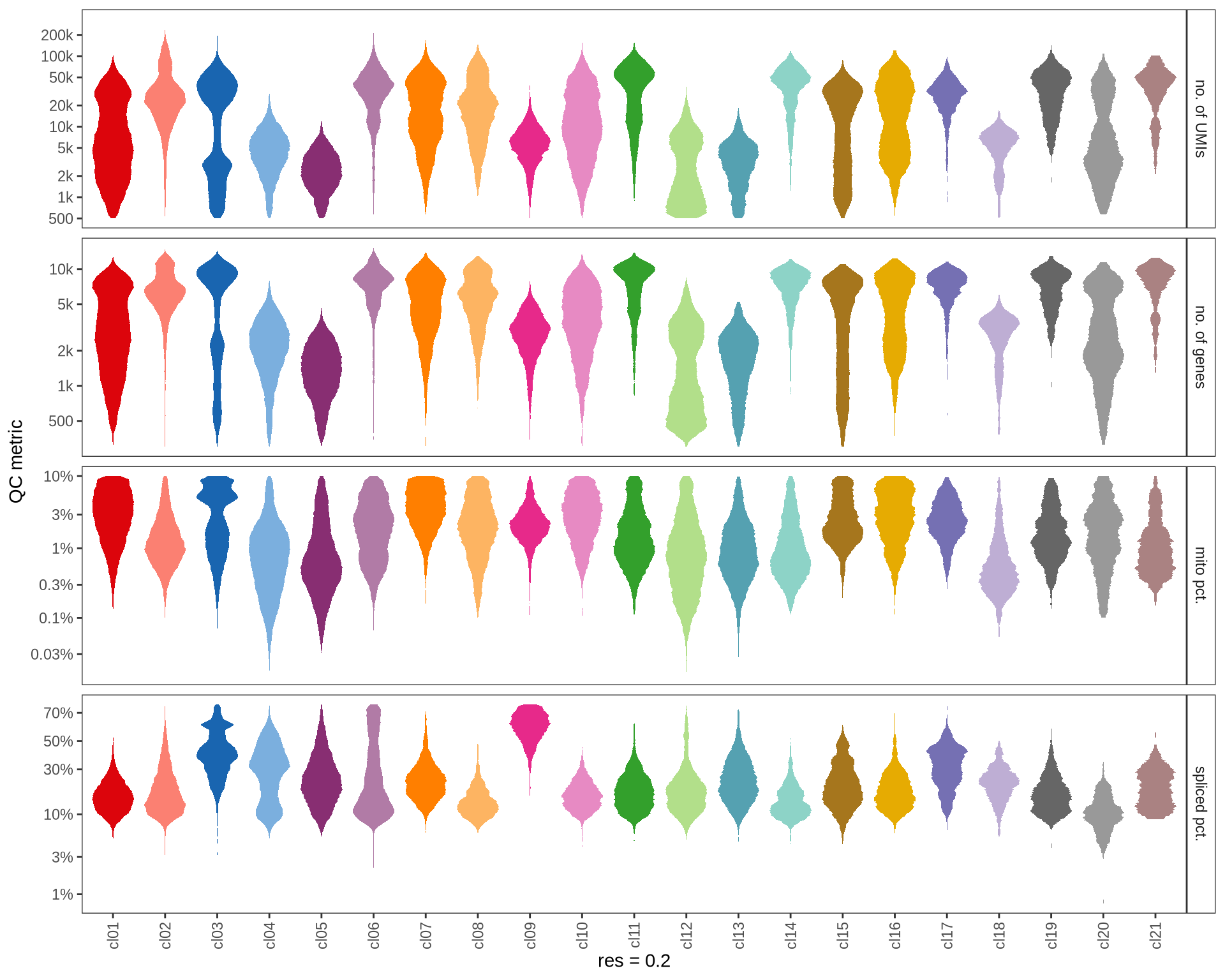
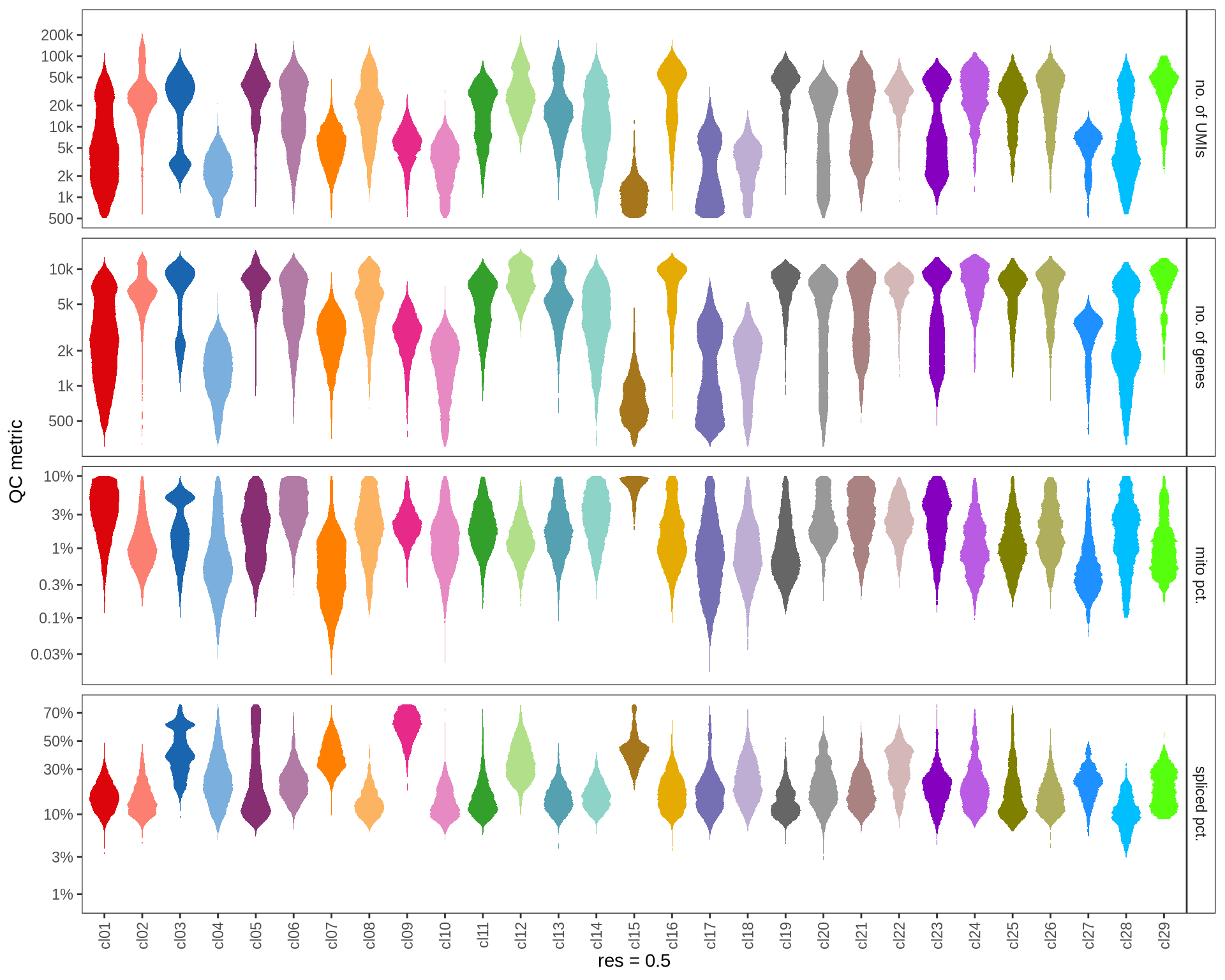
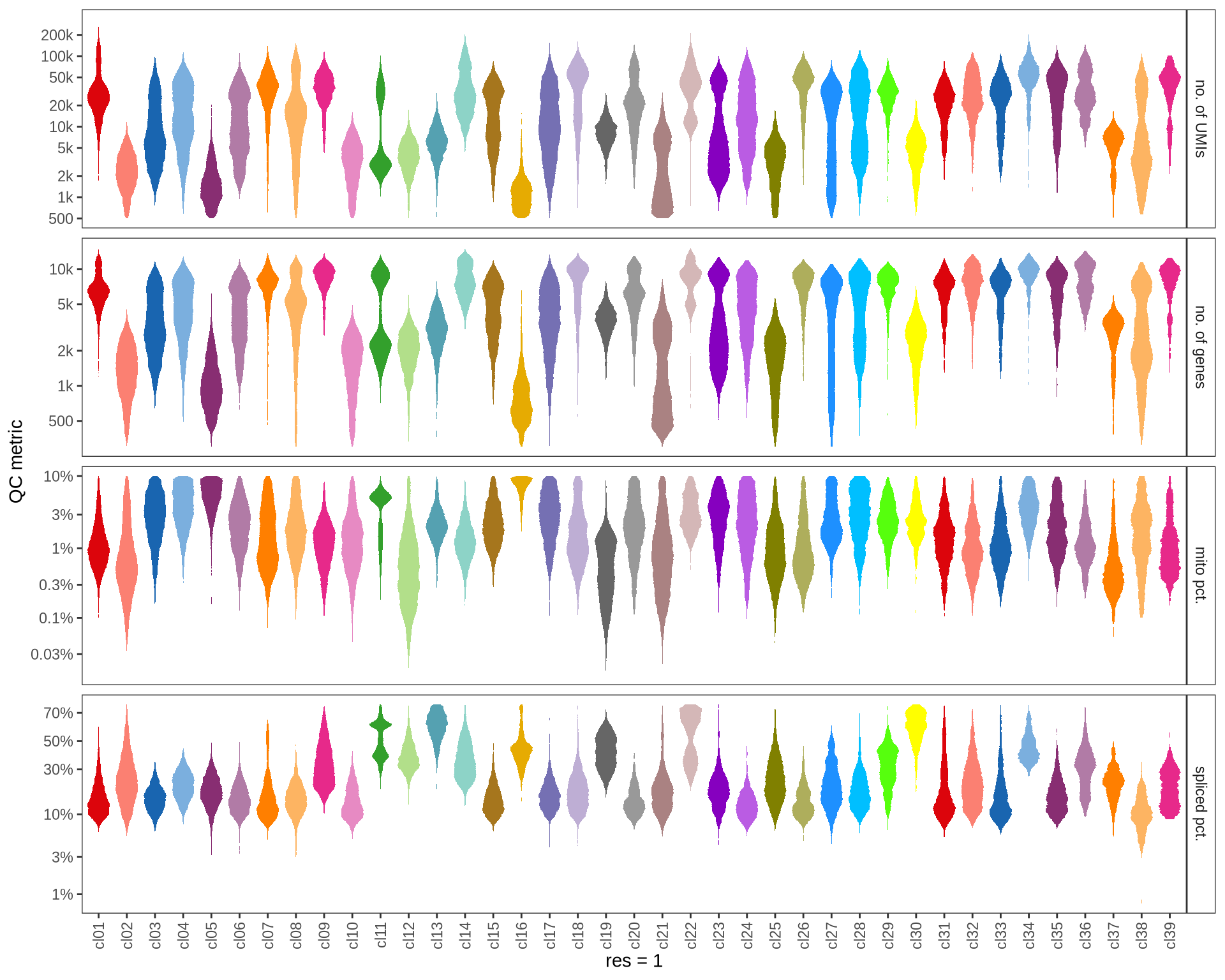
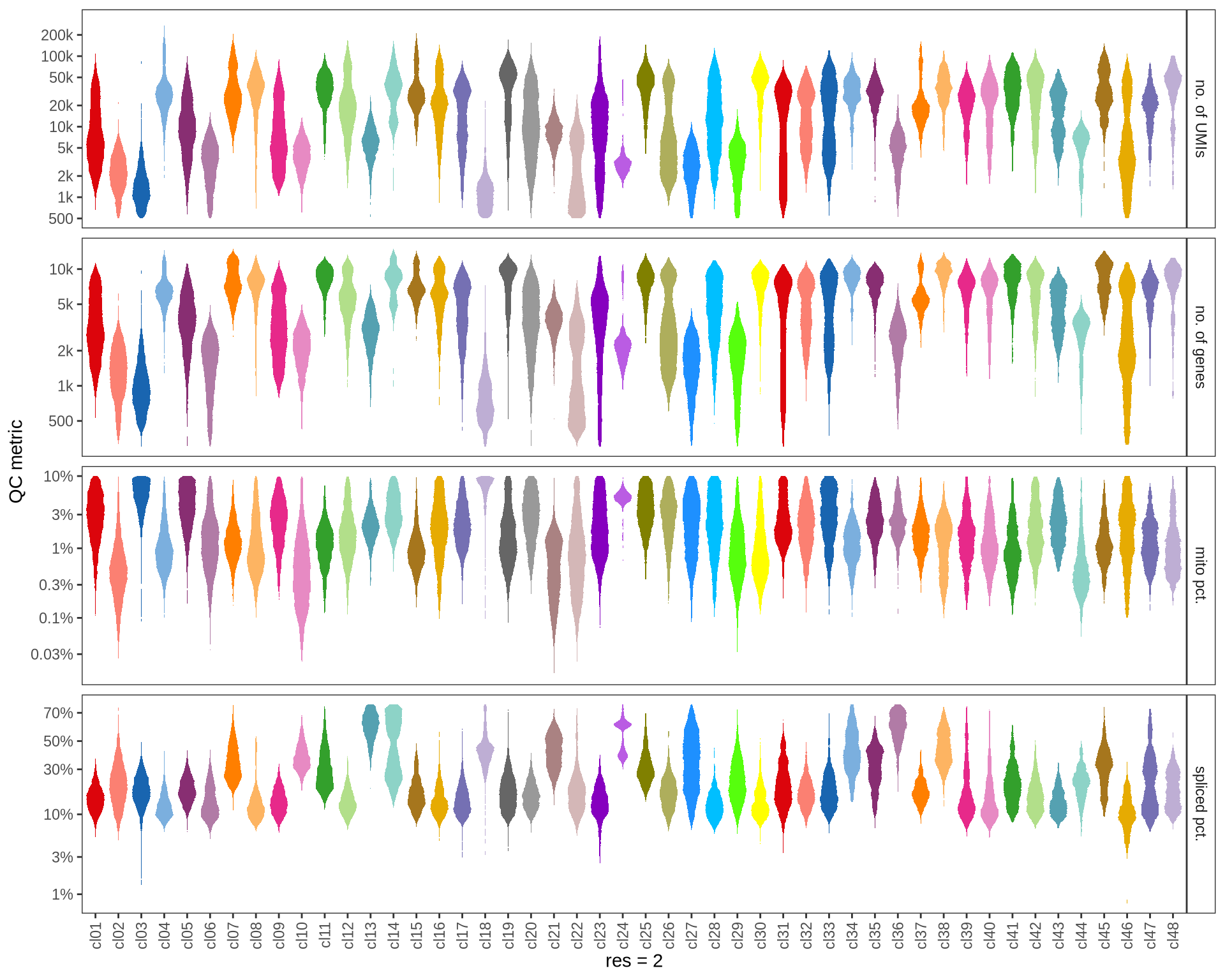
Check QC metrics of clusters
Distributions of QC metrics (library size, number of features, mitochondrial proportion, and spliced proportion) are shown for each cluster across different resolution values.
for (res in res_ls) {
cat('### ', res, '\n')
suppressWarnings(print(plot_cluster_qc_distns(
qc_melt,
clust_dt = int_dt[, .(cell_id, cluster = get(paste0('RNA_snn_res.', res)))],
name = res)))
cat('\n\n')
}0.1
0.2
0.5
1
2
R session info
Details of the R package versions used are given below.
devtools::session_info()## ─ Session info ───────────────────────────────────────────────────────────────
## setting value
## version R version 4.4.3 (2025-02-28)
## os Red Hat Enterprise Linux 8.10 (Ootpa)
## system x86_64, linux-gnu
## ui X11
## language (EN)
## collate en_US.UTF-8
## ctype en_US.UTF-8
## tz Europe/Zurich
## date 2026-03-25
## pandoc 3.8.2.1 @ /home/macnairw/packages/scprocess/.snakemake/conda/4fef11cadd34f9d2d13a0d6139d09340_/bin/ (via rmarkdown)
## quarto NA
##
## ─ Packages ───────────────────────────────────────────────────────────────────
## package * version date (UTC) lib source
## abind 1.4-8 2024-09-12 [1] CRAN (R 4.4.3)
## assertthat * 0.2.1 2019-03-21 [1] CRAN (R 4.4.3)
## basilisk 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## basilisk.utils 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## beachmat 2.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## beeswarm 0.4.0 2021-06-01 [1] CRAN (R 4.4.3)
## Biobase * 2.66.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocGenerics * 0.52.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocManager 1.30.27 2025-11-14 [1] CRAN (R 4.4.3)
## BiocNeighbors 2.0.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## BiocParallel * 1.40.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## BiocSingular 1.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## BiocStyle * 2.34.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## bluster 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## bookdown 0.45 2025-10-03 [1] CRAN (R 4.4.3)
## bslib 0.9.0 2025-01-30 [1] CRAN (R 4.4.3)
## ca 0.71.1 2020-01-24 [1] CRAN (R 4.4.3)
## cachem 1.1.0 2024-05-16 [1] CRAN (R 4.4.3)
## callr 3.7.6 2024-03-25 [1] CRAN (R 4.4.3)
## cellranger 1.1.0 2016-07-27 [1] CRAN (R 4.4.3)
## circlize * 0.4.16 2024-02-20 [1] CRAN (R 4.4.3)
## cli 3.6.5 2025-04-23 [1] CRAN (R 4.4.3)
## clue 0.3-66 2024-11-13 [1] CRAN (R 4.4.3)
## cluster 2.1.8.1 2025-03-12 [1] CRAN (R 4.4.3)
## codetools 0.2-20 2024-03-31 [1] CRAN (R 4.4.3)
## colorspace 2.1-2 2025-09-22 [1] CRAN (R 4.4.3)
## ComplexHeatmap * 2.22.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## cowplot 1.2.0 2025-07-07 [1] CRAN (R 4.4.3)
## crayon 1.5.3 2024-06-20 [1] CRAN (R 4.4.3)
## data.table * 1.17.8 2025-07-10 [1] CRAN (R 4.4.3)
## DelayedArray 0.32.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## deldir 2.0-4 2024-02-28 [1] CRAN (R 4.4.3)
## devtools 2.4.6 2025-10-03 [1] CRAN (R 4.4.3)
## digest 0.6.39 2025-11-19 [1] CRAN (R 4.4.3)
## dir.expiry 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## doParallel 1.0.17 2022-02-07 [1] CRAN (R 4.4.3)
## dotCall64 1.2 2024-10-04 [1] CRAN (R 4.4.3)
## dplyr 1.1.4 2023-11-17 [1] CRAN (R 4.4.3)
## dqrng 0.3.2 2023-11-29 [1] CRAN (R 4.4.3)
## edgeR 4.4.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## ellipsis 0.3.2 2021-04-29 [1] CRAN (R 4.4.3)
## evaluate 1.0.5 2025-08-27 [1] CRAN (R 4.4.3)
## farver 2.1.2 2024-05-13 [1] CRAN (R 4.4.3)
## fastDummies 1.7.5 2025-01-20 [1] CRAN (R 4.4.3)
## fastmap 1.2.0 2024-05-15 [1] CRAN (R 4.4.3)
## filelock 1.0.3 2023-12-11 [1] CRAN (R 4.4.3)
## fitdistrplus 1.2-4 2025-07-03 [1] CRAN (R 4.4.3)
## forcats * 1.0.1 2025-09-25 [1] CRAN (R 4.4.3)
## foreach 1.5.2 2022-02-02 [1] CRAN (R 4.4.3)
## fs 1.6.6 2025-04-12 [1] CRAN (R 4.4.3)
## future * 1.68.0 2025-11-17 [1] CRAN (R 4.4.3)
## future.apply 1.20.0 2025-06-06 [1] CRAN (R 4.4.3)
## generics 0.1.4 2025-05-09 [1] CRAN (R 4.4.3)
## GenomeInfoDb * 1.42.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## GenomeInfoDbData 1.2.13 2026-03-05 [1] Bioconductor
## GenomicRanges * 1.58.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## GetoptLong 1.0.5 2020-12-15 [1] CRAN (R 4.4.3)
## getPass 0.2-4 2023-12-10 [1] CRAN (R 4.4.3)
## ggbeeswarm * 0.7.2 2023-04-29 [1] CRAN (R 4.4.3)
## ggh4x * 0.3.1 2025-05-30 [1] CRAN (R 4.4.3)
## ggplot.multistats * 1.0.1 2024-09-25 [1] CRAN (R 4.4.3)
## ggplot2 * 4.0.1 2025-11-14 [1] CRAN (R 4.4.3)
## ggrepel * 0.9.6 2024-09-07 [1] CRAN (R 4.4.3)
## ggridges 0.5.7 2025-08-27 [1] CRAN (R 4.4.3)
## git2r 0.35.0 2024-10-20 [1] CRAN (R 4.4.3)
## GlobalOptions 0.1.2 2020-06-10 [1] CRAN (R 4.4.3)
## globals 0.18.0 2025-05-08 [1] CRAN (R 4.4.3)
## glue 1.8.0 2024-09-30 [1] CRAN (R 4.4.3)
## goftest 1.2-3 2021-10-07 [1] CRAN (R 4.4.3)
## gridExtra 2.3 2017-09-09 [1] CRAN (R 4.4.3)
## gtable 0.3.6 2024-10-25 [1] CRAN (R 4.4.3)
## harmony * 1.2.4 2025-10-10 [1] CRAN (R 4.4.3)
## hexbin 1.28.5 2024-11-13 [1] CRAN (R 4.4.3)
## htmltools 0.5.8.1 2024-04-04 [1] CRAN (R 4.4.3)
## htmlwidgets 1.6.4 2023-12-06 [1] CRAN (R 4.4.3)
## httpuv 1.6.16 2025-04-16 [1] CRAN (R 4.4.3)
## httr 1.4.7 2023-08-15 [1] CRAN (R 4.4.3)
## ica 1.0-3 2022-07-08 [1] CRAN (R 4.4.3)
## igraph 2.1.4 2025-01-23 [1] CRAN (R 4.4.3)
## IRanges * 2.40.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## irlba 2.3.5.1 2022-10-03 [1] CRAN (R 4.4.3)
## iterators 1.0.14 2022-02-05 [1] CRAN (R 4.4.3)
## jquerylib 0.1.4 2021-04-26 [1] CRAN (R 4.4.3)
## jsonlite 2.0.0 2025-03-27 [1] CRAN (R 4.4.3)
## KernSmooth 2.23-26 2025-01-01 [1] CRAN (R 4.4.3)
## knitr 1.50 2025-03-16 [1] CRAN (R 4.4.3)
## later 1.4.4 2025-08-27 [1] CRAN (R 4.4.3)
## lattice 0.22-7 2025-04-02 [1] CRAN (R 4.4.3)
## lazyeval 0.2.2 2019-03-15 [1] CRAN (R 4.4.3)
## lifecycle 1.0.4 2023-11-07 [1] CRAN (R 4.4.3)
## limma 3.62.1 2024-11-03 [1] Bioconductor 3.20 (R 4.4.2)
## listenv 0.10.0 2025-11-02 [1] CRAN (R 4.4.3)
## lmtest 0.9-40 2022-03-21 [1] CRAN (R 4.4.3)
## locfit 1.5-9.12 2025-03-05 [1] CRAN (R 4.4.3)
## magrittr * 2.0.4 2025-09-12 [1] CRAN (R 4.4.3)
## MASS 7.3-65 2025-02-28 [1] CRAN (R 4.4.3)
## Matrix * 1.7-4 2025-08-28 [1] CRAN (R 4.4.3)
## MatrixGenerics * 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## matrixStats * 1.5.0 2025-01-07 [1] CRAN (R 4.4.3)
## memoise 2.0.1 2021-11-26 [1] CRAN (R 4.4.3)
## metapod 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## mgcv 1.9-4 2025-11-07 [1] CRAN (R 4.4.3)
## mime 0.13 2025-03-17 [1] CRAN (R 4.4.3)
## miniUI 0.1.2 2025-04-17 [1] CRAN (R 4.4.3)
## nlme 3.1-168 2025-03-31 [1] CRAN (R 4.4.3)
## otel 0.2.0 2025-08-29 [1] CRAN (R 4.4.3)
## parallelly 1.45.1 2025-07-24 [1] CRAN (R 4.4.3)
## patchwork * 1.3.2 2025-08-25 [1] CRAN (R 4.4.3)
## pbapply 1.7-4 2025-07-20 [1] CRAN (R 4.4.3)
## pillar 1.11.1 2025-09-17 [1] CRAN (R 4.4.3)
## pkgbuild 1.4.8 2025-05-26 [1] CRAN (R 4.4.3)
## pkgconfig 2.0.3 2019-09-22 [1] CRAN (R 4.4.3)
## pkgload 1.4.1 2025-09-23 [1] CRAN (R 4.4.3)
## plotly 4.11.0 2025-06-19 [1] CRAN (R 4.4.3)
## plyr 1.8.9 2023-10-02 [1] CRAN (R 4.4.3)
## png 0.1-8 2022-11-29 [1] CRAN (R 4.4.3)
## polyclip 1.10-7 2024-07-23 [1] CRAN (R 4.4.3)
## processx 3.8.6 2025-02-21 [1] CRAN (R 4.4.3)
## progressr 0.18.0 2025-11-06 [1] CRAN (R 4.4.3)
## promises 1.5.0 2025-11-01 [1] CRAN (R 4.4.3)
## ps 1.9.1 2025-04-12 [1] CRAN (R 4.4.3)
## purrr 1.2.0 2025-11-04 [1] CRAN (R 4.4.3)
## R.methodsS3 1.8.2 2022-06-13 [1] CRAN (R 4.4.3)
## R.oo 1.27.1 2025-05-02 [1] CRAN (R 4.4.3)
## R.utils 2.13.0 2025-02-24 [1] CRAN (R 4.4.3)
## R6 2.6.1 2025-02-15 [1] CRAN (R 4.4.3)
## RANN 2.6.2 2024-08-25 [1] CRAN (R 4.4.3)
## RColorBrewer * 1.1-3 2022-04-03 [1] CRAN (R 4.4.3)
## Rcpp * 1.1.0 2025-07-02 [1] CRAN (R 4.4.3)
## RcppAnnoy 0.0.22 2024-01-23 [1] CRAN (R 4.4.3)
## RcppHNSW 0.6.0 2024-02-04 [1] CRAN (R 4.4.3)
## readxl * 1.4.5 2025-03-07 [1] CRAN (R 4.4.3)
## registry 0.5-1 2019-03-05 [1] CRAN (R 4.4.3)
## remotes 2.5.0 2024-03-17 [1] CRAN (R 4.4.3)
## reshape2 1.4.5 2025-11-12 [1] CRAN (R 4.4.3)
## reticulate 1.44.1 2025-11-14 [1] CRAN (R 4.4.3)
## rhdf5 * 2.50.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## rhdf5filters 1.18.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## Rhdf5lib 1.28.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## rjson 0.2.23 2024-09-16 [1] CRAN (R 4.4.3)
## rlang 1.1.6 2025-04-11 [1] CRAN (R 4.4.3)
## rmarkdown 2.30 2025-09-28 [1] CRAN (R 4.4.3)
## rmdformats 1.0.4 2022-05-17 [1] CRAN (R 4.4.3)
## ROCR 1.0-11 2020-05-02 [1] CRAN (R 4.4.3)
## rprojroot 2.1.1 2025-08-26 [1] CRAN (R 4.4.3)
## RSpectra 0.16-2 2024-07-18 [1] CRAN (R 4.4.3)
## rstudioapi 0.17.1 2024-10-22 [1] CRAN (R 4.4.3)
## rsvd 1.0.5 2021-04-16 [1] CRAN (R 4.4.1)
## Rtsne 0.17 2023-12-07 [1] CRAN (R 4.4.3)
## S4Arrays 1.6.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## S4Vectors * 0.44.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## S7 0.2.1 2025-11-14 [1] CRAN (R 4.4.3)
## sass 0.4.10 2025-04-11 [1] CRAN (R 4.4.3)
## ScaledMatrix 1.14.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## scales * 1.4.0 2025-04-24 [1] CRAN (R 4.4.3)
## scattermore 1.2 2023-06-12 [1] CRAN (R 4.4.3)
## scran * 1.34.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## sctransform 0.4.2 2025-04-30 [1] CRAN (R 4.4.3)
## scuttle * 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## seriation * 1.5.8 2025-08-20 [1] CRAN (R 4.4.3)
## sessioninfo 1.2.3 2025-02-05 [1] CRAN (R 4.4.3)
## Seurat * 5.3.1 2025-10-29 [1] CRAN (R 4.4.3)
## SeuratObject * 5.2.0 2025-08-27 [1] CRAN (R 4.4.3)
## shape 1.4.6.1 2024-02-23 [1] CRAN (R 4.4.3)
## shiny 1.11.1 2025-07-03 [1] CRAN (R 4.4.3)
## SingleCellExperiment * 1.28.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## sp * 2.2-0 2025-02-01 [1] CRAN (R 4.4.3)
## spam 2.11-1 2025-01-20 [1] CRAN (R 4.4.3)
## SparseArray 1.6.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.3)
## spatstat.data 3.1-9 2025-10-18 [1] CRAN (R 4.4.3)
## spatstat.explore 3.6-0 2025-11-22 [1] CRAN (R 4.4.3)
## spatstat.geom 3.6-1 2025-11-20 [1] CRAN (R 4.4.3)
## spatstat.random 3.4-3 2025-11-21 [1] CRAN (R 4.4.3)
## spatstat.sparse 3.1-0 2024-06-21 [1] CRAN (R 4.4.3)
## spatstat.univar 3.1-5 2025-11-17 [1] CRAN (R 4.4.3)
## spatstat.utils 3.2-0 2025-09-20 [1] CRAN (R 4.4.3)
## statmod 1.5.1 2025-10-09 [1] CRAN (R 4.4.3)
## strex * 2.0.1 2024-10-03 [1] CRAN (R 4.4.3)
## stringi 1.8.7 2025-03-27 [1] CRAN (R 4.4.3)
## stringr * 1.6.0 2025-11-04 [1] CRAN (R 4.4.3)
## SummarizedExperiment * 1.36.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## survival 3.8-3 2024-12-17 [1] CRAN (R 4.4.3)
## tensor 1.5.1 2025-06-17 [1] CRAN (R 4.4.3)
## tibble 3.3.0 2025-06-08 [1] CRAN (R 4.4.3)
## tidyr 1.3.1 2024-01-24 [1] CRAN (R 4.4.3)
## tidyselect 1.2.1 2024-03-11 [1] CRAN (R 4.4.3)
## TSP 1.2.6 2025-11-27 [1] CRAN (R 4.4.3)
## UCSC.utils 1.2.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## usethis 3.2.1 2025-09-06 [1] CRAN (R 4.4.3)
## uwot * 0.2.4 2025-11-10 [1] CRAN (R 4.4.3)
## vctrs 0.6.5 2023-12-01 [1] CRAN (R 4.4.3)
## vipor 0.4.7 2023-12-18 [1] CRAN (R 4.4.3)
## viridis * 0.6.5 2024-01-29 [1] CRAN (R 4.4.3)
## viridisLite * 0.4.2 2023-05-02 [1] CRAN (R 4.4.3)
## whisker 0.4.1 2022-12-05 [1] CRAN (R 4.4.3)
## withr 3.0.2 2024-10-28 [1] CRAN (R 4.4.3)
## workflowr * 1.7.2 2025-08-18 [1] CRAN (R 4.4.3)
## xfun 0.54 2025-10-30 [1] CRAN (R 4.4.3)
## xtable 1.8-4 2019-04-21 [1] CRAN (R 4.4.3)
## XVector 0.46.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## yaml 2.3.11 2025-11-28 [1] CRAN (R 4.4.3)
## zellkonverter * 1.16.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## zlibbioc 1.52.0 2024-10-29 [1] Bioconductor 3.20 (R 4.4.2)
## zoo 1.8-14 2025-04-10 [1] CRAN (R 4.4.3)
##
## [1] /home/macnairw/packages/scprocess/.snakemake/conda/4fef11cadd34f9d2d13a0d6139d09340_/lib/R/library
## * ── Packages attached to the search path.
##
## ──────────────────────────────────────────────────────────────────────────────