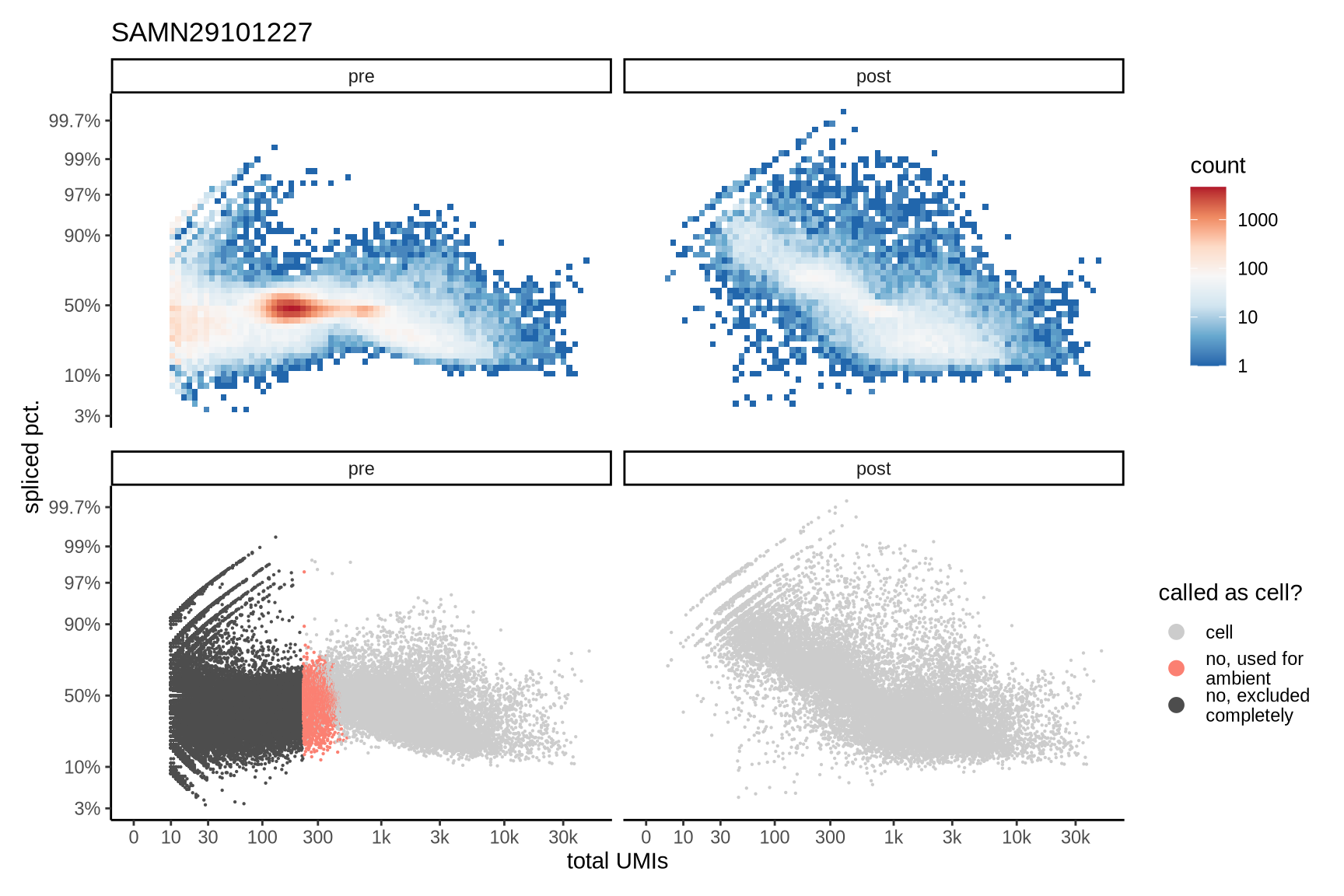
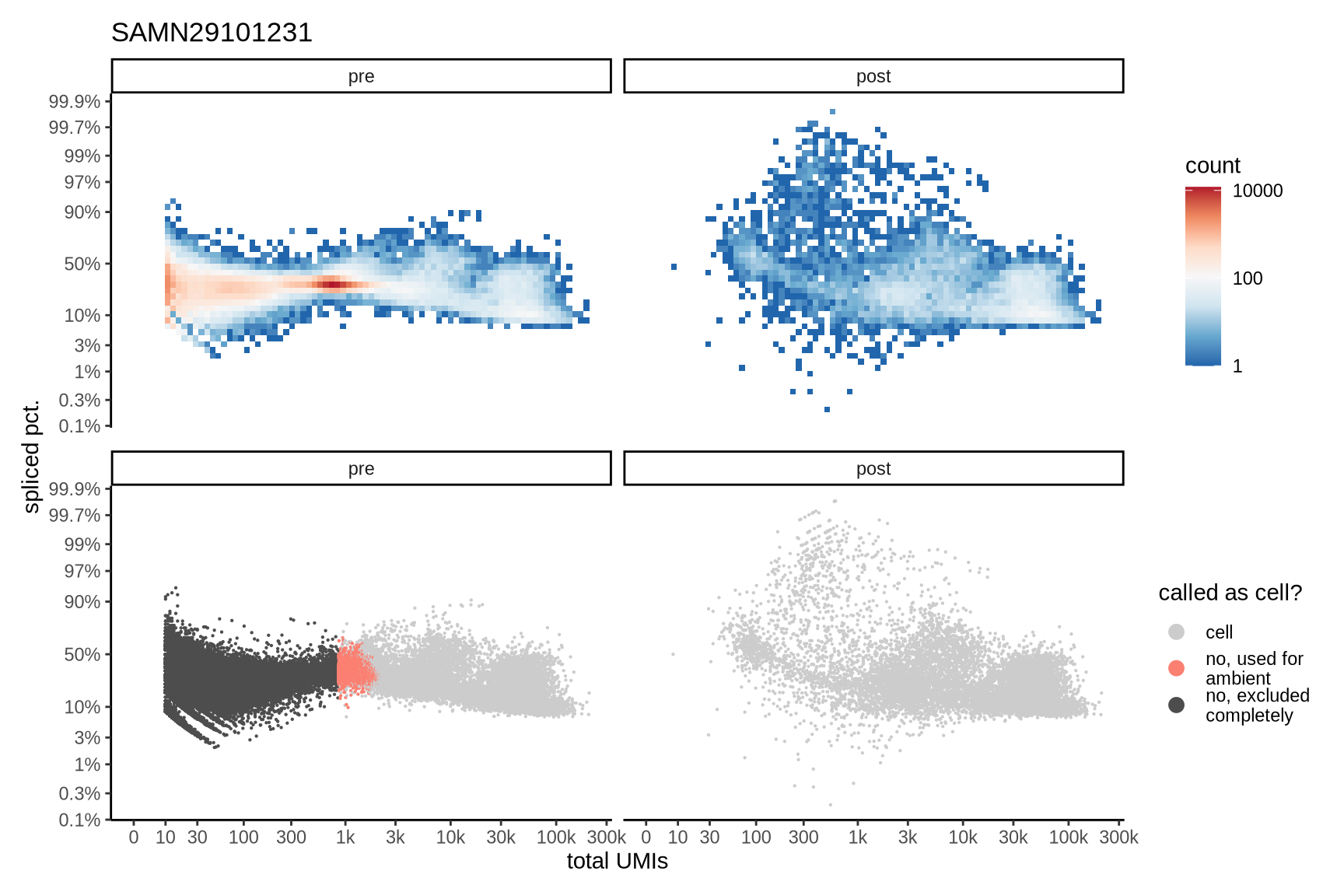
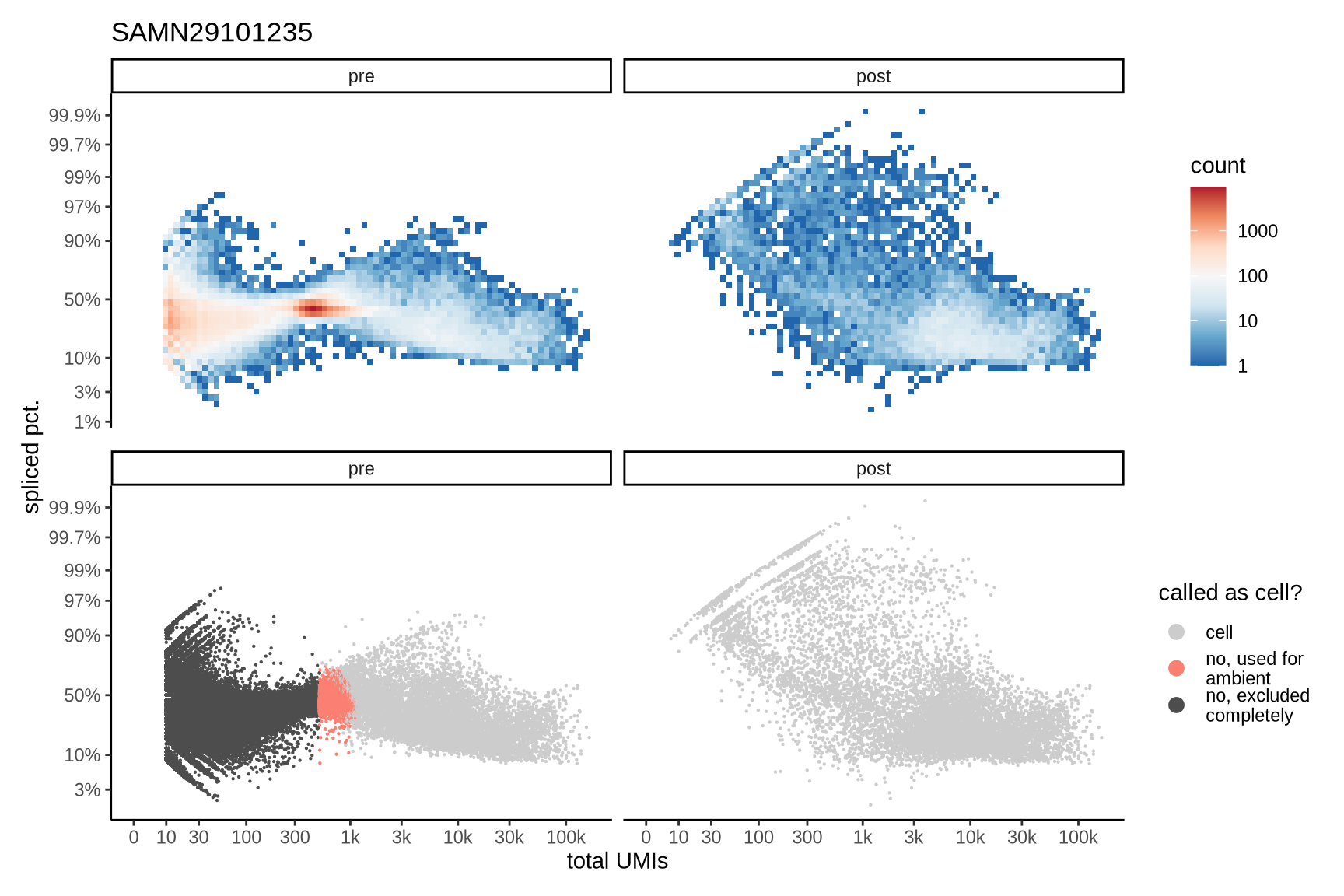
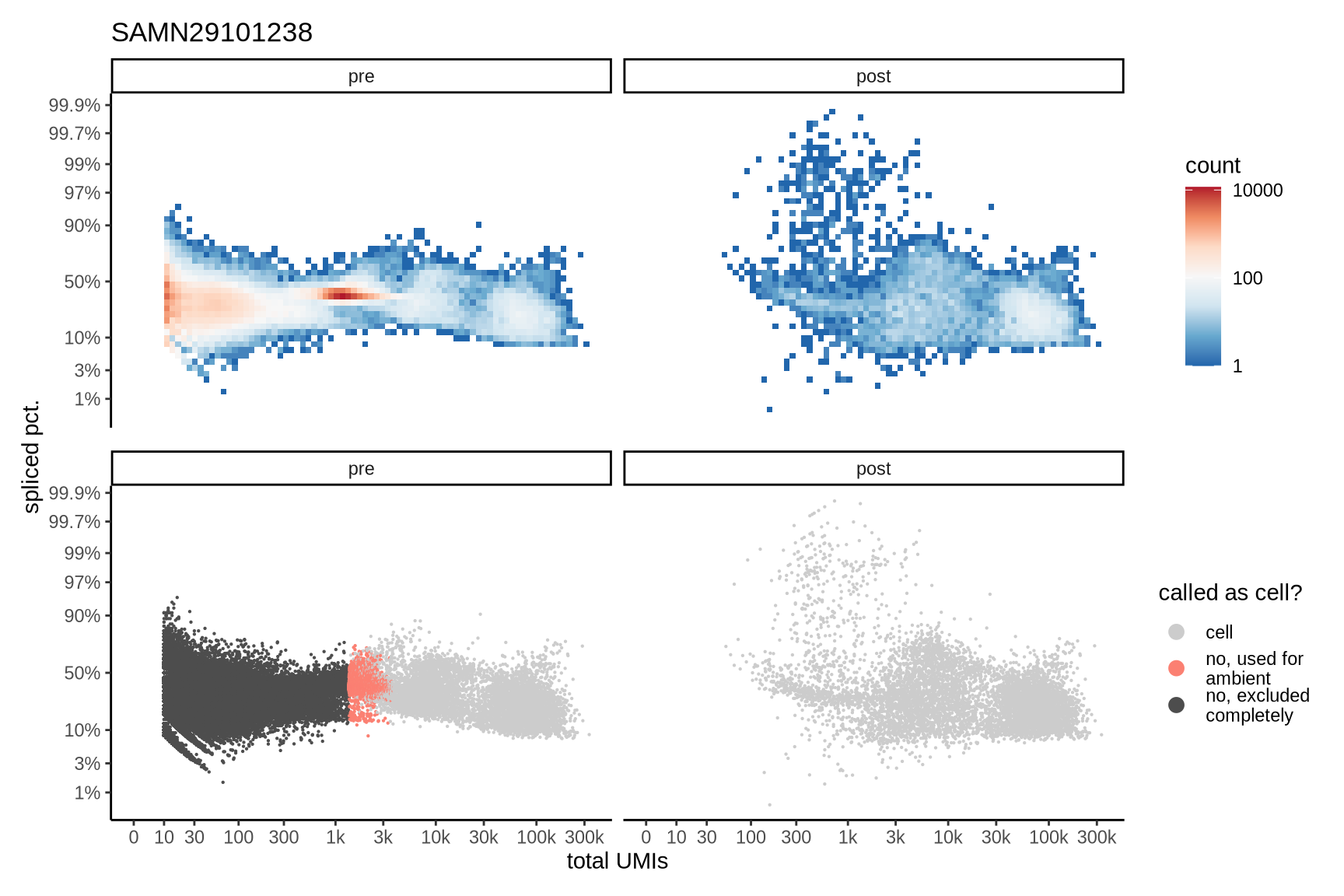
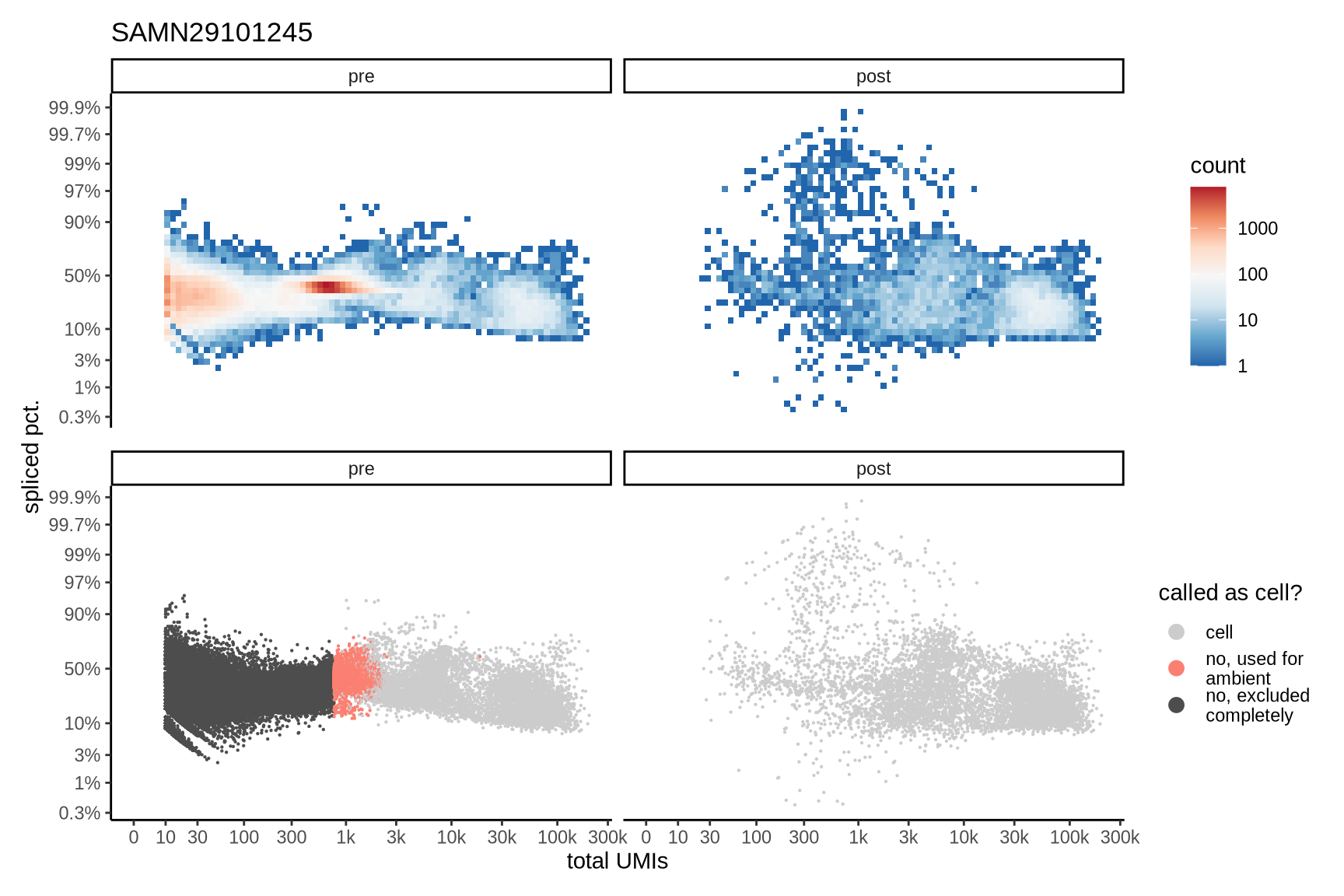
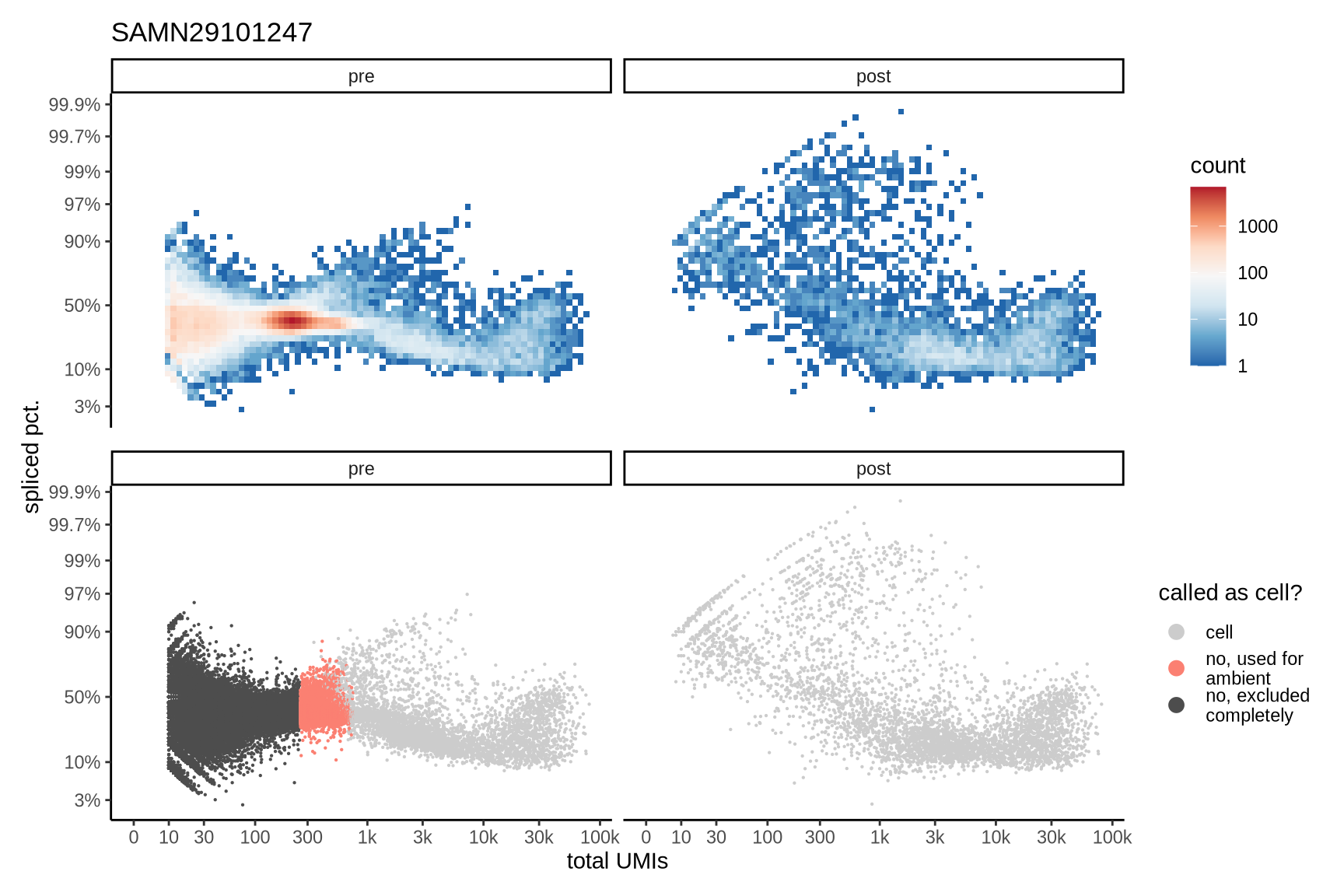
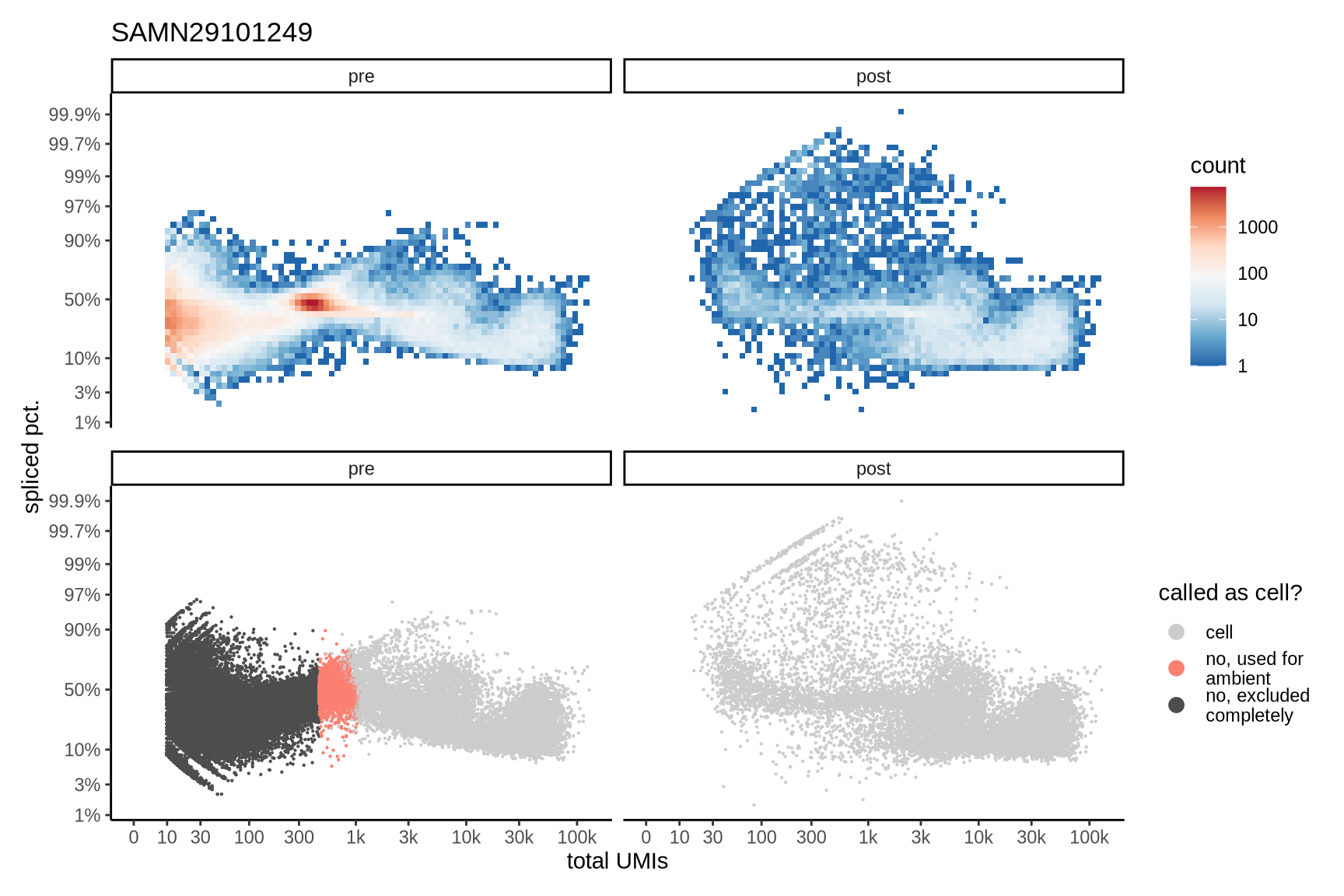
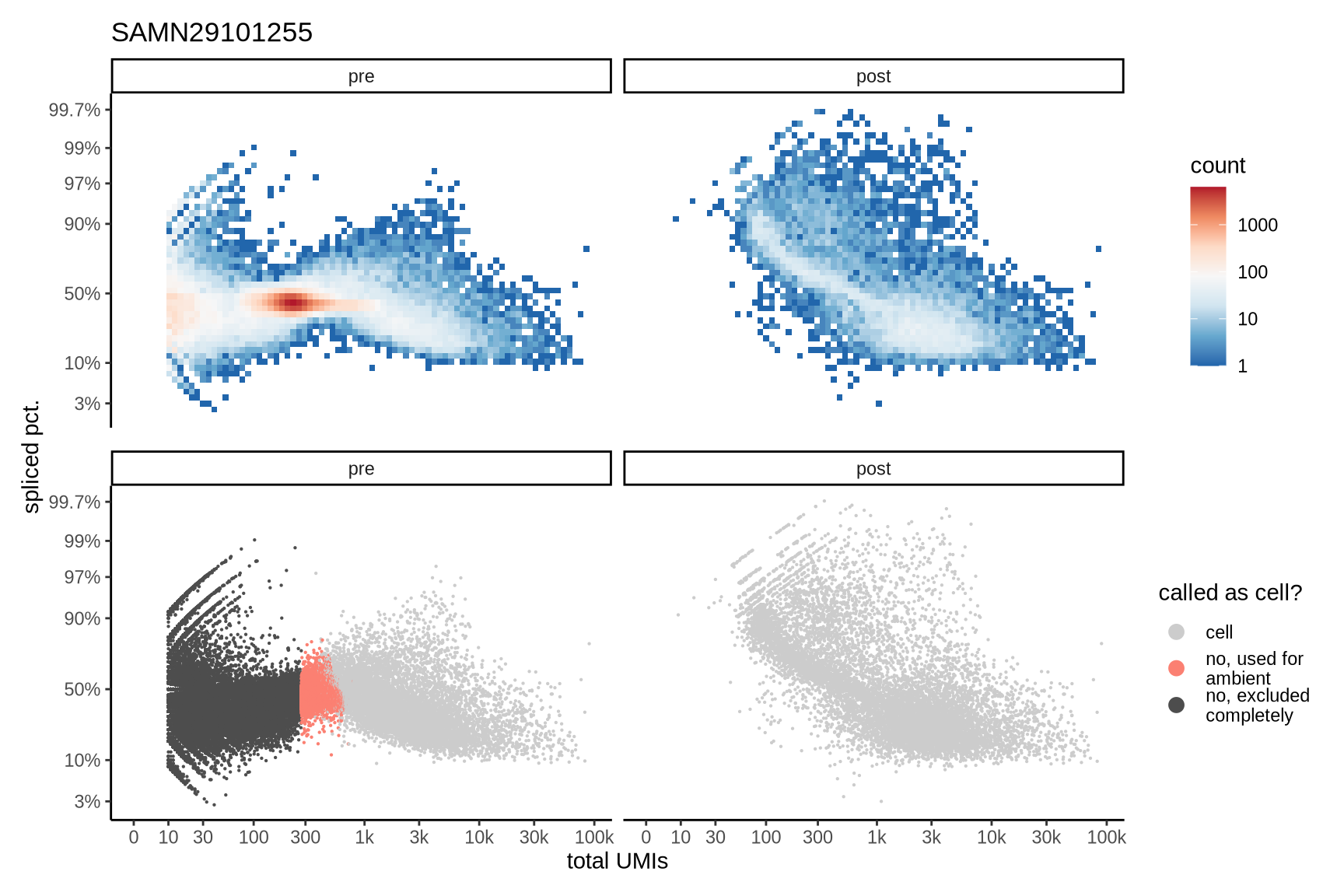
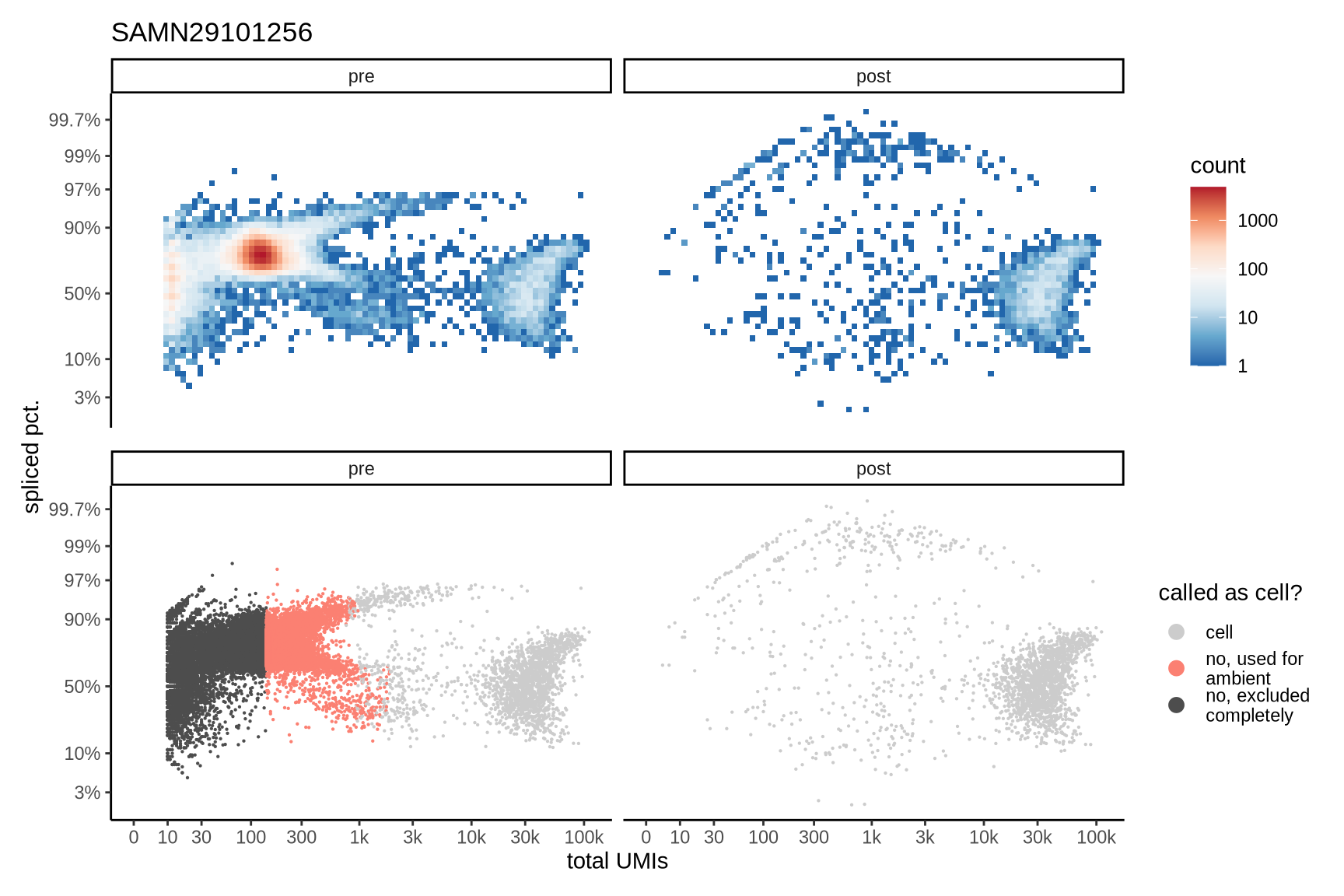
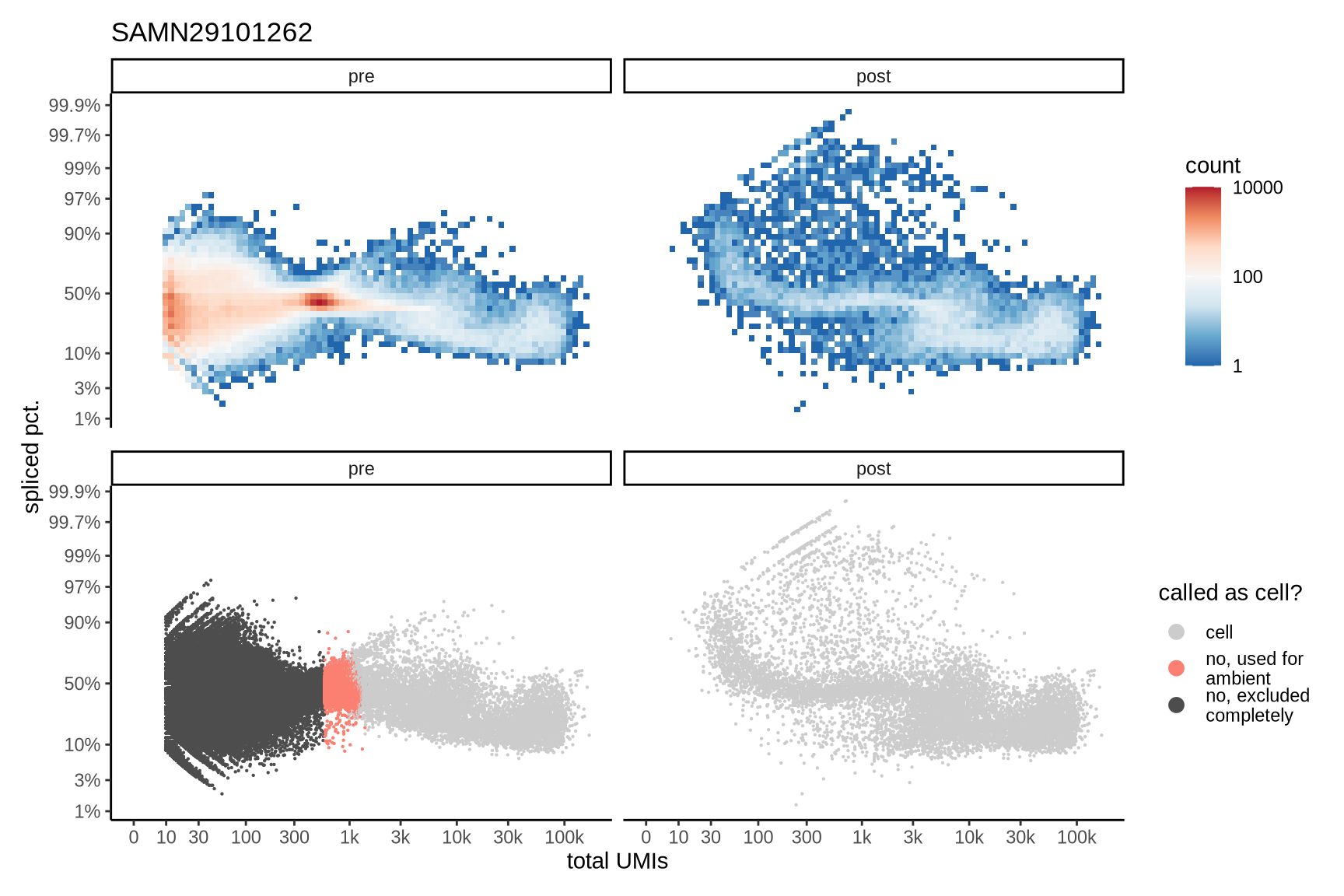
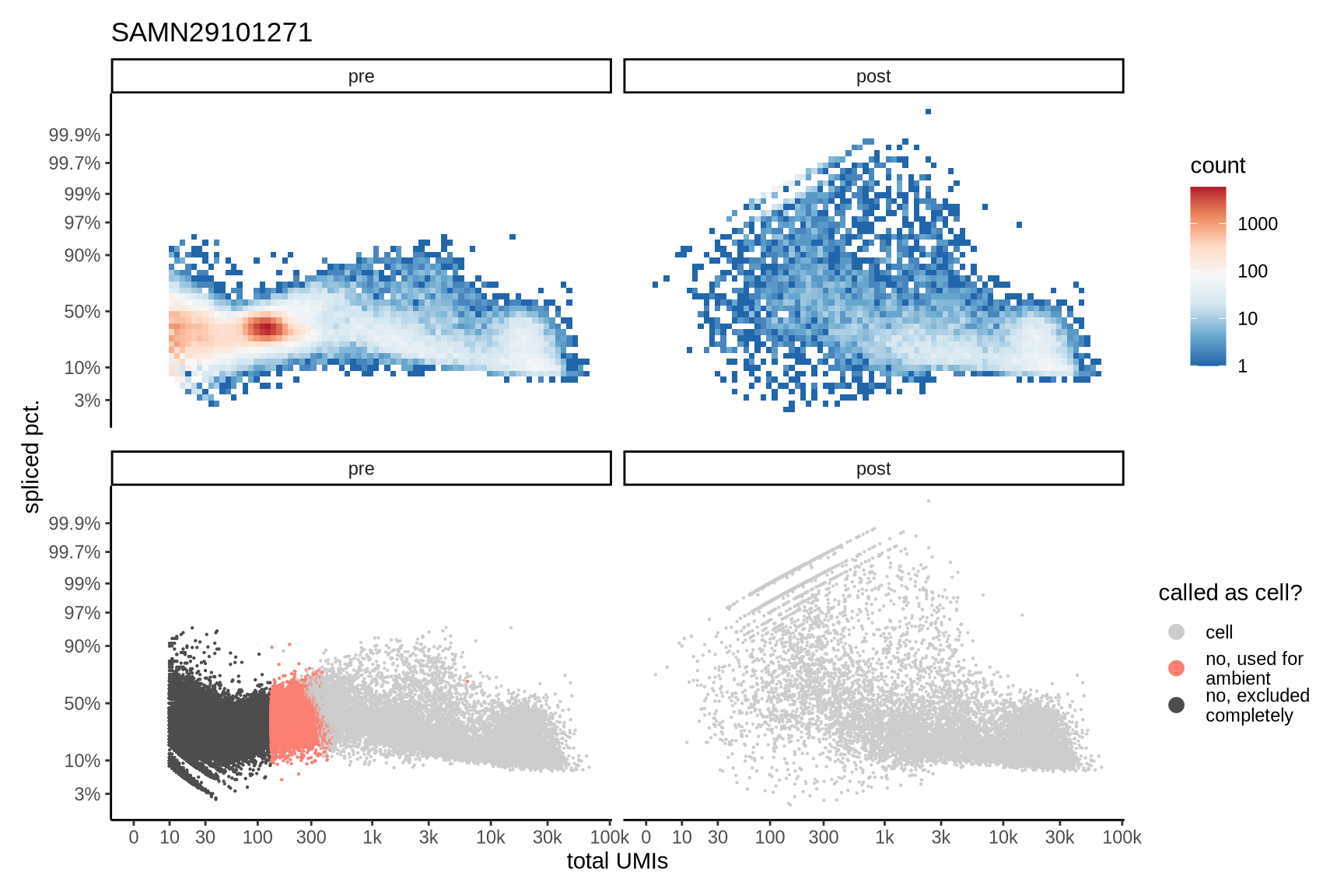
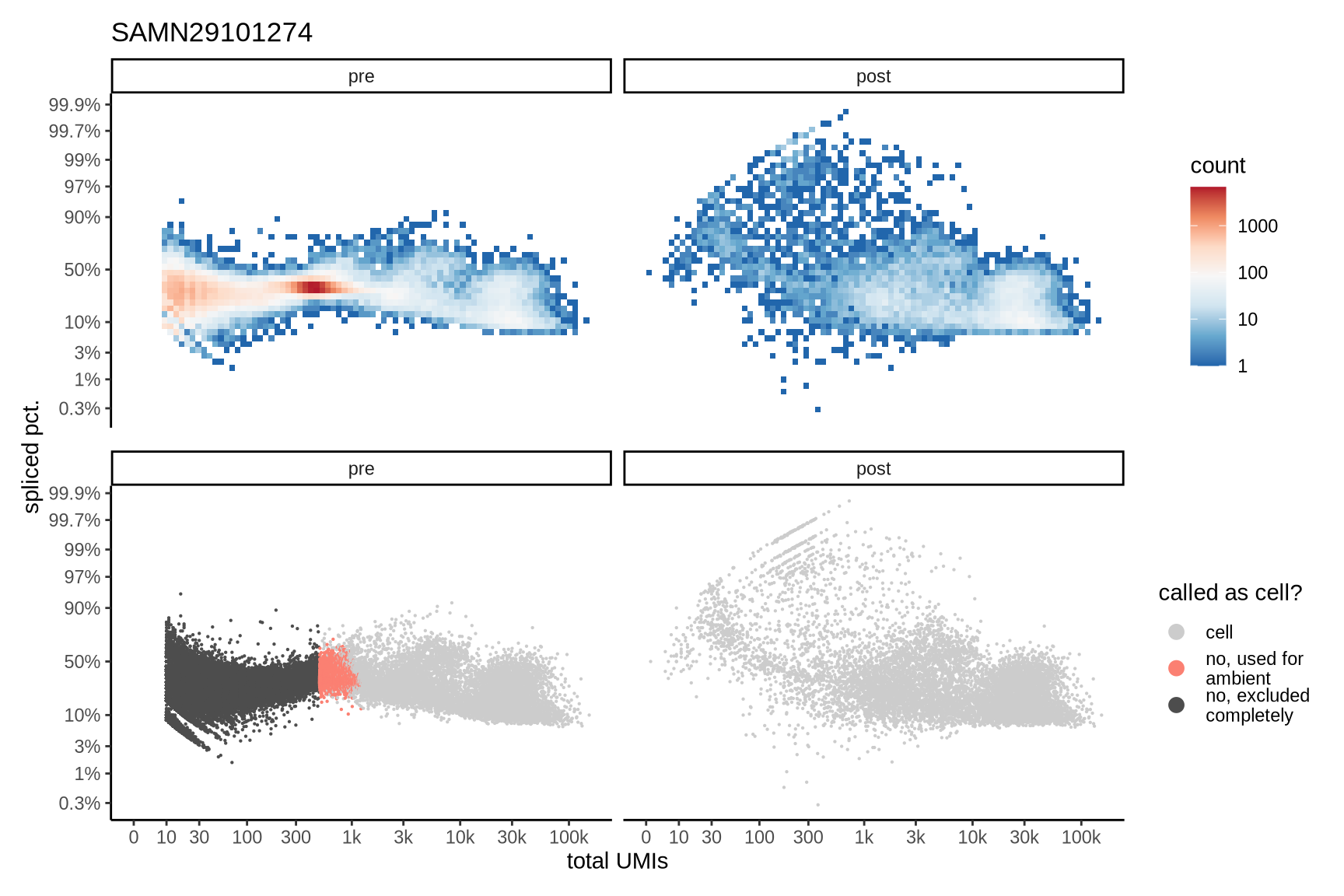
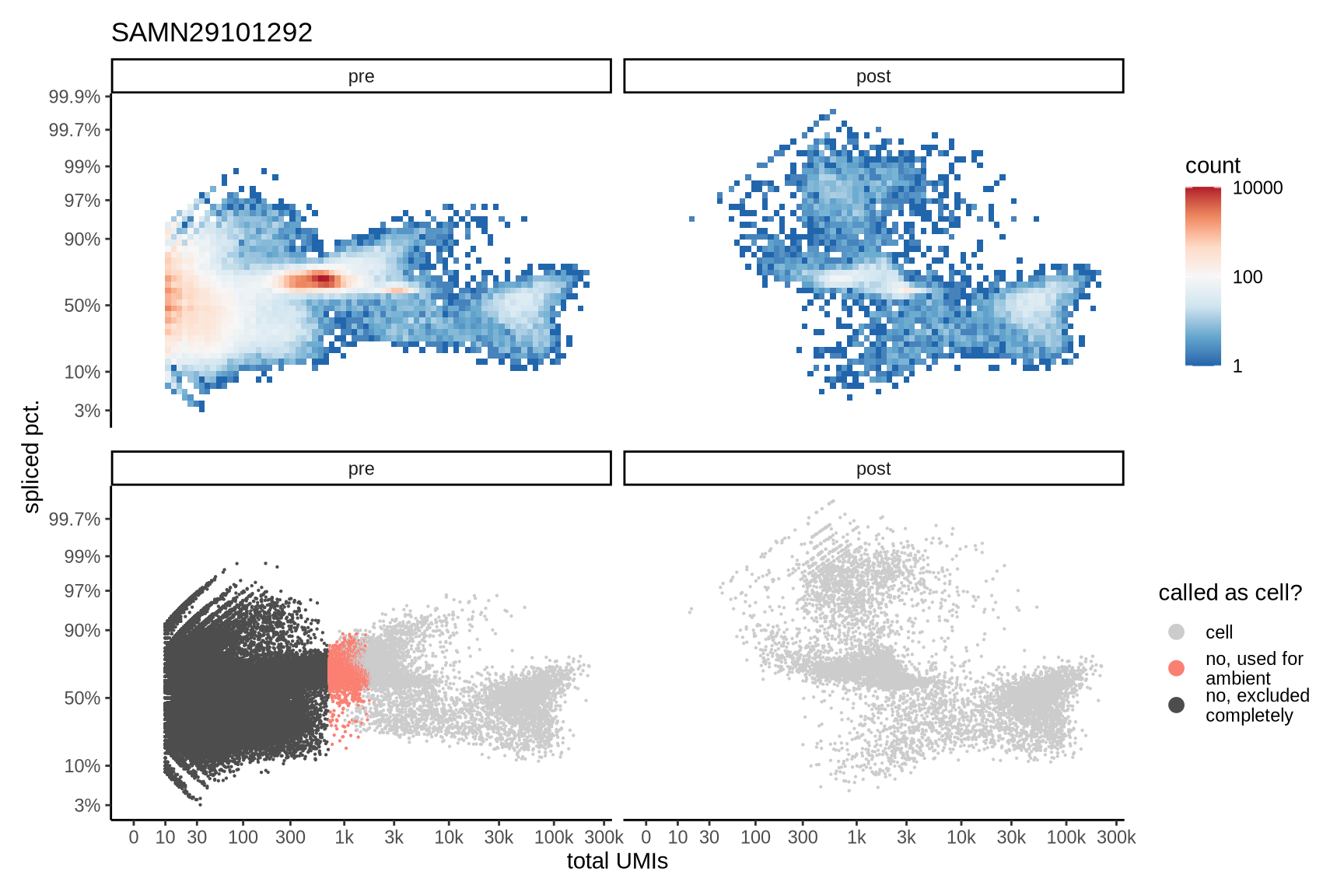
Distributions of spliced pct. in empty and cell barcodes
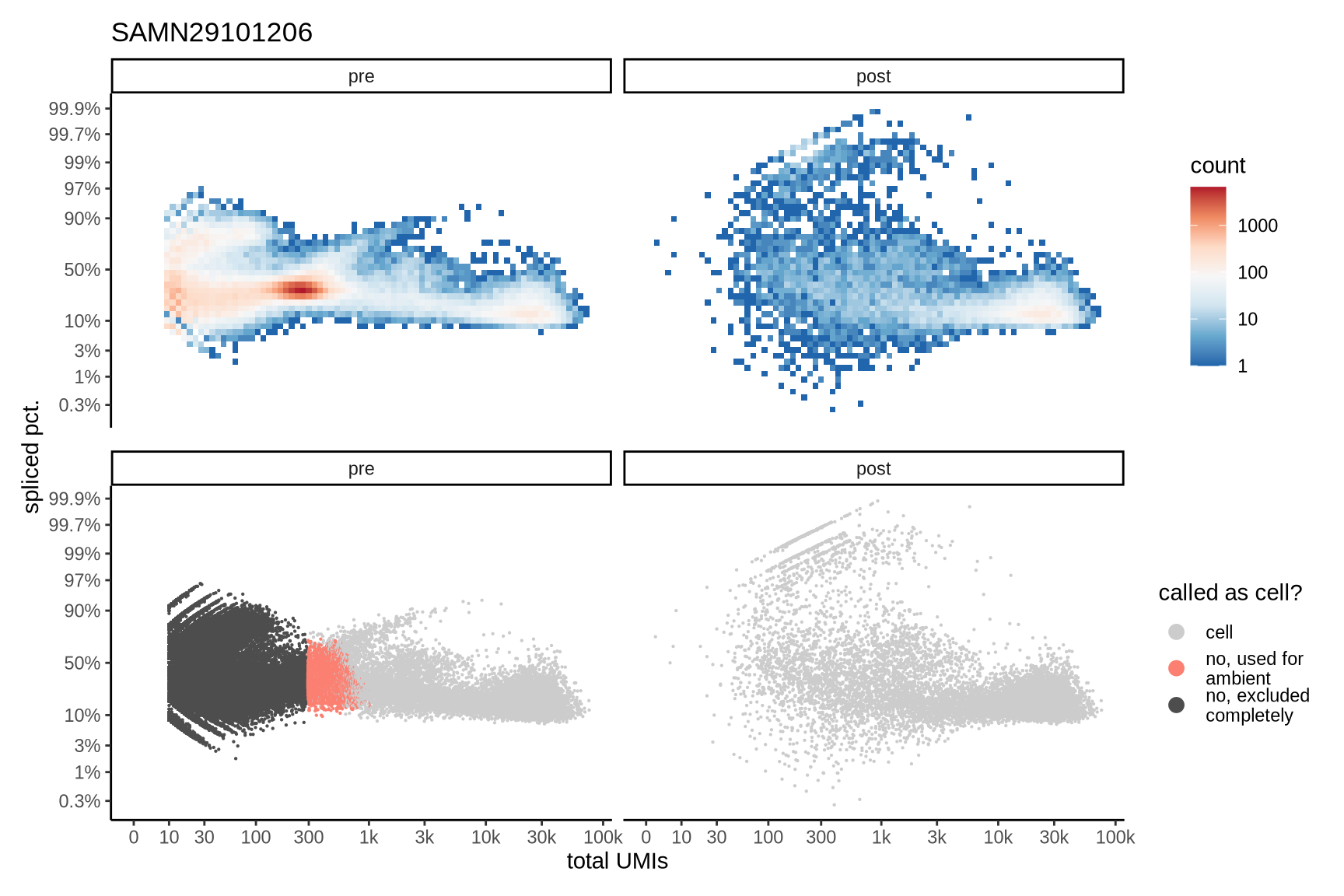
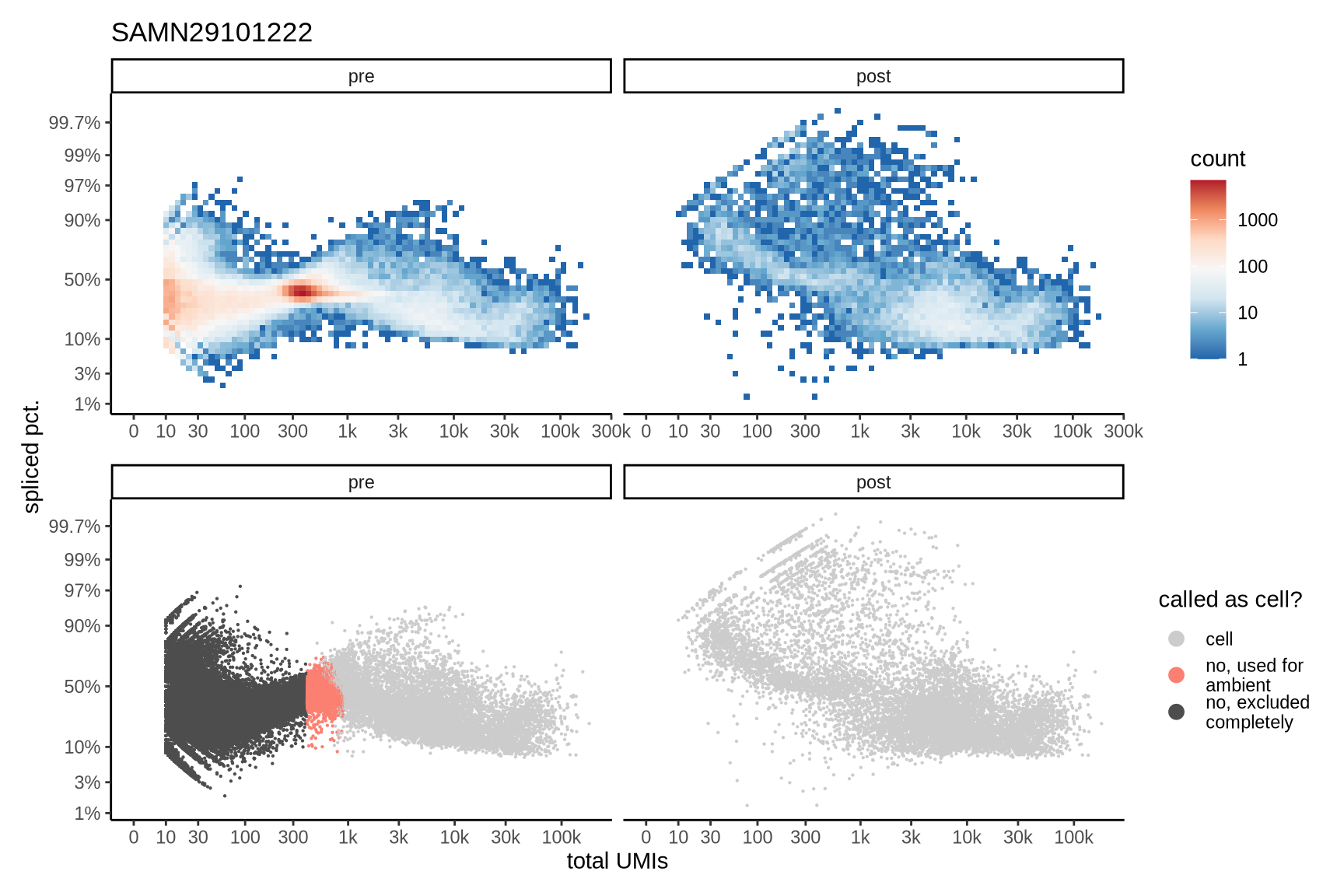
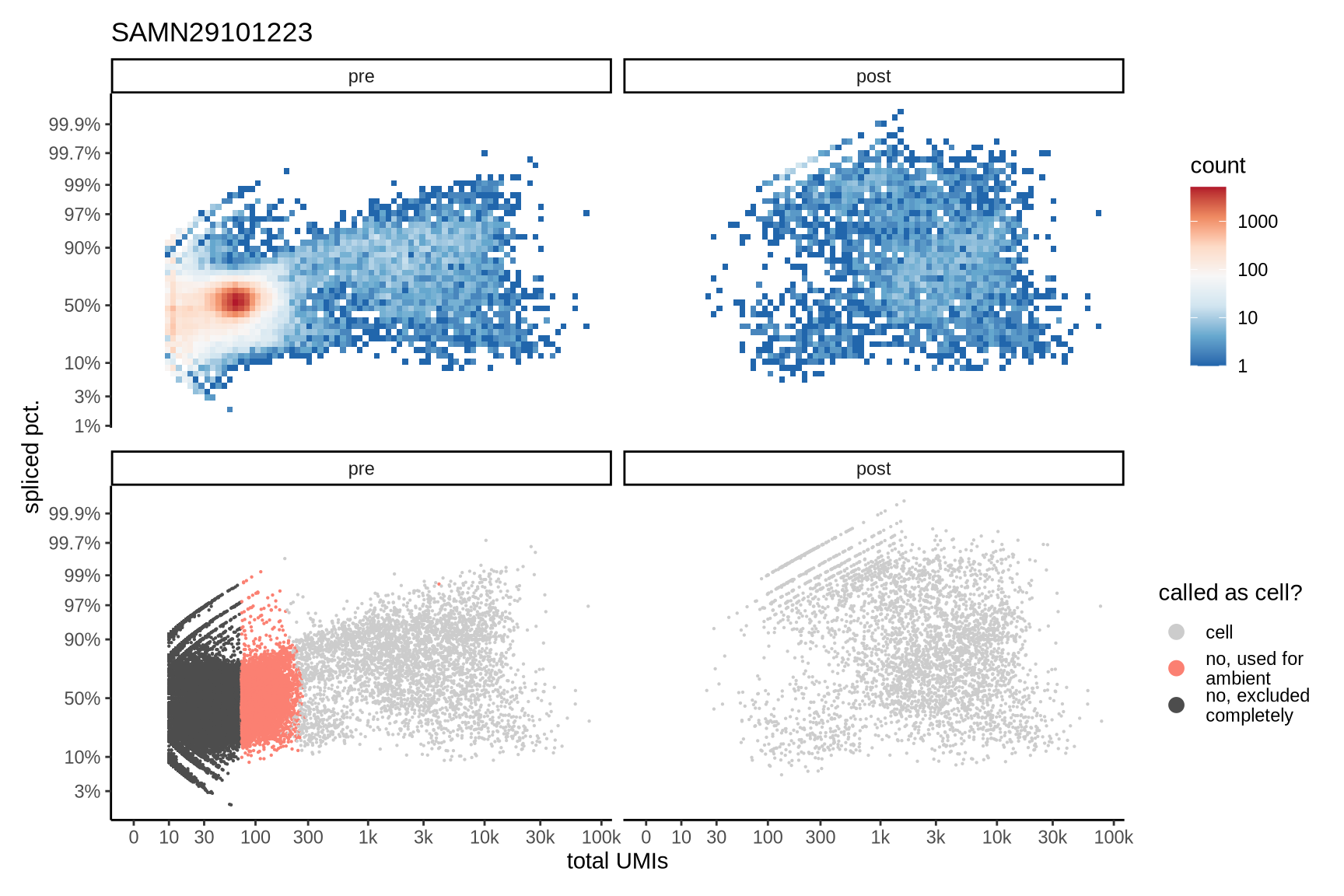
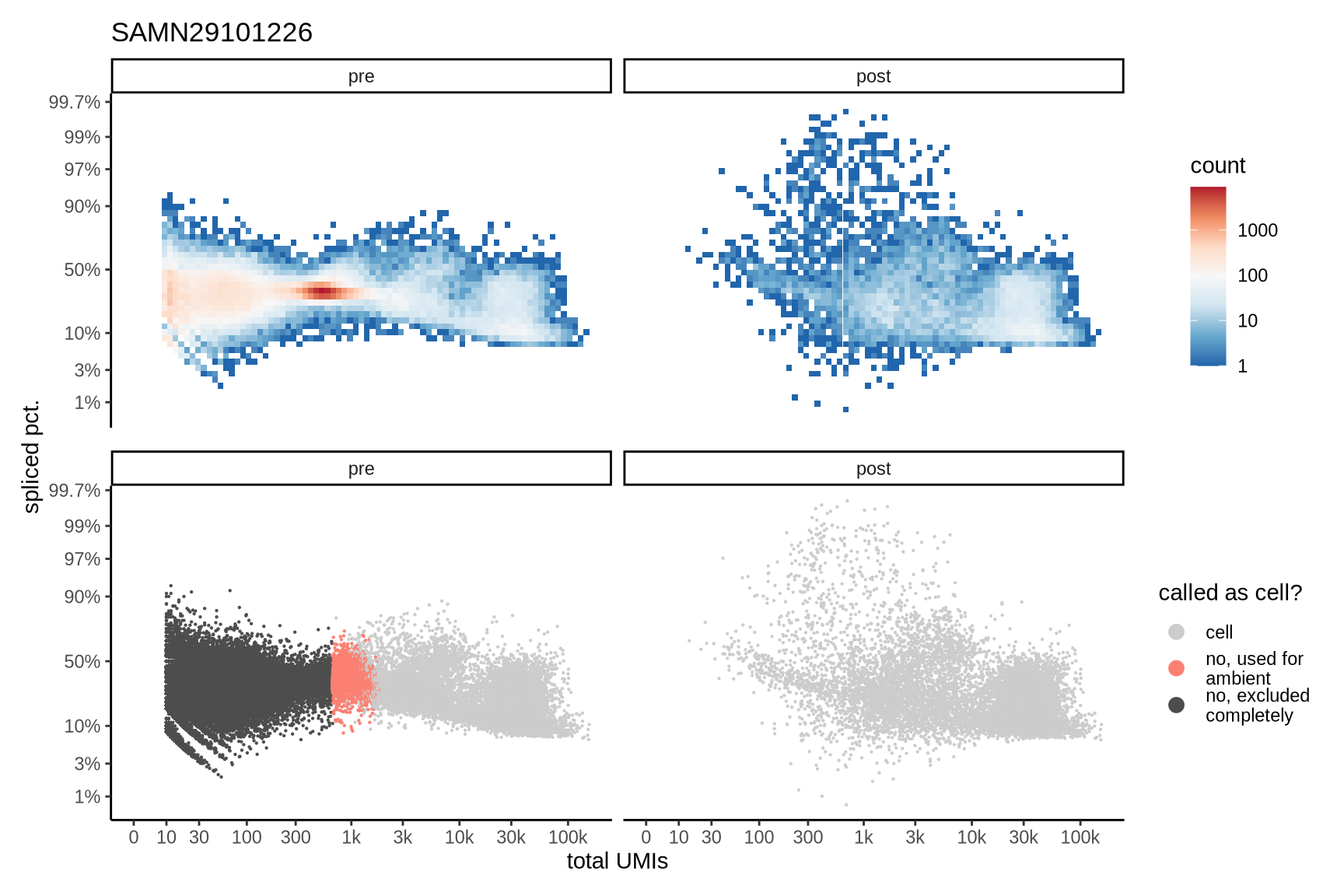
The plots show the relationship between the number of UMIs and the percentage of spliced reads, separated by sample, both before and after ambient RNA removal. Ambient RNA is typically characterized by a high proportion of spliced reads, making the spliced percentage a useful metric for evaluating the effectiveness of an ambient RNA removal method. By examining changes in spliced percentages, we can assess how well the method performed in reducing ambient RNA contamination.
for (ss in s_lvls) {
cat("### ", ss, "\n")
print( plot_spliced_vs_umis(ss, usa_dt_ls[[ss]], ok_bcs_ls[[ss]], tot_inc_ls[[ss]]) )
cat("\n\n")
}SAMN29101206
SAMN29101222
SAMN29101223
SAMN29101226
SAMN29101227
SAMN29101231
SAMN29101235
SAMN29101238
SAMN29101245
SAMN29101247
SAMN29101249
SAMN29101255
SAMN29101256
SAMN29101262
SAMN29101271
SAMN29101274
SAMN29101278
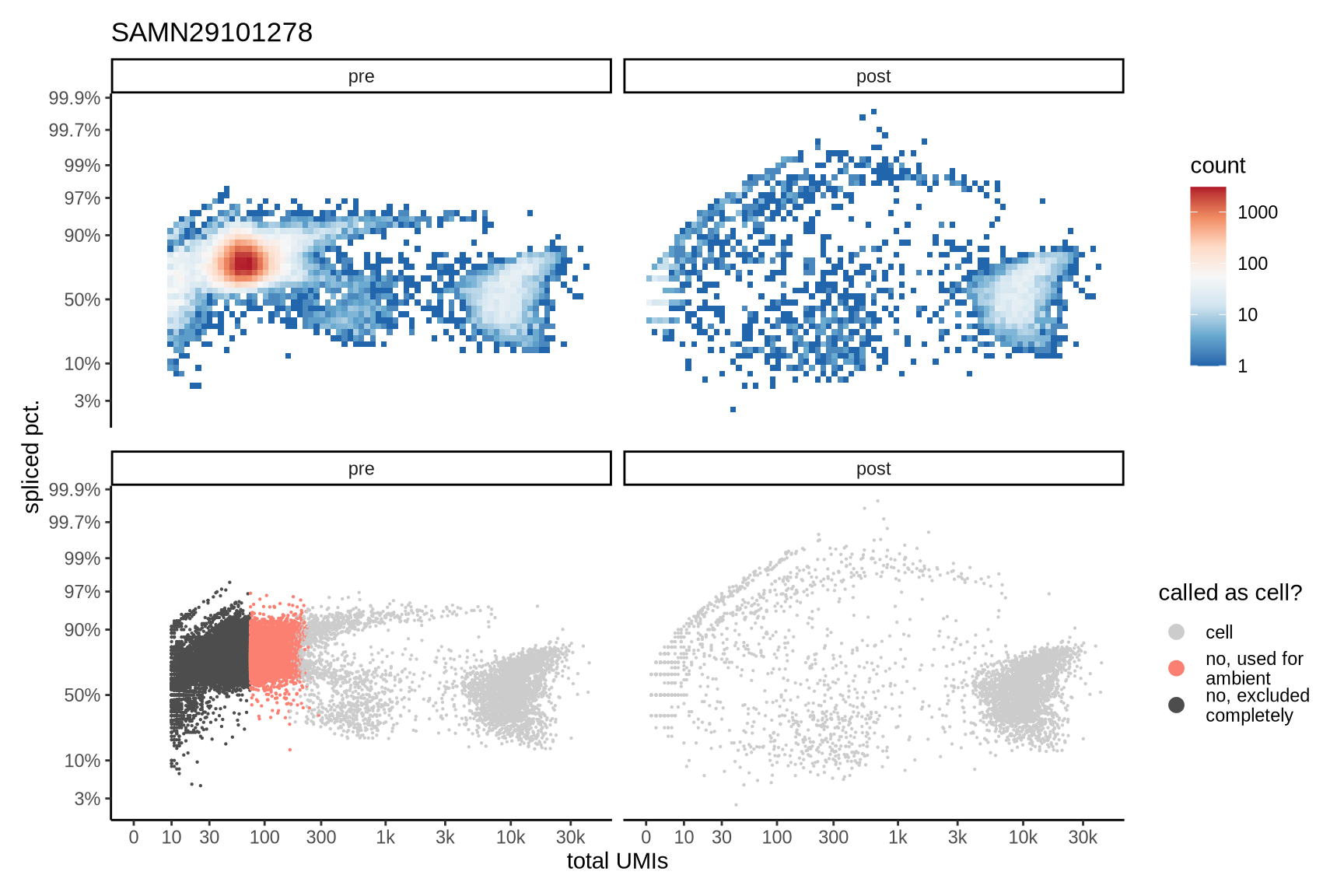
SAMN29101279
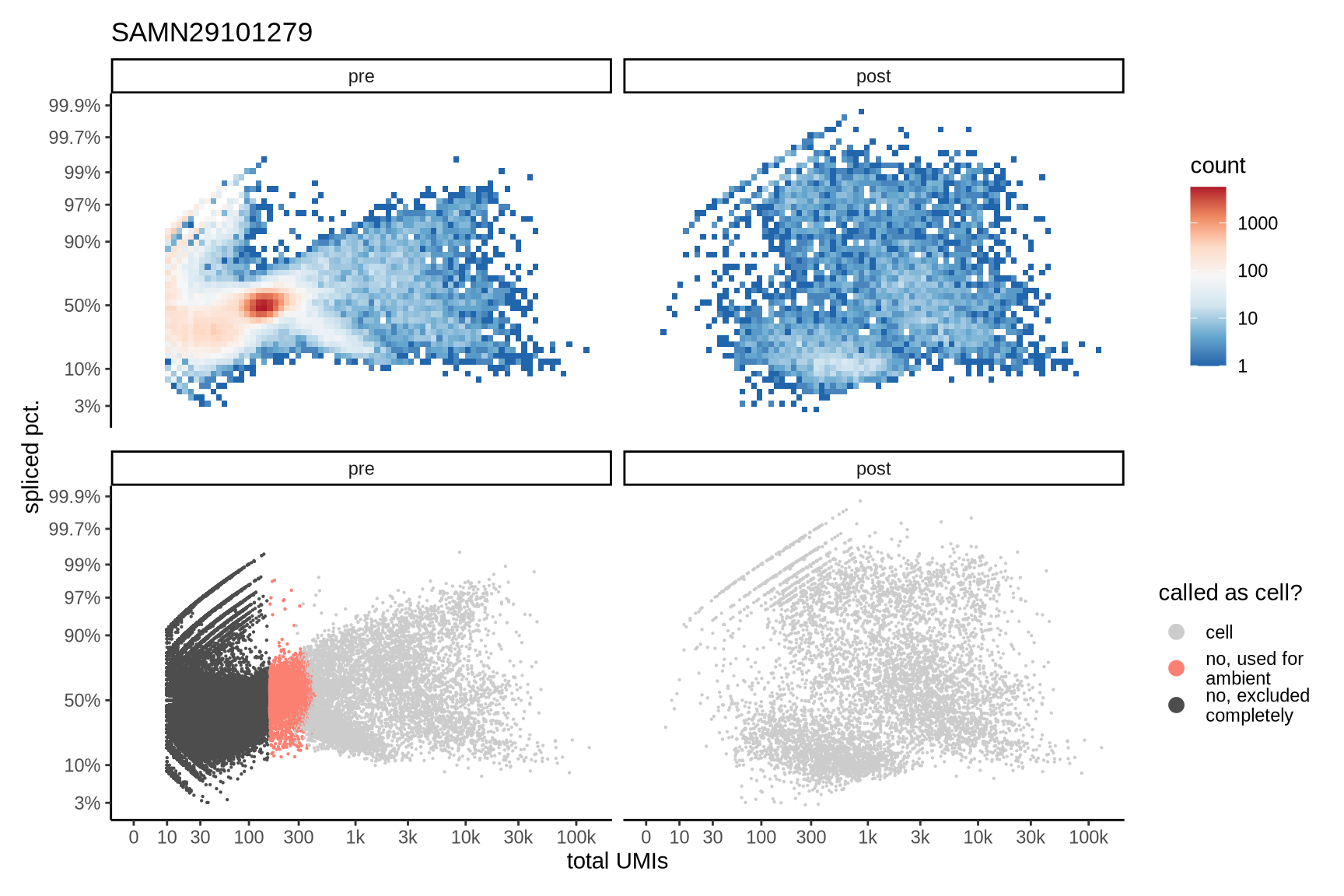
SAMN29101284
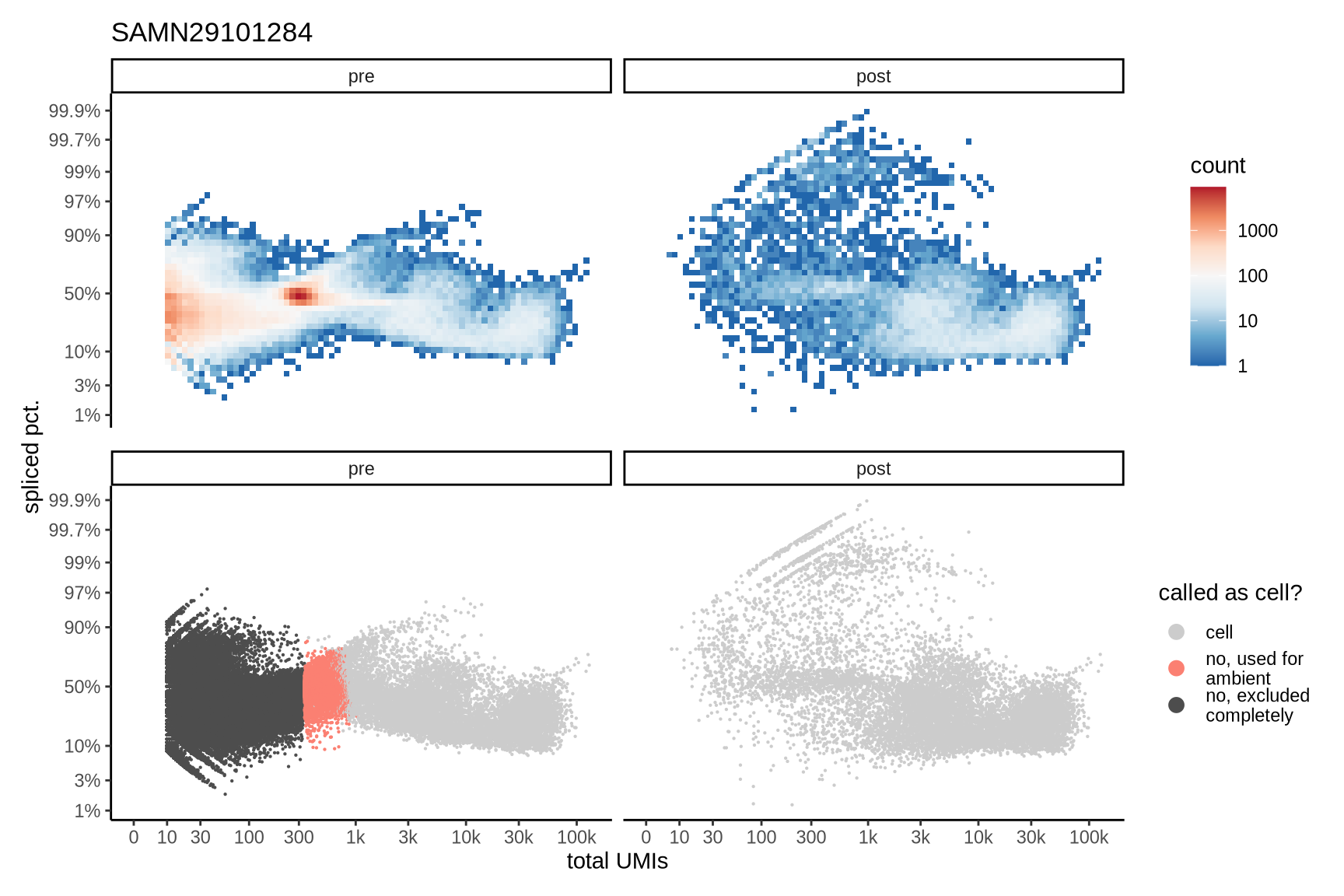
SAMN29101290
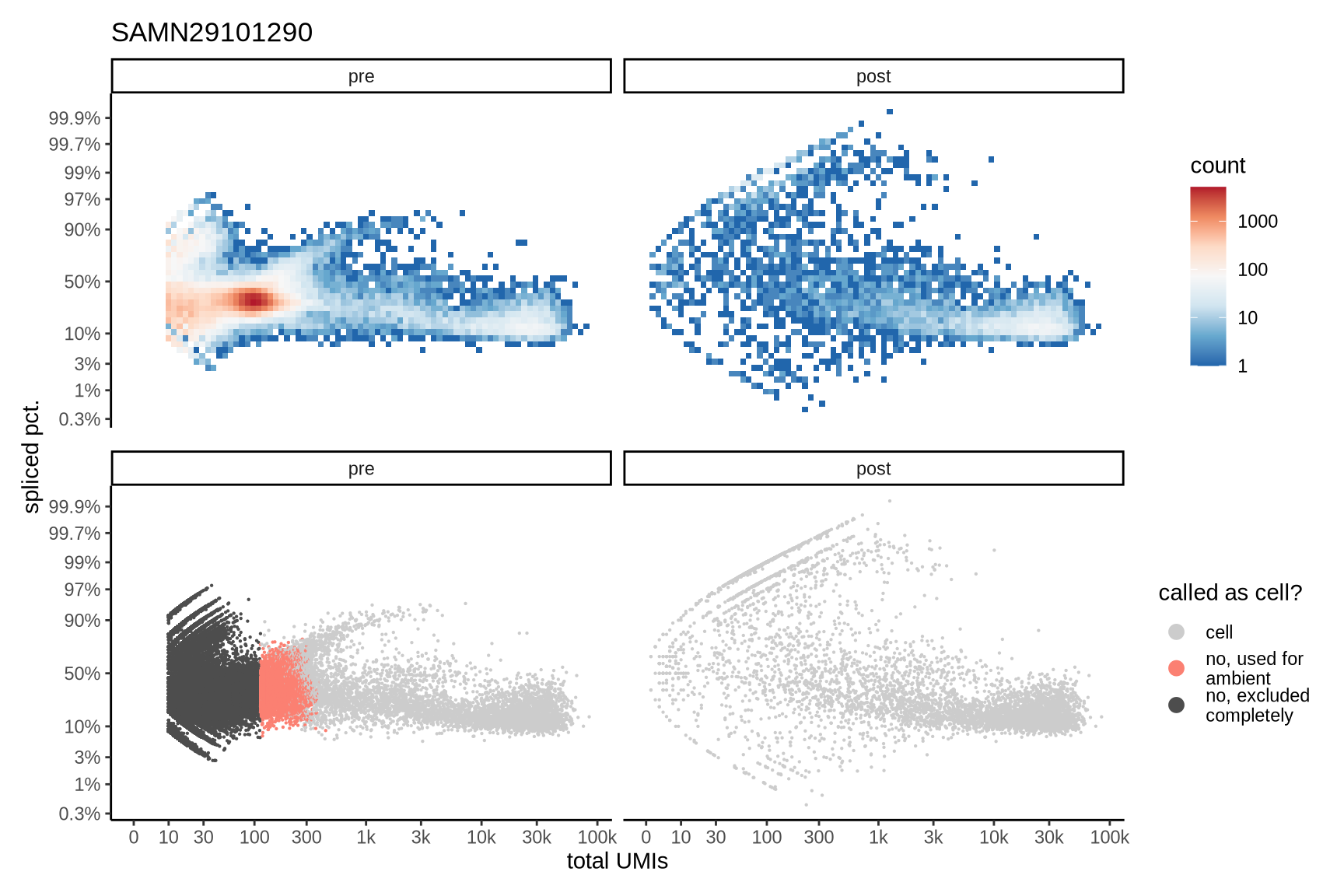
SAMN29101292
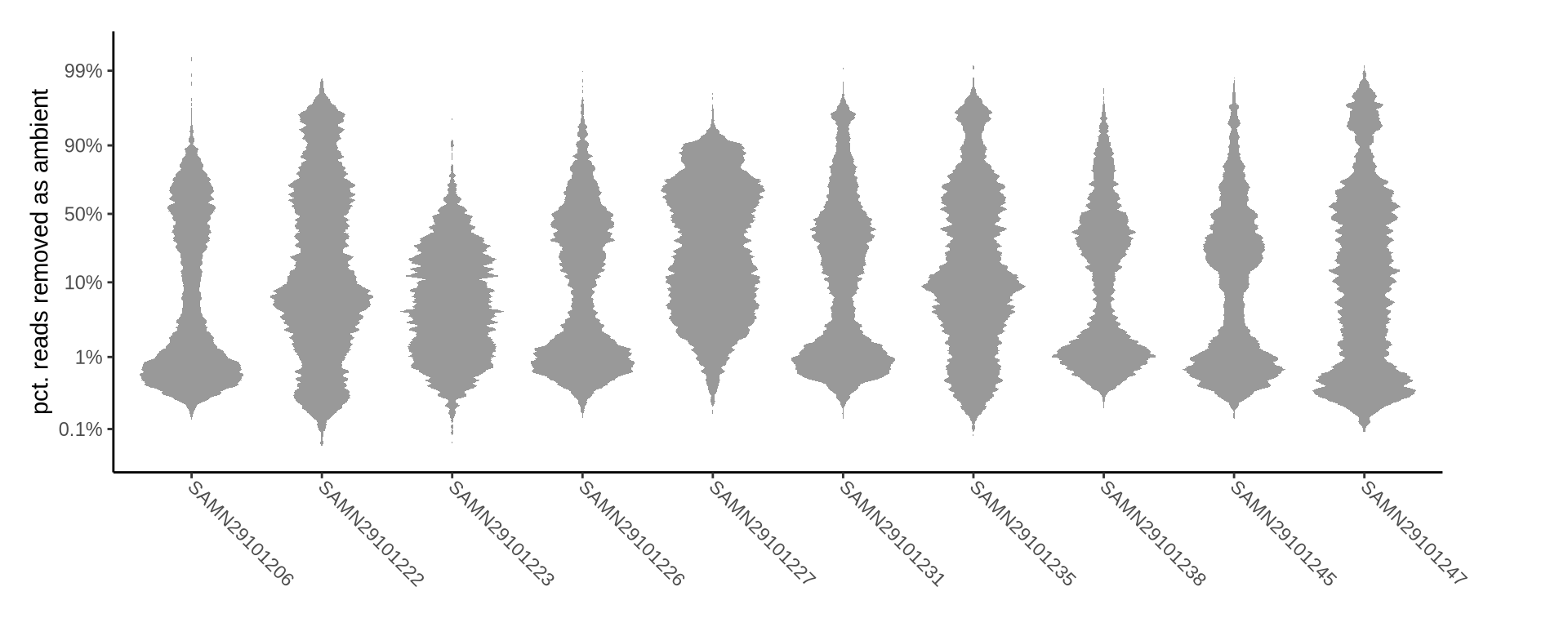
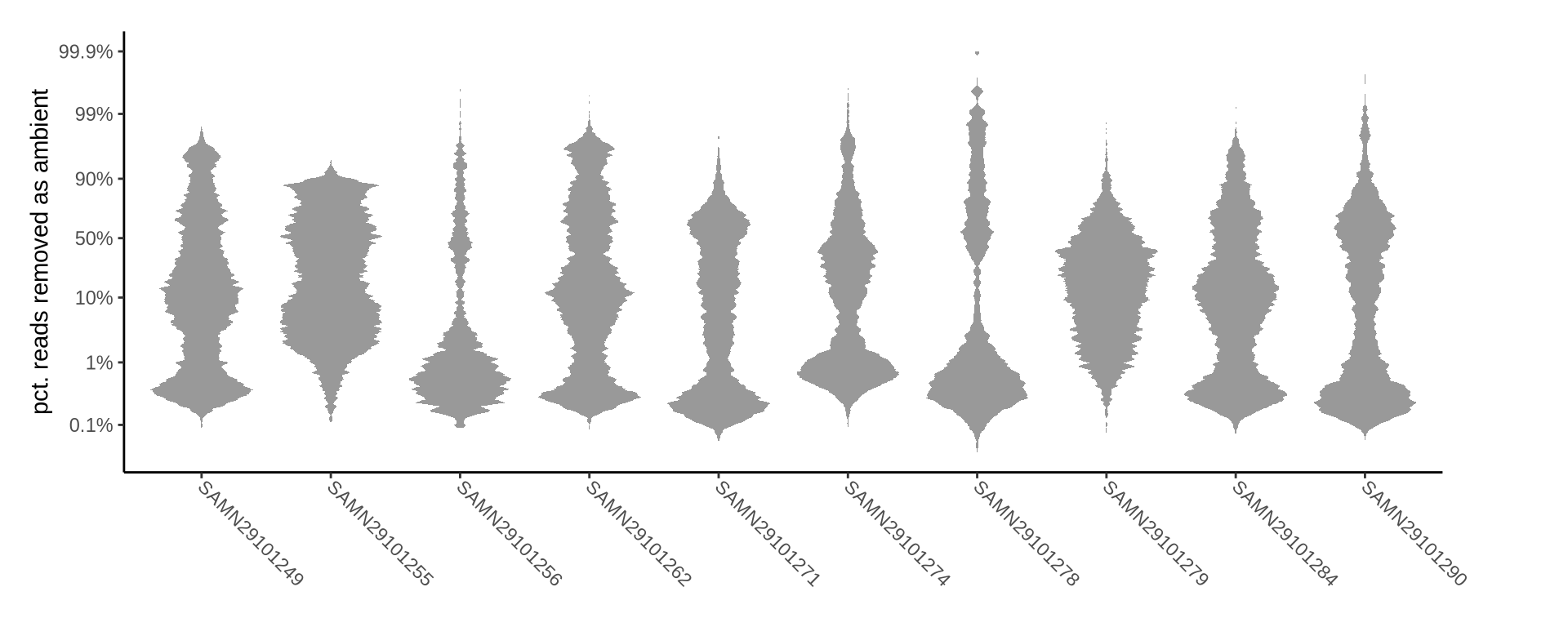
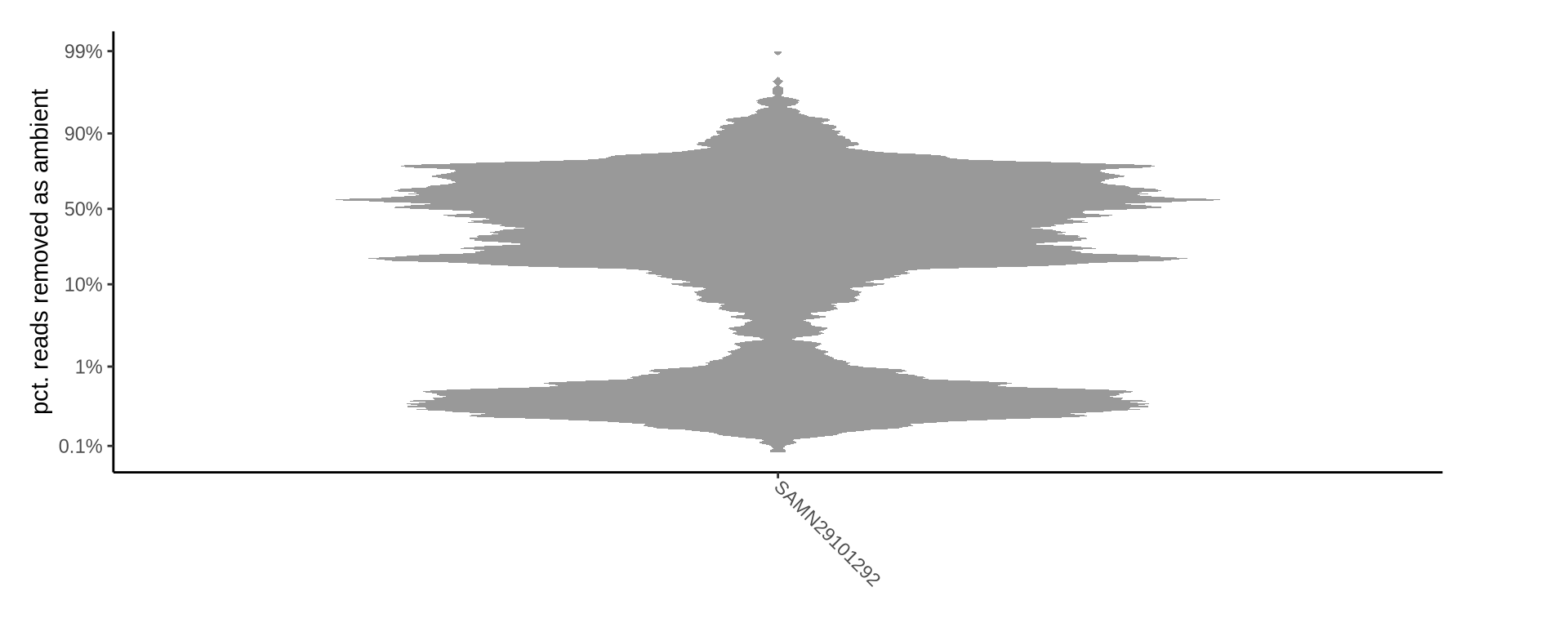
How many reads were removed as ambient?
Plots show what proportion of reads were removed from all barcodes called as cells.
for (ii in seq_along(chunks_ls)) {
chunk = chunks_ls[[ii]]
cat("### samples ", min(chunk), "-", max(chunk), "\n")
print(plot_reads_removed_as_ambient(usa_dt_ls[chunk], ok_bcs_ls[chunk]))
cat("\n\n")
}samples 1 - 10

samples 11 - 20

samples 21 - 21
Samples excluded by ambient removal step
CellBender includes all barcodes in the analysis up to the
total_droplets threshold covering both cell-containing
droplets and empty droplets. If CellBender calls the majority of these
included droplets as cells, it may indicate an underlying issue. This
typically occurs in low-quality samples where cell-containing barcodes
and empty droplets cannot be clearly distinguished in the barcode rank
plot. The table below shows the proportion of included droplets that
were classified as cells by CellBender. Samples where this proportion
exceeded % were excluded from further analysis.
calc_ambient_exclusions(stats_dt, run_var) %>% knitr::kable()| sample_id | total_droplets | kept_droplets | pct_kept | bad_run |
|---|---|---|---|---|
| SAMN29101206 | 31502 | 11215 | 35.6 | FALSE |
| SAMN29101222 | 30827 | 9997 | 32.4 | FALSE |
| SAMN29101223 | 29994 | 3873 | 12.9 | FALSE |
| SAMN29101226 | 28542 | 9092 | 31.9 | FALSE |
| SAMN29101227 | 34672 | 17569 | 50.7 | FALSE |
| SAMN29101231 | 34114 | 9491 | 27.8 | FALSE |
| SAMN29101235 | 30217 | 9934 | 32.9 | FALSE |
| SAMN29101238 | 28991 | 7576 | 26.1 | FALSE |
| SAMN29101245 | 30969 | 6730 | 21.7 | FALSE |
| SAMN29101247 | 29702 | 4553 | 15.3 | FALSE |
| SAMN29101249 | 30880 | 11836 | 38.3 | FALSE |
| SAMN29101255 | 32647 | 10274 | 31.5 | FALSE |
| SAMN29101256 | 26631 | 2210 | 8.3 | FALSE |
| SAMN29101262 | 30521 | 10031 | 32.9 | FALSE |
| SAMN29101271 | 35460 | 10099 | 28.5 | FALSE |
| SAMN29101274 | 33647 | 9469 | 28.1 | FALSE |
| SAMN29101278 | 27334 | 3934 | 14.4 | FALSE |
| SAMN29101279 | 28209 | 6383 | 22.6 | FALSE |
| SAMN29101284 | 32748 | 11180 | 34.1 | FALSE |
| SAMN29101290 | 27368 | 5559 | 20.3 | FALSE |
| SAMN29101292 | 27814 | 7708 | 27.7 | FALSE |
R session info
Details of the R package versions used are given below.
devtools::session_info()## ─ Session info ───────────────────────────────────────────────────────────────
## setting value
## version R version 4.3.3 (2024-02-29)
## os Red Hat Enterprise Linux 8.10 (Ootpa)
## system x86_64, linux-gnu
## ui X11
## language (EN)
## collate en_US.UTF-8
## ctype en_US.UTF-8
## tz Europe/Zurich
## date 2026-03-25
## pandoc 3.8.2.1 @ /home/macnairw/packages/scprocess/.snakemake/conda/fd31be42acf0c42e462db0f4d9adae50_/bin/ (via rmarkdown)
## quarto NA
##
## ─ Packages ───────────────────────────────────────────────────────────────────
## package * version date (UTC) lib source
## abind 1.4-5 2016-07-21 [2] CRAN (R 4.3.3)
## assertthat * 0.2.1 2019-03-21 [2] CRAN (R 4.3.3)
## beachmat 2.18.0 2023-10-24 [2] Bioconductor
## beeswarm 0.4.0 2021-06-01 [2] CRAN (R 4.3.3)
## Biobase * 2.62.0 2023-10-24 [2] Bioconductor
## BiocGenerics * 0.48.1 2023-11-01 [2] Bioconductor
## BiocManager 1.30.26 2025-06-05 [2] CRAN (R 4.3.3)
## BiocParallel * 1.36.0 2023-10-24 [2] Bioconductor
## BiocStyle * 2.30.0 2023-10-24 [2] Bioconductor
## bitops 1.0-9 2024-10-03 [2] CRAN (R 4.3.3)
## bookdown 0.44 2025-08-21 [2] CRAN (R 4.3.3)
## bslib 0.9.0 2025-01-30 [2] CRAN (R 4.3.3)
## ca 0.71.1 2020-01-24 [2] CRAN (R 4.3.3)
## cachem 1.1.0 2024-05-16 [2] CRAN (R 4.3.3)
## callr 3.7.6 2024-03-25 [2] CRAN (R 4.3.3)
## cellranger 1.1.0 2016-07-27 [2] CRAN (R 4.3.3)
## circlize * 0.4.16 2024-02-20 [2] CRAN (R 4.3.3)
## cli 3.6.5 2025-04-23 [2] CRAN (R 4.3.3)
## clue 0.3-66 2024-11-13 [2] CRAN (R 4.3.3)
## cluster 2.1.8.1 2025-03-12 [2] CRAN (R 4.3.3)
## codetools 0.2-20 2024-03-31 [2] CRAN (R 4.3.3)
## colorspace 2.1-1 2024-07-26 [2] CRAN (R 4.3.3)
## combinat 0.0-8 2012-10-29 [2] CRAN (R 4.3.3)
## ComplexHeatmap * 2.18.0 2023-10-24 [2] Bioconductor
## cowplot 1.2.0 2025-07-07 [2] CRAN (R 4.3.3)
## crayon 1.5.3 2024-06-20 [2] CRAN (R 4.3.3)
## data.table * 1.17.8 2025-07-10 [2] CRAN (R 4.3.3)
## decontX * 1.0.0 2023-10-24 [2] Bioconductor
## DelayedArray 0.28.0 2023-10-24 [2] Bioconductor
## DelayedMatrixStats 1.24.0 2023-10-24 [2] Bioconductor
## deldir 2.0-4 2024-02-28 [2] CRAN (R 4.3.3)
## DEoptimR 1.1-4 2025-07-27 [2] CRAN (R 4.3.3)
## devtools 2.4.5 2022-10-11 [2] CRAN (R 4.3.3)
## digest 0.6.37 2024-08-19 [2] CRAN (R 4.3.3)
## doParallel 1.0.17 2022-02-07 [2] CRAN (R 4.3.3)
## dotCall64 1.2 2024-10-04 [2] CRAN (R 4.3.3)
## dplyr * 1.1.4 2023-11-17 [2] CRAN (R 4.3.3)
## dqrng 0.3.2 2023-11-29 [2] CRAN (R 4.3.3)
## DropletUtils * 1.22.0 2023-10-24 [2] Bioconductor
## edgeR 4.0.16 2024-02-18 [2] Bioconductor 3.18 (R 4.3.3)
## ellipsis 0.3.2 2021-04-29 [2] CRAN (R 4.3.3)
## evaluate 1.0.5 2025-08-27 [2] CRAN (R 4.3.3)
## farver 2.1.2 2024-05-13 [2] CRAN (R 4.3.3)
## fastDummies 1.7.5 2025-01-20 [2] CRAN (R 4.3.3)
## fastmap 1.2.0 2024-05-15 [2] CRAN (R 4.3.3)
## fitdistrplus 1.2-4 2025-07-03 [2] CRAN (R 4.3.3)
## forcats * 1.0.0 2023-01-29 [2] CRAN (R 4.3.3)
## foreach 1.5.2 2022-02-02 [2] CRAN (R 4.3.3)
## fs 1.6.6 2025-04-12 [2] CRAN (R 4.3.3)
## future 1.67.0 2025-07-29 [2] CRAN (R 4.3.3)
## future.apply 1.20.0 2025-06-06 [2] CRAN (R 4.3.3)
## generics 0.1.4 2025-05-09 [2] CRAN (R 4.3.3)
## GenomeInfoDb * 1.38.1 2023-11-08 [2] Bioconductor
## GenomeInfoDbData 1.2.11 2026-03-05 [2] Bioconductor
## GenomicRanges * 1.54.1 2023-10-29 [2] Bioconductor
## GetoptLong 1.0.5 2020-12-15 [2] CRAN (R 4.3.3)
## getPass 0.2-4 2023-12-10 [2] CRAN (R 4.3.3)
## ggbeeswarm * 0.7.2 2023-04-29 [2] CRAN (R 4.3.3)
## ggh4x * 0.3.1 2025-05-30 [2] CRAN (R 4.3.3)
## ggplot2 * 3.5.2 2025-04-09 [2] CRAN (R 4.3.3)
## ggrepel * 0.9.6 2024-09-07 [2] CRAN (R 4.3.3)
## ggridges 0.5.7 2025-08-27 [2] CRAN (R 4.3.3)
## git2r 0.35.0 2024-10-20 [2] CRAN (R 4.3.3)
## GlobalOptions 0.1.2 2020-06-10 [2] CRAN (R 4.3.3)
## globals 0.18.0 2025-05-08 [2] CRAN (R 4.3.3)
## glue 1.8.0 2024-09-30 [2] CRAN (R 4.3.3)
## goftest 1.2-3 2021-10-07 [2] CRAN (R 4.3.3)
## gridExtra * 2.3 2017-09-09 [2] CRAN (R 4.3.3)
## gtable 0.3.6 2024-10-25 [2] CRAN (R 4.3.3)
## HDF5Array 1.30.0 2023-10-24 [2] Bioconductor
## hms 1.1.3 2023-03-21 [2] CRAN (R 4.3.3)
## htmltools 0.5.8.1 2024-04-04 [2] CRAN (R 4.3.3)
## htmlwidgets 1.6.4 2023-12-06 [2] CRAN (R 4.3.3)
## httpuv 1.6.16 2025-04-16 [2] CRAN (R 4.3.3)
## httr 1.4.7 2023-08-15 [2] CRAN (R 4.3.3)
## ica 1.0-3 2022-07-08 [2] CRAN (R 4.3.3)
## igraph 2.1.4 2025-01-23 [2] CRAN (R 4.3.3)
## inline 0.3.21 2025-01-09 [2] CRAN (R 4.3.3)
## IRanges * 2.36.0 2023-10-24 [2] Bioconductor
## irlba 2.3.5.1 2022-10-03 [2] CRAN (R 4.3.3)
## iterators 1.0.14 2022-02-05 [2] CRAN (R 4.3.3)
## jquerylib 0.1.4 2021-04-26 [2] CRAN (R 4.3.3)
## jsonlite 2.0.0 2025-03-27 [2] CRAN (R 4.3.3)
## KernSmooth 2.23-26 2025-01-01 [2] CRAN (R 4.3.3)
## knitr 1.50 2025-03-16 [2] CRAN (R 4.3.3)
## later 1.4.4 2025-08-27 [2] CRAN (R 4.3.3)
## lattice 0.22-7 2025-04-02 [2] CRAN (R 4.3.3)
## lazyeval 0.2.2 2019-03-15 [2] CRAN (R 4.3.3)
## lifecycle 1.0.4 2023-11-07 [2] CRAN (R 4.3.3)
## limma 3.58.1 2023-10-31 [2] Bioconductor
## listenv 0.9.1 2024-01-29 [2] CRAN (R 4.3.3)
## lmtest 0.9-40 2022-03-21 [2] CRAN (R 4.3.3)
## locfit 1.5-9.12 2025-03-05 [2] CRAN (R 4.3.3)
## loo 2.8.0 2024-07-03 [2] CRAN (R 4.3.3)
## lubridate * 1.9.4 2024-12-08 [2] CRAN (R 4.3.3)
## magrittr * 2.0.3 2022-03-30 [2] CRAN (R 4.3.3)
## MASS 7.3-60.0.1 2024-01-13 [2] CRAN (R 4.3.3)
## Matrix * 1.6-5 2024-01-11 [2] CRAN (R 4.3.3)
## MatrixGenerics * 1.14.0 2023-10-24 [2] Bioconductor
## matrixStats * 1.5.0 2025-01-07 [2] CRAN (R 4.3.3)
## MCMCprecision 0.4.2 2025-07-22 [2] CRAN (R 4.3.3)
## memoise 2.0.1 2021-11-26 [2] CRAN (R 4.3.3)
## mime 0.13 2025-03-17 [2] CRAN (R 4.3.3)
## miniUI 0.1.2 2025-04-17 [2] CRAN (R 4.3.3)
## nlme 3.1-168 2025-03-31 [2] CRAN (R 4.3.3)
## parallelly 1.45.1 2025-07-24 [2] CRAN (R 4.3.3)
## patchwork * 1.3.2 2025-08-25 [2] CRAN (R 4.3.3)
## pbapply 1.7-4 2025-07-20 [2] CRAN (R 4.3.3)
## pillar 1.11.0 2025-07-04 [2] CRAN (R 4.3.3)
## pkgbuild 1.4.8 2025-05-26 [2] CRAN (R 4.3.3)
## pkgconfig 2.0.3 2019-09-22 [2] CRAN (R 4.3.3)
## pkgload 1.4.0 2024-06-28 [2] CRAN (R 4.3.3)
## plotly 4.11.0 2025-06-19 [2] CRAN (R 4.3.3)
## plyr 1.8.9 2023-10-02 [2] CRAN (R 4.3.3)
## png 0.1-8 2022-11-29 [2] CRAN (R 4.3.3)
## polyclip 1.10-7 2024-07-23 [2] CRAN (R 4.3.3)
## processx 3.8.6 2025-02-21 [2] CRAN (R 4.3.3)
## profvis 0.4.0 2024-09-20 [2] CRAN (R 4.3.3)
## progressr 0.15.1 2024-11-22 [2] CRAN (R 4.3.3)
## promises 1.3.3 2025-05-29 [2] CRAN (R 4.3.3)
## ps 1.9.1 2025-04-12 [2] CRAN (R 4.3.3)
## purrr * 1.1.0 2025-07-10 [2] CRAN (R 4.3.3)
## QuickJSR 1.8.0 2025-06-09 [2] CRAN (R 4.3.3)
## R.methodsS3 1.8.2 2022-06-13 [2] CRAN (R 4.3.3)
## R.oo 1.27.1 2025-05-02 [2] CRAN (R 4.3.3)
## R.utils 2.13.0 2025-02-24 [2] CRAN (R 4.3.3)
## R6 2.6.1 2025-02-15 [2] CRAN (R 4.3.3)
## RANN 2.6.2 2024-08-25 [2] CRAN (R 4.3.3)
## RColorBrewer * 1.1-3 2022-04-03 [2] CRAN (R 4.3.3)
## Rcpp * 1.1.0 2025-07-02 [2] CRAN (R 4.3.3)
## RcppAnnoy 0.0.22 2024-01-23 [2] CRAN (R 4.3.3)
## RcppHNSW 0.6.0 2024-02-04 [2] CRAN (R 4.3.3)
## RcppParallel * 5.1.9 2024-08-19 [2] CRAN (R 4.3.3)
## RcppZiggurat * 0.1.8 2025-03-30 [2] CRAN (R 4.3.3)
## RCurl 1.98-1.17 2025-03-22 [2] CRAN (R 4.3.3)
## readr * 2.1.5 2024-01-10 [2] CRAN (R 4.3.3)
## readxl * 1.4.5 2025-03-07 [2] CRAN (R 4.3.3)
## registry 0.5-1 2019-03-05 [2] CRAN (R 4.3.3)
## remotes 2.5.0 2024-03-17 [2] CRAN (R 4.3.3)
## reshape2 1.4.4 2020-04-09 [2] CRAN (R 4.3.3)
## reticulate 1.43.0 2025-07-21 [2] CRAN (R 4.3.3)
## Rfast * 2.1.0 2023-11-09 [2] CRAN (R 4.3.3)
## rhdf5 * 2.46.1 2023-11-29 [2] Bioconductor 3.18 (R 4.3.3)
## rhdf5filters 1.14.1 2023-11-06 [2] Bioconductor
## Rhdf5lib 1.24.0 2023-10-24 [2] Bioconductor
## rjson 0.2.23 2024-09-16 [2] CRAN (R 4.3.3)
## rlang 1.1.6 2025-04-11 [2] CRAN (R 4.3.3)
## rmarkdown 2.29 2024-11-04 [2] CRAN (R 4.3.3)
## rmdformats 1.0.4 2022-05-17 [2] CRAN (R 4.3.3)
## robustbase * 0.99-6 2025-09-04 [2] CRAN (R 4.3.3)
## ROCR 1.0-11 2020-05-02 [2] CRAN (R 4.3.3)
## rprojroot 2.1.1 2025-08-26 [2] CRAN (R 4.3.3)
## RSpectra 0.16-2 2024-07-18 [2] CRAN (R 4.3.3)
## rstan 2.32.7 2025-03-10 [2] CRAN (R 4.3.3)
## rstantools 2.5.0 2025-09-01 [2] CRAN (R 4.3.3)
## rstudioapi 0.17.1 2024-10-22 [2] CRAN (R 4.3.3)
## Rtsne 0.17 2023-12-07 [2] CRAN (R 4.3.3)
## S4Arrays 1.2.0 2023-10-24 [2] Bioconductor
## S4Vectors * 0.40.2 2023-11-23 [2] Bioconductor 3.18 (R 4.3.3)
## sass 0.4.10 2025-04-11 [2] CRAN (R 4.3.3)
## scales * 1.4.0 2025-04-24 [2] CRAN (R 4.3.3)
## scattermore 1.2 2023-06-12 [2] CRAN (R 4.3.3)
## sctransform 0.4.2 2025-04-30 [2] CRAN (R 4.3.3)
## scuttle 1.12.0 2023-10-24 [2] Bioconductor
## seriation * 1.5.8 2025-08-20 [2] CRAN (R 4.3.3)
## sessioninfo 1.2.3 2025-02-05 [2] CRAN (R 4.3.3)
## Seurat 5.3.0 2025-04-23 [2] CRAN (R 4.3.3)
## SeuratObject 5.2.0 2025-08-27 [2] CRAN (R 4.3.3)
## shape 1.4.6.1 2024-02-23 [2] CRAN (R 4.3.3)
## shiny 1.11.1 2025-07-03 [2] CRAN (R 4.3.3)
## SingleCellExperiment * 1.24.0 2023-10-24 [2] Bioconductor
## sp 2.2-0 2025-02-01 [2] CRAN (R 4.3.3)
## spam 2.11-1 2025-01-20 [2] CRAN (R 4.3.3)
## SparseArray 1.2.2 2023-11-07 [2] Bioconductor
## sparseMatrixStats 1.14.0 2023-10-24 [2] Bioconductor
## spatstat.data 3.1-8 2025-08-18 [2] CRAN (R 4.3.3)
## spatstat.explore 3.5-2 2025-07-22 [2] CRAN (R 4.3.3)
## spatstat.geom 3.5-0 2025-07-20 [2] CRAN (R 4.3.3)
## spatstat.random 3.4-1 2025-05-20 [2] CRAN (R 4.3.3)
## spatstat.sparse 3.1-0 2024-06-21 [2] CRAN (R 4.3.3)
## spatstat.univar 3.1-4 2025-07-13 [2] CRAN (R 4.3.3)
## spatstat.utils 3.1-5 2025-07-17 [2] CRAN (R 4.3.3)
## StanHeaders 2.32.10 2024-07-15 [2] CRAN (R 4.3.3)
## statmod 1.5.0 2023-01-06 [2] CRAN (R 4.3.3)
## strex * 2.0.1 2024-10-03 [2] CRAN (R 4.3.3)
## stringi 1.8.7 2025-03-27 [2] CRAN (R 4.3.3)
## stringr * 1.5.2 2025-09-08 [2] CRAN (R 4.3.3)
## SummarizedExperiment * 1.32.0 2023-10-24 [2] Bioconductor
## survival 3.8-3 2024-12-17 [2] CRAN (R 4.3.3)
## tensor 1.5.1 2025-06-17 [2] CRAN (R 4.3.3)
## testit * 0.13 2021-04-14 [2] CRAN (R 4.3.3)
## tibble * 3.3.0 2025-06-08 [2] CRAN (R 4.3.3)
## tidyr * 1.3.1 2024-01-24 [2] CRAN (R 4.3.3)
## tidyselect 1.2.1 2024-03-11 [2] CRAN (R 4.3.3)
## tidyverse * 2.0.0 2023-02-22 [2] CRAN (R 4.3.3)
## timechange 0.3.0 2024-01-18 [2] CRAN (R 4.3.3)
## TSP 1.2-5 2025-05-27 [2] CRAN (R 4.3.3)
## tzdb 0.5.0 2025-03-15 [2] CRAN (R 4.3.3)
## urlchecker 1.0.1 2021-11-30 [2] CRAN (R 4.3.3)
## usethis 3.2.1 2025-09-06 [2] CRAN (R 4.3.3)
## uwot 0.2.3 2025-02-24 [2] CRAN (R 4.3.3)
## vctrs 0.6.5 2023-12-01 [2] CRAN (R 4.3.3)
## vipor 0.4.7 2023-12-18 [2] CRAN (R 4.3.3)
## viridis * 0.6.5 2024-01-29 [2] CRAN (R 4.3.3)
## viridisLite * 0.4.2 2023-05-02 [2] CRAN (R 4.3.3)
## whisker 0.4.1 2022-12-05 [2] CRAN (R 4.3.3)
## withr 3.0.2 2024-10-28 [2] CRAN (R 4.3.3)
## workflowr * 1.7.2 2025-08-18 [2] CRAN (R 4.3.3)
## xfun 0.53 2025-08-19 [2] CRAN (R 4.3.3)
## xtable 1.8-4 2019-04-21 [2] CRAN (R 4.3.3)
## XVector 0.42.0 2023-10-24 [2] Bioconductor
## yaml * 2.3.10 2024-07-26 [2] CRAN (R 4.3.3)
## zlibbioc 1.48.0 2023-10-24 [2] Bioconductor
## zoo 1.8-14 2025-04-10 [2] CRAN (R 4.3.3)
##
## [1] /home/macnairw/R/x86_64-conda-linux-gnu-library/4.3
## [2] /home/macnairw/packages/scprocess/.snakemake/conda/fd31be42acf0c42e462db0f4d9adae50_/lib/R/library
## * ── Packages attached to the search path.
##
## ──────────────────────────────────────────────────────────────────────────────